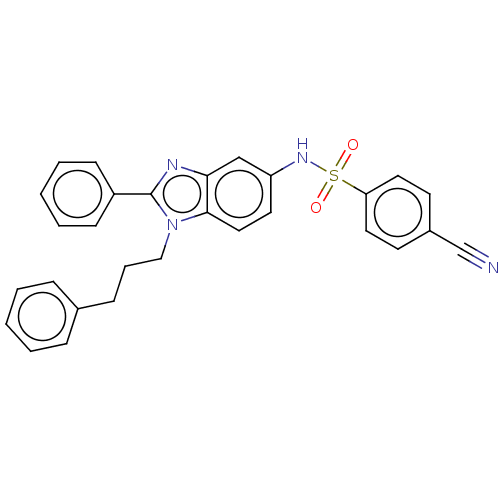
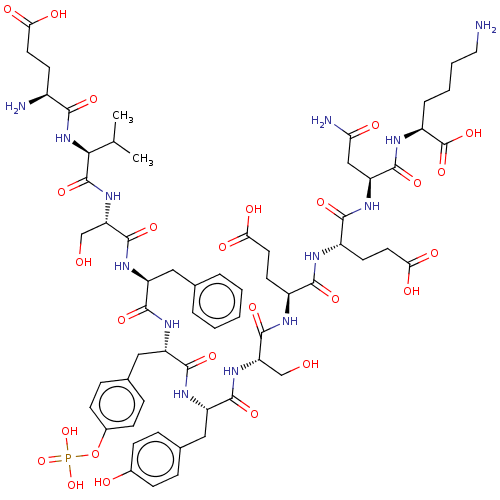
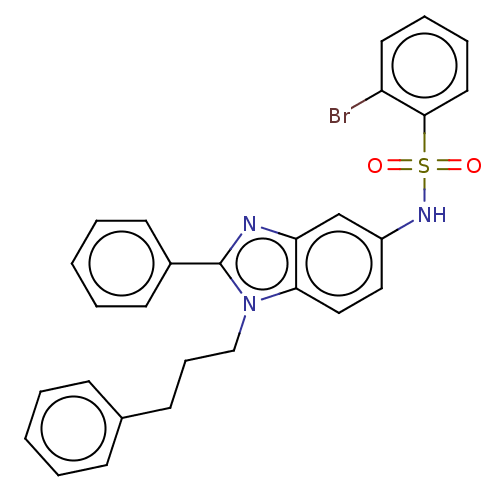
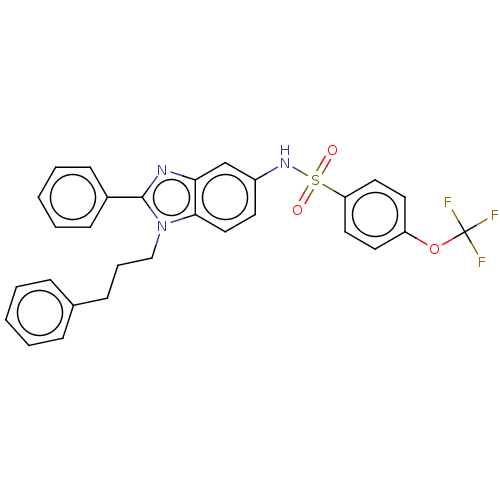
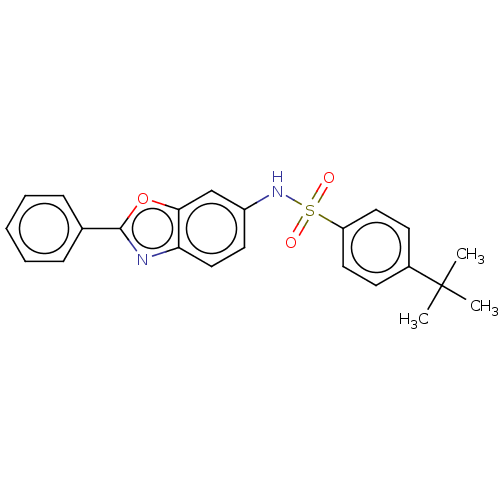
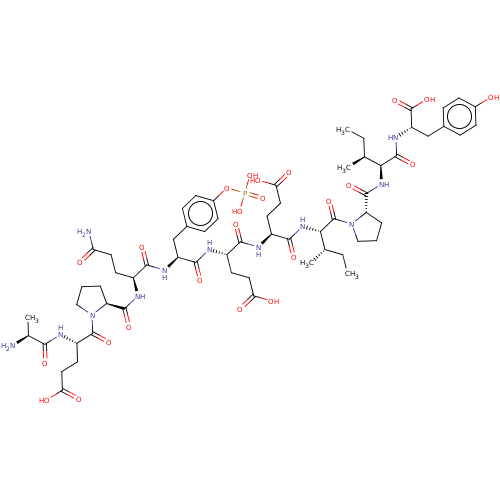
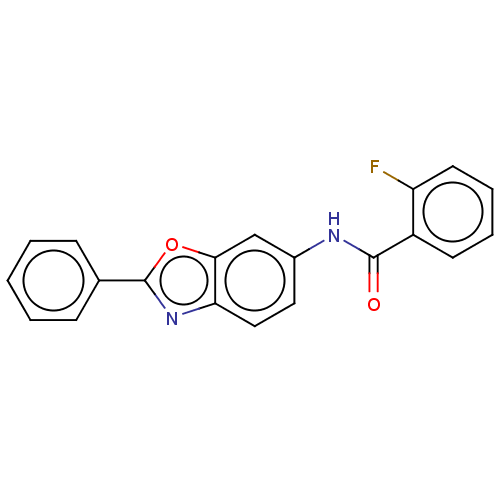
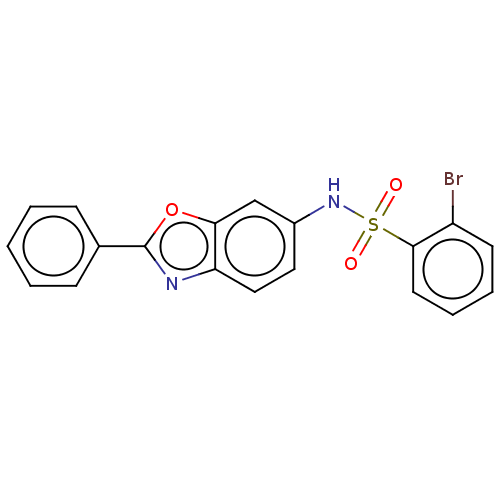
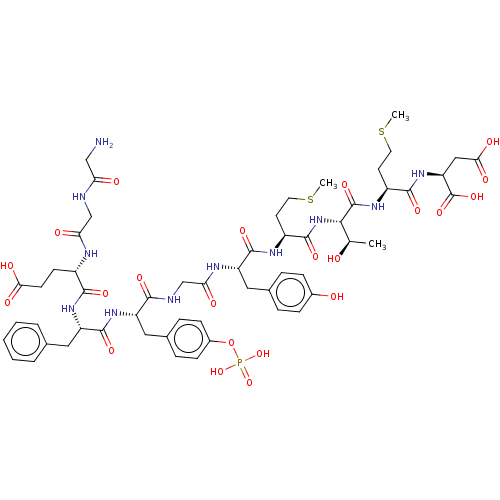
Report error Found 25 Enz. Inhib. hit(s) with all data for entry = 7947
Affinity DataIC50: 1.32E+3nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.75E+3nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 2.63E+3nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 6.10E+3nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 9.59E+3nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.28E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.63E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.88E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 2.08E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 2.19E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 2.62E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 3.04E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 3.32E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 3.54E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 3.63E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 8.5 T: 2°CAssay Description:Biochemical activity of G9a was measured as described [Kubicek et al., Mol. Cell, 25:473-481]. Assays were performed in white, opaque 384-well plates...More data for this Ligand-Target Pair