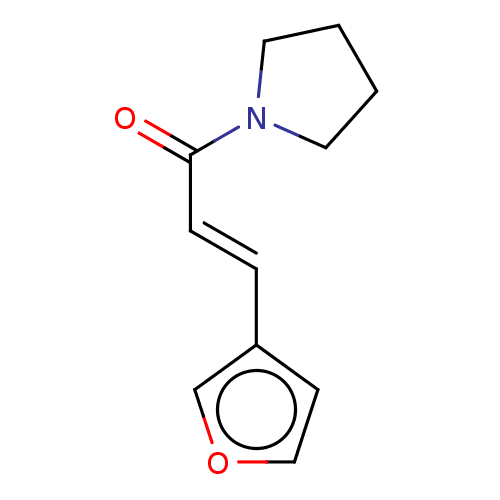
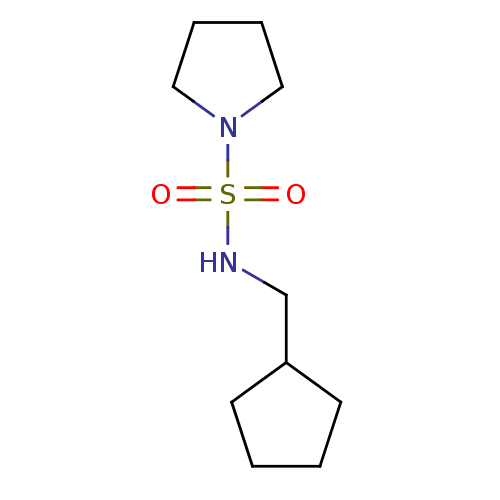
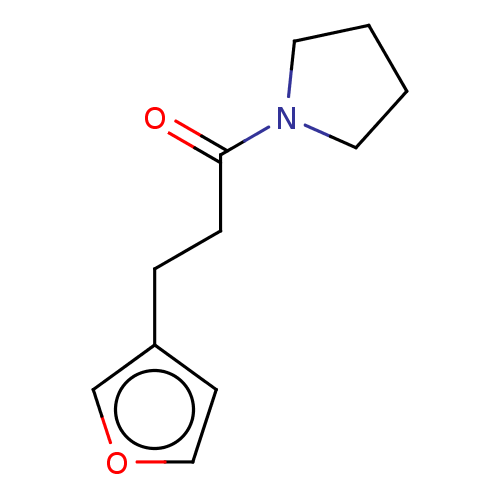
Report error Found 8 Enz. Inhib. hit(s) with all data for entry = 8024
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
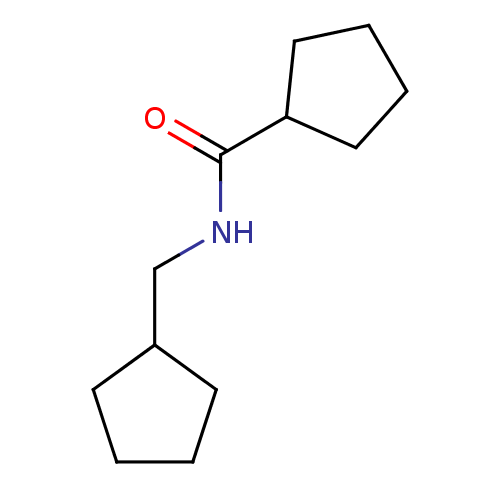
Affinity DataKd: 6.00E+3nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
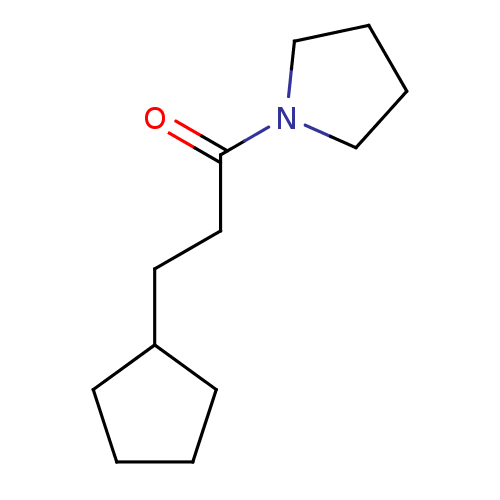
Affinity DataKd: 1.10E+4nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
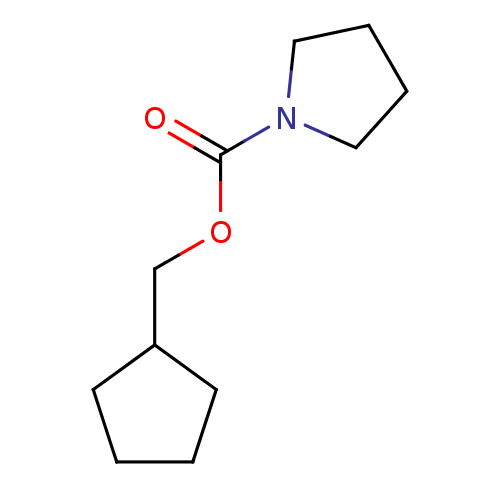
Affinity DataKd: 1.20E+4nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
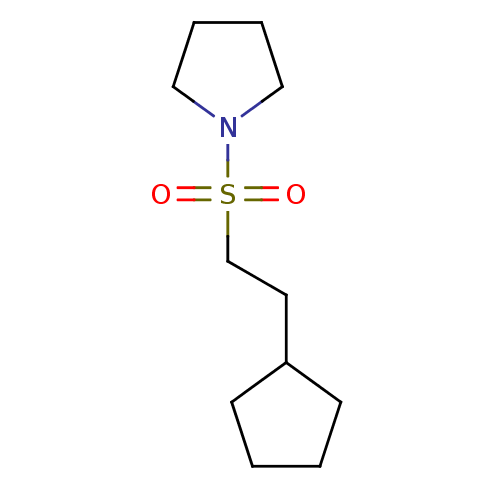
Affinity DataKd: 1.50E+4nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
Affinity DataKd: 1.60E+4nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
Affinity DataKd: 2.00E+4nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
Affinity DataKd: 2.00E+4nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
TargetHTH-type transcriptional regulator EthR(Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of Cambridge
University of Cambridge
Affinity DataKd: 2.20E+4nMAssay Description:An aqueous solution of the fragment to be tested (1 mM) was prepared containing NaCl (300 mM), Tris.HCl (20 mM, pH = 8.0), d6-DMSO (10% v/v) and glyc...More data for this Ligand-Target Pair
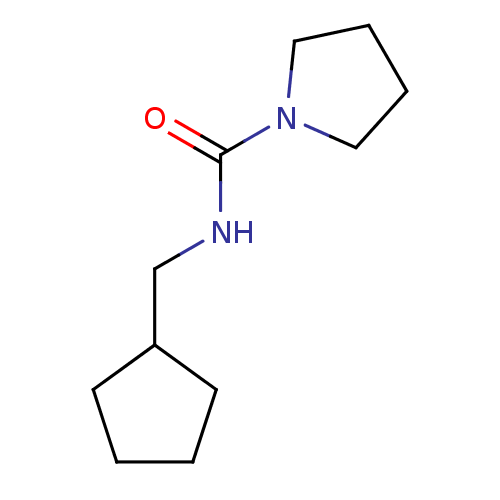
 3D Structure (crystal)
3D Structure (crystal)