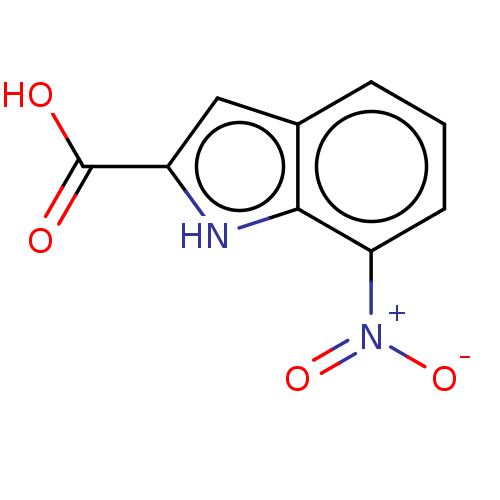
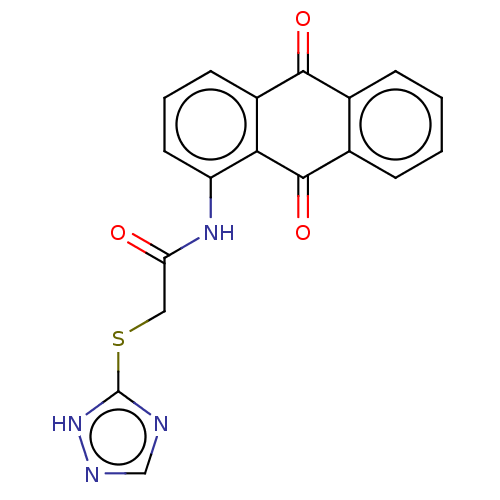
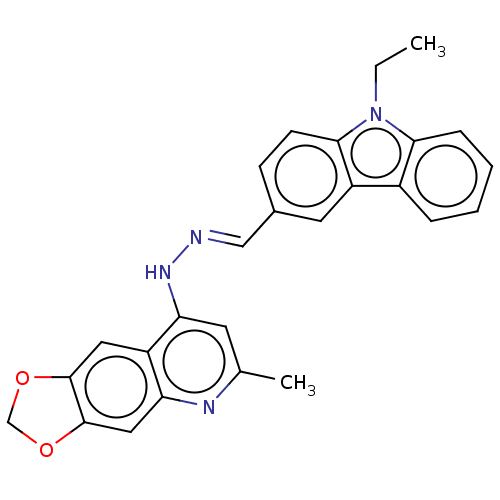
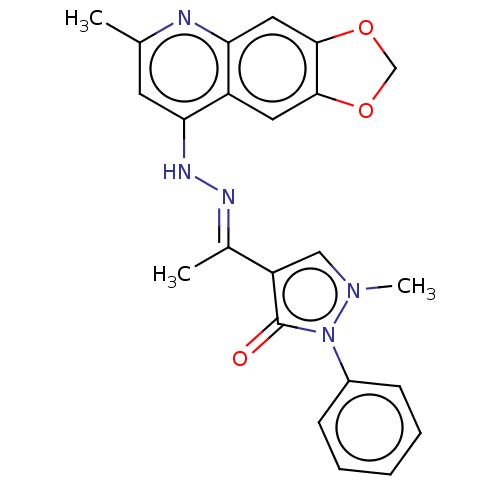
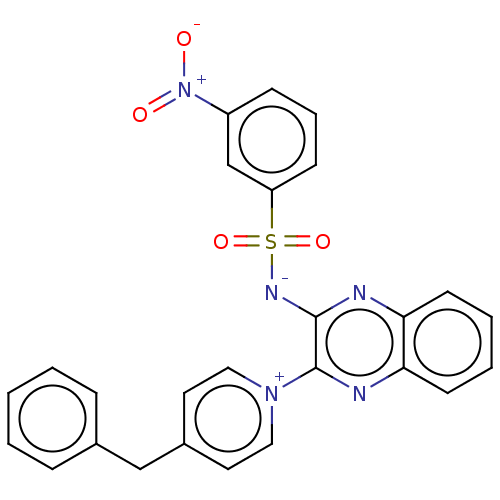
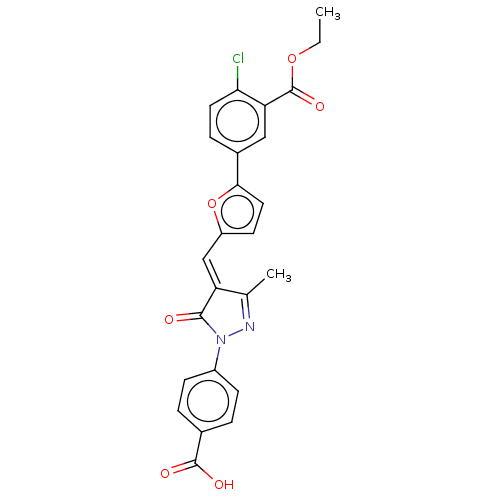
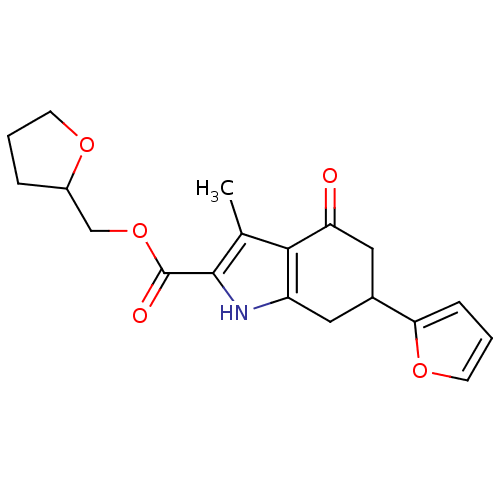
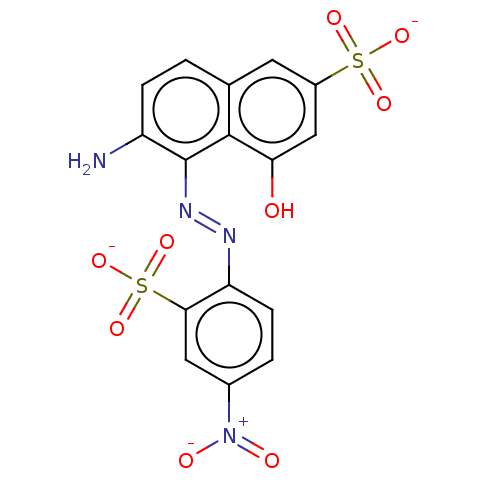
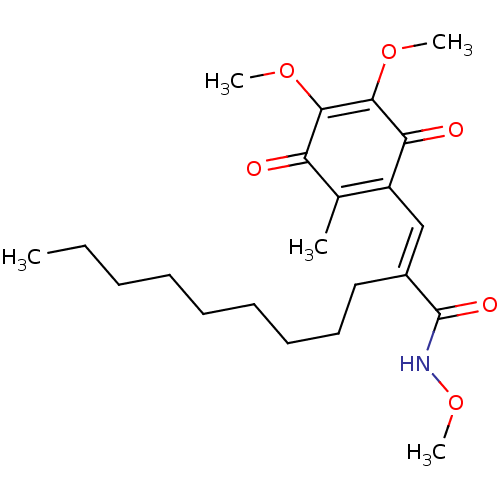
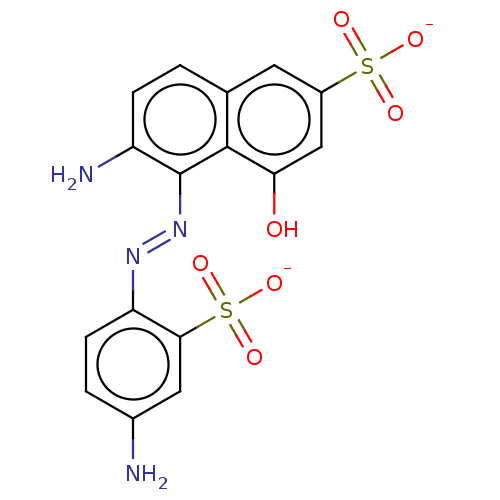
Report error Found 55 Enz. Inhib. hit(s) with all data for entry = 50002115
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataKd: 0.880nMAssay Description:Binding affinity to full length human APE1 expressed in Escherichia coli BL21/DE3More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataKd: 1.60nMAssay Description:Binding affinity to APE1 (unknown origin) by SPR assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataKd: 10nMAssay Description:Binding affinity to full length human APE1 expressed in Escherichia coli BL21/DE3More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 55nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 55nMAssay Description:Inhibition of human APE1 assessed as reduction in abasic sites DNA cleavage using 6-FAM/TAMRA labelled 17-Tphi as substrate by fluorescence-based ass...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 55nMAssay Description:Inhibition of APE1 (unknown origin)More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 57nMAssay Description:Inhibition of human APE1 assessed as reduction in abasic sites DNA cleavage using 6-FAM/TAMRA labelled 17-Tphi as substrate by fluorescence-based ass...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Inhibition of full length human APE1 expressed in Escherichia coli BL21/DE3 assessed as inhibition of incision of depurinated supercoiled pUC18 plasm...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataKd: 89nMAssay Description:Binding affinity to full length human APE1 expressed in Escherichia coli BL21/DE3More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Inhibition of recombinant human APE1 using double-stranded DNA as substrate preincubated for 15 mins followed by substrate addition measured at 1 min...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 320nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of human APE1 using a 5'-TAMRA labelled THF abasic site double-stranded oligodeoxynucleotide as substrate in presence of Mg2+ by fluoresce...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of human APE1 using a 5'-TAMRA labelled THF abasic site double-stranded oligodeoxynucleotide as substrate in presence of Mg2+ by fluoresce...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 800nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 900nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of human APE1 measured every 5 mins for 60 mins by FAM fluorophore based fluorescence assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of N-terminal hexa-His tagged human APE1 expressed in Escherichia coli BL21 (Rosetta) using fluorescein-dabcyl-containing oligonucleotide ...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of N-terminal hexa-His tagged human APE1 expressed in Escherichia coli BL21 (Rosetta) using fluorescein-dabcyl-containing oligonucleotide ...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of human APE1 measured every 5 mins for 60 mins by FAM fluorophore based fluorescence assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of recombinant human APE1 using a 5'-FAM/3'-Dabsyl labelled double-stranded DNA as substrate after 30 mins by fluorescence-based assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of recombinant human APE1 using 5'-F-GCCCCCXGGGGACGTACGATATCCCGCTCC-3' as substrate preincubated for 10 mins followed by substrate additio...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of human APE1 preincubated for 15 mins followed by substrate addition by HTS assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of nuclease activity of APE1 in human glioblastoma cells assessed as reduction in AP site DNA cleavageMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of N-terminal hexa-His tagged human APE1 expressed in Escherichia coli BL21 (Rosetta) using fluorescein-dabcyl-containing oligonucleotide ...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of human APE1 using a 5'-TAMRA labelled THF abasic site double-stranded oligodeoxynucleotide as substrate in presence of Mg2+ by fluoresce...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of recombinant His-6 tagged human APE1 expressed in Escherichia coli C41(DE3) using 5'-(F)-CGACTXTTGAATTGACACGCCATGTCGATCAATTCAATAGTCG-(D)...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human APE1 using 5'-F-GCCCCCXGGGGACGTACGATATCCCGCTCC-3' as substrate preincubated for 10 mins followed by substrate additio...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human APE1 using 5'-F-GCCCCCXGGGGACGTACGATATCCCGCTCC-3' as substrate preincubated for 10 mins followed by substrate additio...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 3.06E+3nMAssay Description:Inhibition of APE1 in human HeLa cells assessed as reduction in AP site cleavage 5'-[gamma-32P]CTCGCAAGUGGGTACCGA-3' as substrate after 1 hrMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of APE1 in human HeLa cellsMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibition of recombinant His-6 tagged human APE1 expressed in Escherichia coli C41(DE3) using 5'-(F)-CGACTXTTGAATTGACACGCCATGTCGATCAATTCAATAGTCG-(D)...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 4.40E+3nMAssay Description:Inhibition of recombinant His-6 tagged human APE1 expressed in Escherichia coli C41(DE3) using 5'-(F)-CGACTXTTGAATTGACACGCCATGTCGATCAATTCAATAGTCG-(D)...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of full length human APE1 expressed in Escherichia coli BL21/DE3 assessed as inhibition of incision of depurinated supercoiled pUC18 plasm...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 6.80E+3nMAssay Description:Inhibition of recombinant human APE1 using a 5'-FAM/3'-Dabsyl labelled double-stranded DNA as substrate after 30 mins by fluorescence-based assayMore data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 6.80E+3nMAssay Description:Inhibition of human APE1 using a 5'-TAMRA labelled THF abasic site double-stranded oligodeoxynucleotide as substrate in presence of Mg2+ by fluoresce...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 7.07E+3nMAssay Description:Inhibition of human APE1 using a 5'-TAMRA labelled THF abasic site double-stranded oligodeoxynucleotide as substrate in presence of Mg2+ by fluoresce...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 7.76E+3nMAssay Description:Inhibition of recombinant human APE1 using double-stranded DNA as substrate preincubated for 15 mins followed by substrate addition measured at 1 min...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 8.50E+3nMAssay Description:Inhibition of redox activity of full length N-terminal hexa-His SUMO-fused human APE1 using HEX-labeled THF oligonucleotide as substrate preincubated...More data for this Ligand-Target Pair
TargetDNA-(apurinic or apyrimidinic site) endonuclease(Human)
Russian Academy of Sciences
Curated by ChEMBL
Russian Academy of Sciences
Curated by ChEMBL
Affinity DataIC50: 8.85E+3nMAssay Description:Inhibition of recombinant human APE1 using double-stranded DNA as substrate preincubated for 15 mins followed by substrate addition measured at 1 min...More data for this Ligand-Target Pair