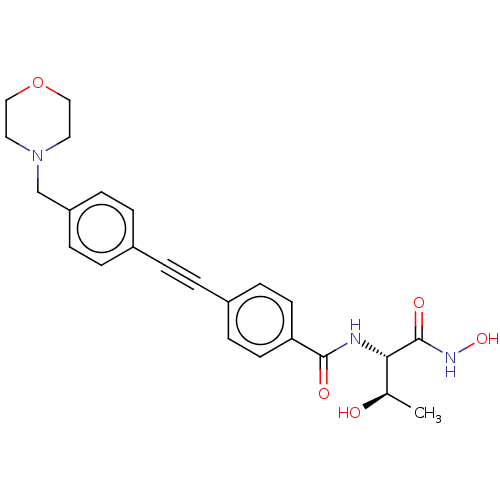
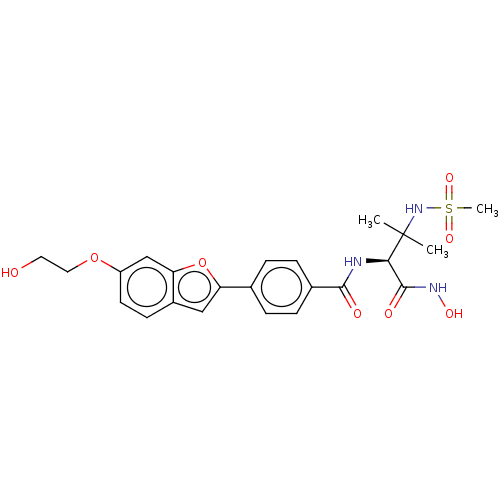
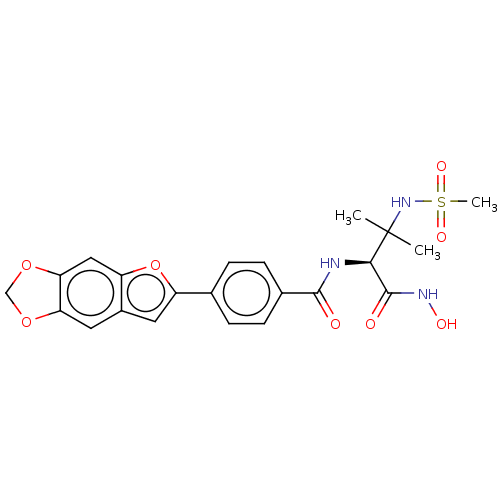
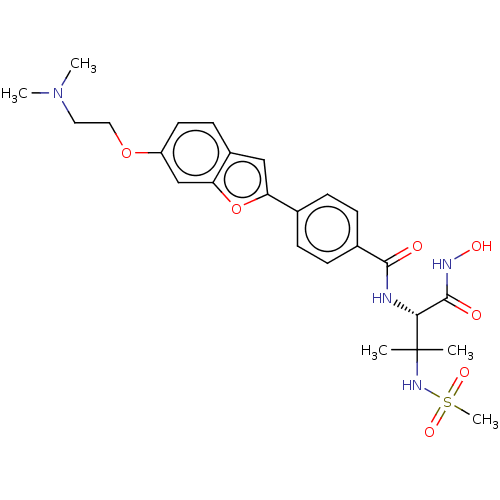
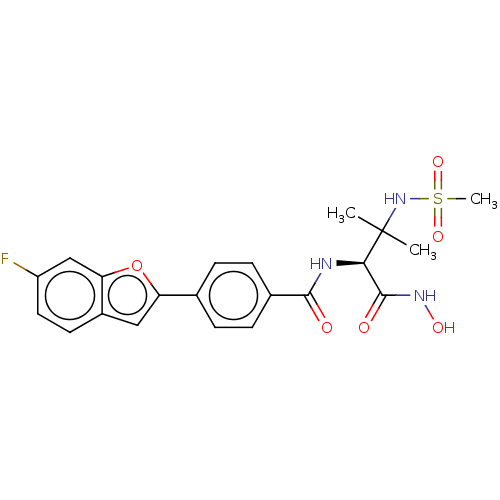
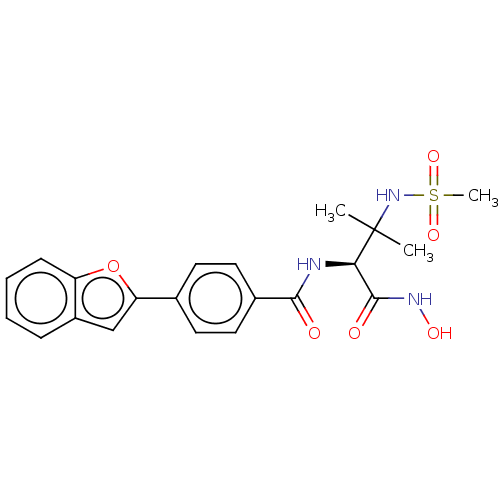
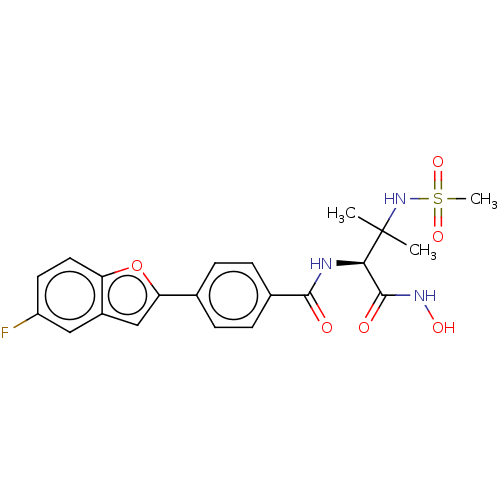
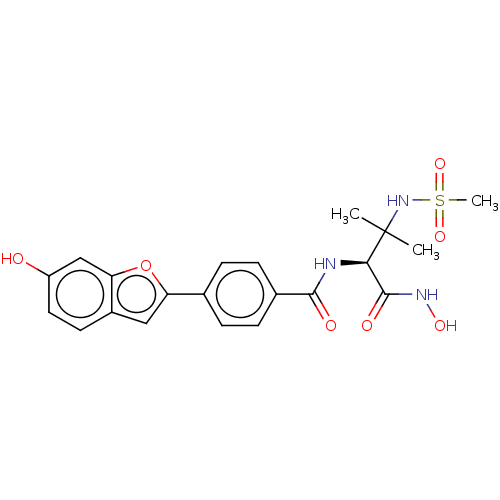
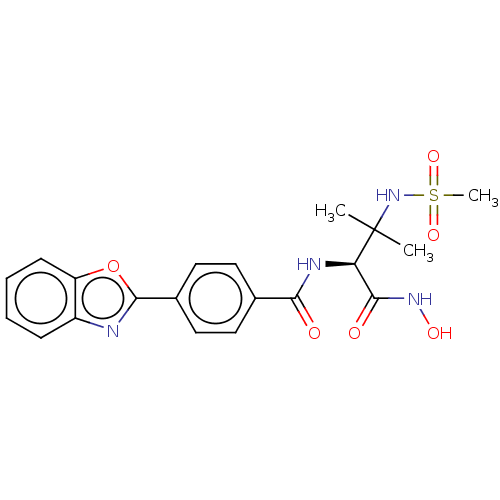
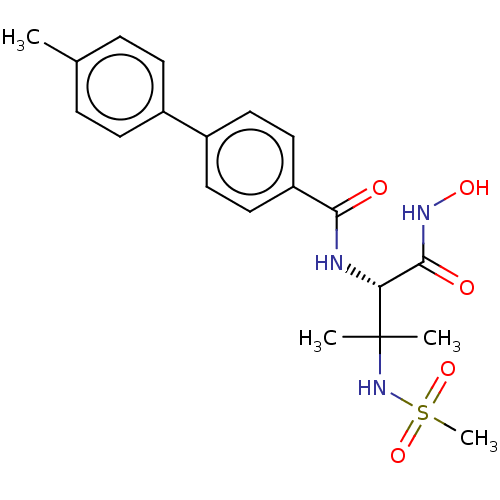
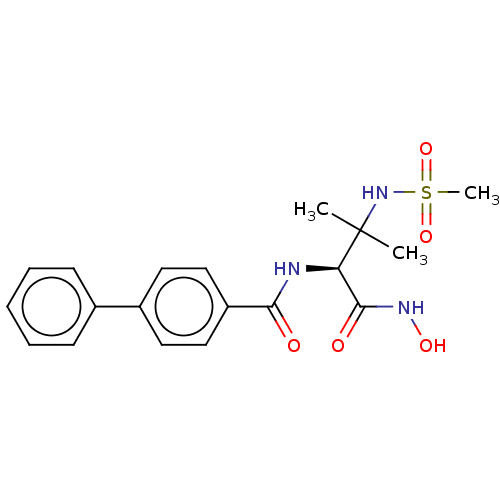
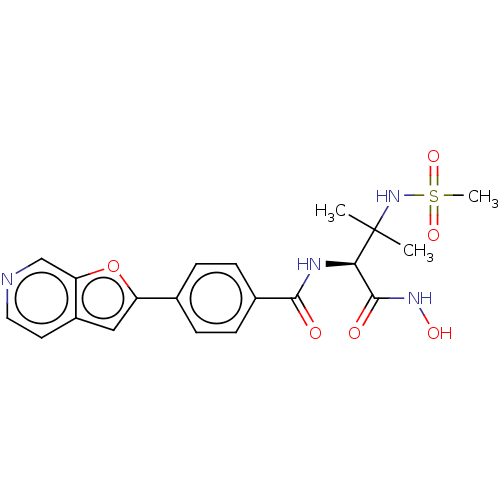
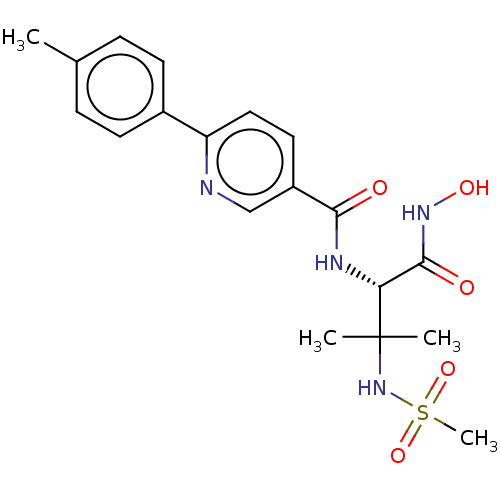
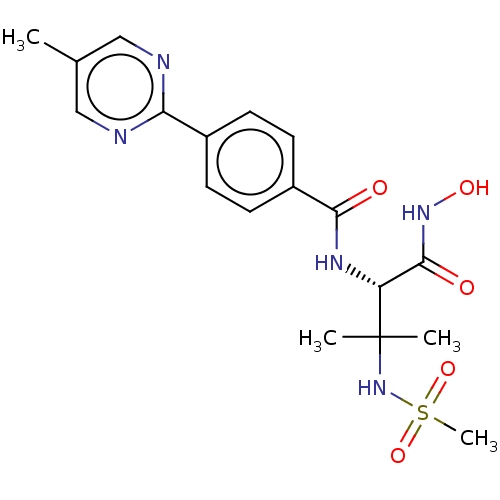
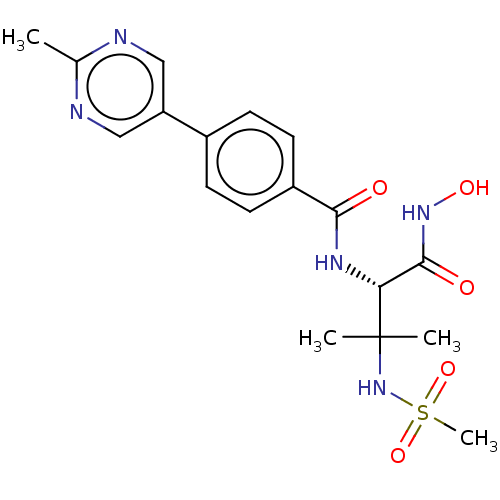
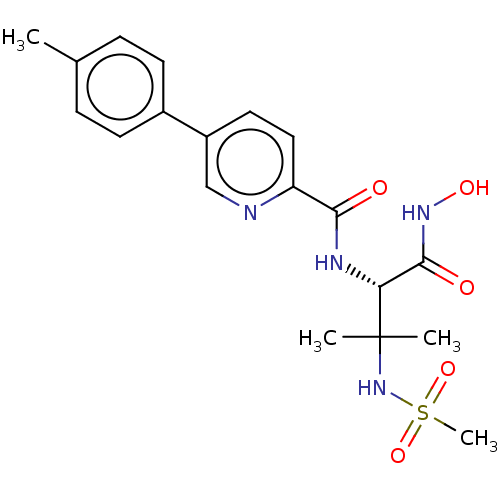
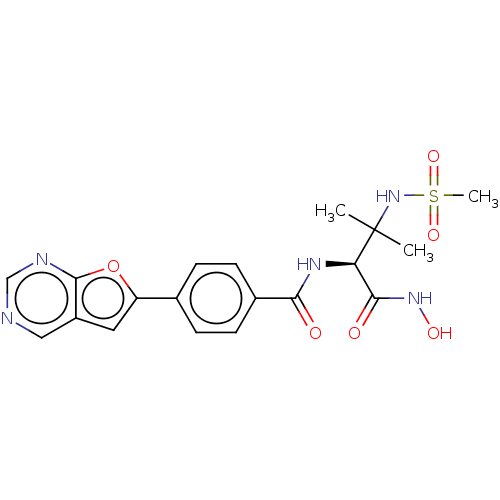
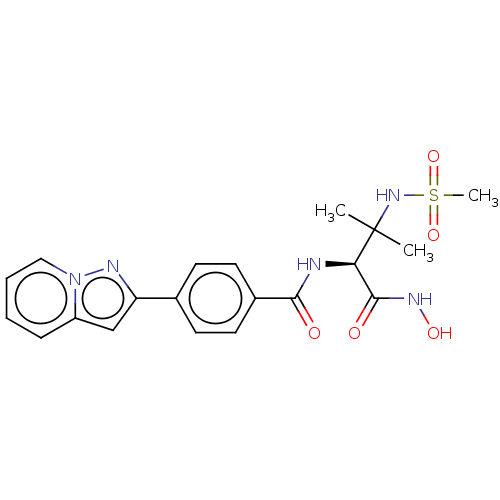
Report error Found 18 Enz. Inhib. hit(s) with all data for entry = 50006637
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 79nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 81nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 140nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 410nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 420nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Kyorin Pharmaceutical
Curated by ChEMBL
Kyorin Pharmaceutical
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-(R-3-hydroxymyristoyl)GlcNAc as substrate after 60 mins by OPA reagent based fluorescence assayMore data for this Ligand-Target Pair