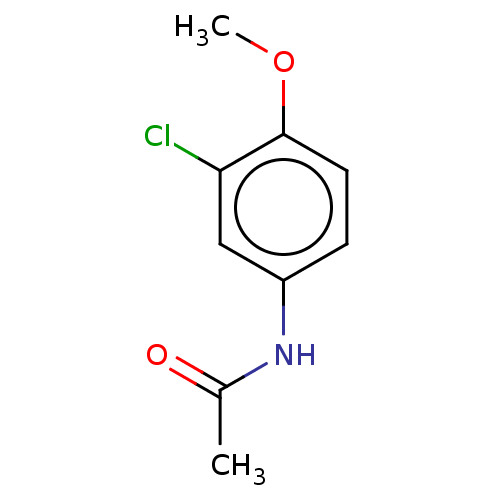
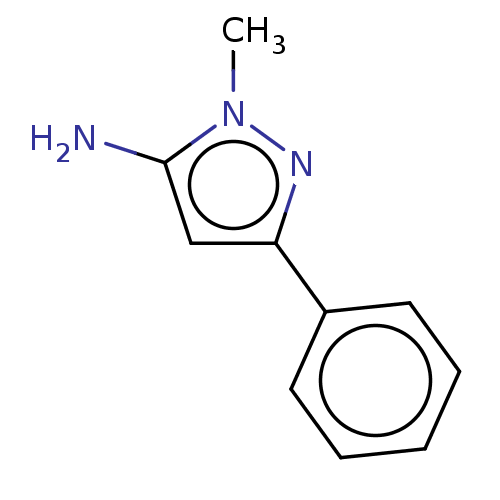
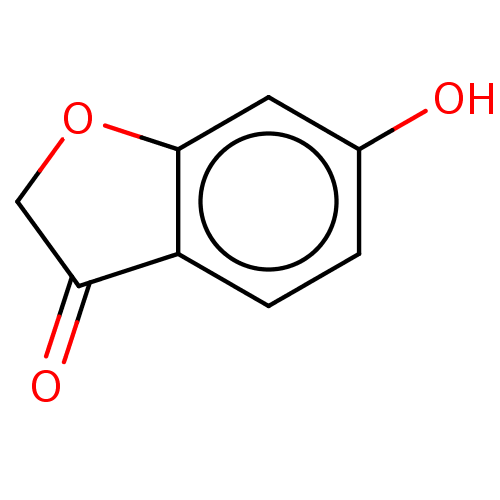
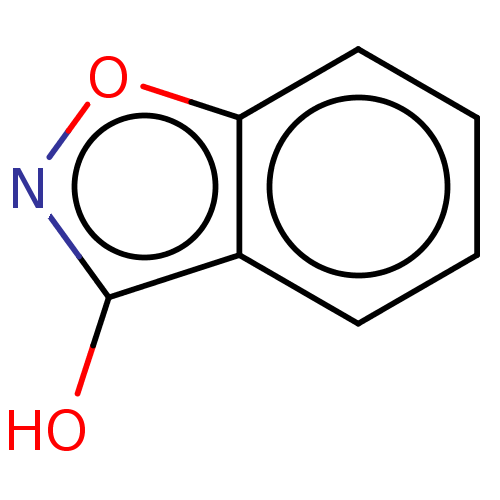
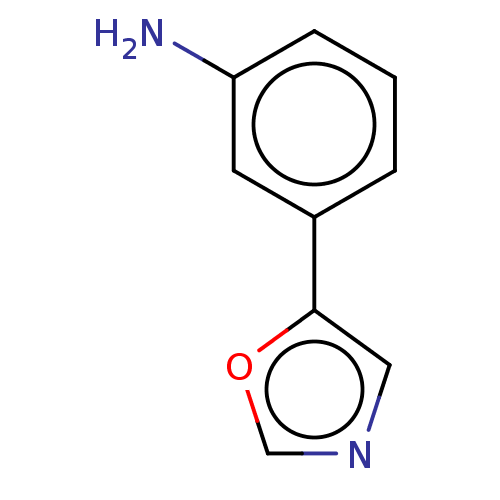
Report error Found 39 Enz. Inhib. hit(s) with all data for entry = 50049012
Affinity DataKd: 180nMAssay Description:Binding affinity to PHGDH (unknown origin) by surface plasma resonance methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of PHGDH (unknown origin) by FLINT methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of human liver N-terminal His6-tagged PHGDH expressed in Escherichia coli Rosetta (DE3)pLysS using 3 -phosphoglycerate as substarte measur...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMAssay Description:Inhibition of human liver N-terminal His6-tagged PHGDH expressed in Escherichia coli Rosetta (DE3)pLysS using 3 -phosphoglycerate as substarte measur...More data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+3nMAssay Description:Inhibition of human liver N-terminal His6-tagged PHGDH expressed in Escherichia coli Rosetta (DE3)pLysS using 3 -phosphoglycerate as substarte measur...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of PHGDH (unknown origin) expressed in Escherichia coli BL21 using 3-phosphoglycerate as substrate preincubated for 4 hrs followed by subs...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+4nMAssay Description:Inhibition of human liver N-terminal His6-tagged PHGDH expressed in Escherichia coli Rosetta (DE3)pLysS using 3 -phosphoglycerate as substarte measur...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+4nMAssay Description:Inhibition of PHGDH (unknown origin) expressed in Escherichia coli BL21 using 3 -phosphoglycerate as substrate preincubated for 30 mins followed by s...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+4nMAssay Description:Inhibition of human liver N-terminal His6-tagged PHGDH expressed in Escherichia coli Rosetta (DE3)pLysS using 3 -phosphoglycerate as substarte measur...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+4nMAssay Description:Inhibition of human liver N-terminal His6-tagged PHGDH expressed in Escherichia coli Rosetta (DE3)pLysS using 3 -phosphoglycerate as substarte measur...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of full length PHGDH (unknown origin) using 3-phosphoglycerate as substrate incubated for 60 mins in presence of PSAT1, PSPH by resazurin ...More data for this Ligand-Target Pair
Affinity DataKd: 1.50E+6nMAssay Description:Binding affinity to human N-terminal His6-tagged truncated PHGDH (3 to 314 residues) expressed in Escherichia coli Rosetta (DE3) by ITC assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.60E+6nMAssay Description:Binding affinity to human N-terminal His6-tagged truncated PHGDH (3 to 314 residues) expressed in Escherichia coli Rosetta (DE3) by ITC assayMore data for this Ligand-Target Pair
Affinity DataKd: 2.80E+6nMAssay Description:Binding affinity to human N-terminal His6-tagged truncated PHGDH (3 to 314 residues) expressed in Escherichia coli Rosetta (DE3) by ITC assayMore data for this Ligand-Target Pair
Affinity DataKd: 5.80E+6nMAssay Description:Binding affinity to human N-terminal His6-tagged truncated PHGDH (3 to 314 residues) expressed in Escherichia coli Rosetta (DE3) by ITC assayMore data for this Ligand-Target Pair
Affinity DataKd: 5.90E+6nMAssay Description:Binding affinity to human N-terminal His6-tagged truncated PHGDH (3 to 314 residues) expressed in Escherichia coli Rosetta (DE3) by ITC assayMore data for this Ligand-Target Pair
Affinity DataKd: 6.50E+6nMAssay Description:Binding affinity to human N-terminal His6-tagged truncated PHGDH (3 to 314 residues) expressed in Escherichia coli Rosetta (DE3) by ITC assayMore data for this Ligand-Target Pair
Affinity DataKd: 9.30E+6nMAssay Description:Binding affinity to human N-terminal His6-tagged truncated PHGDH (3 to 314 residues) expressed in Escherichia coli Rosetta (DE3) by ITC assayMore data for this Ligand-Target Pair
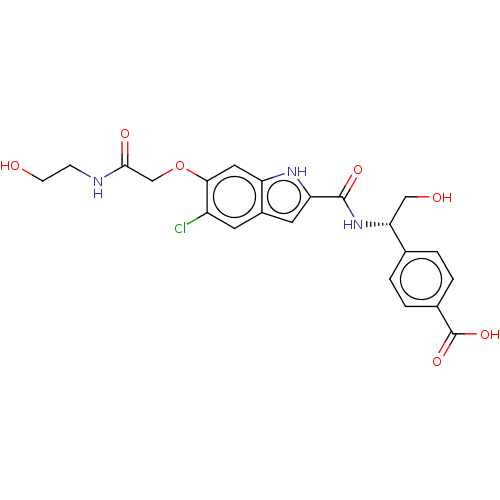
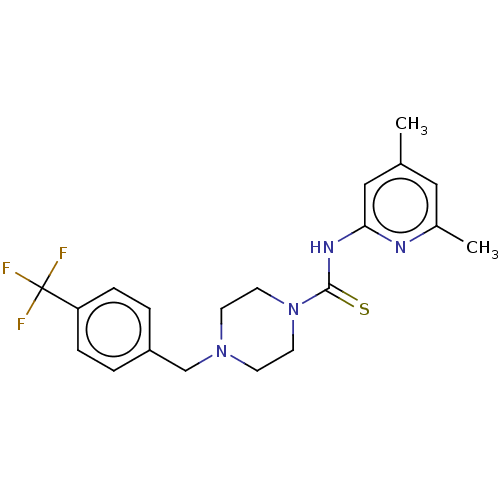
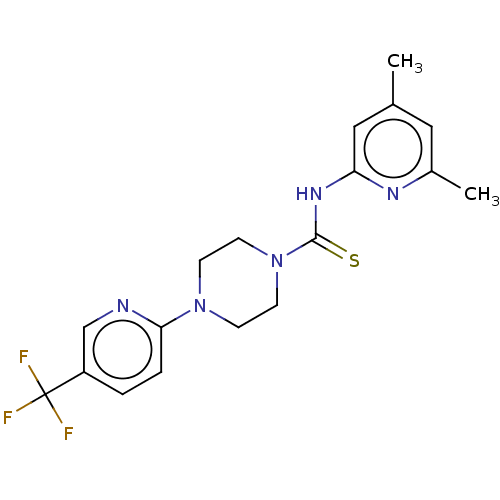
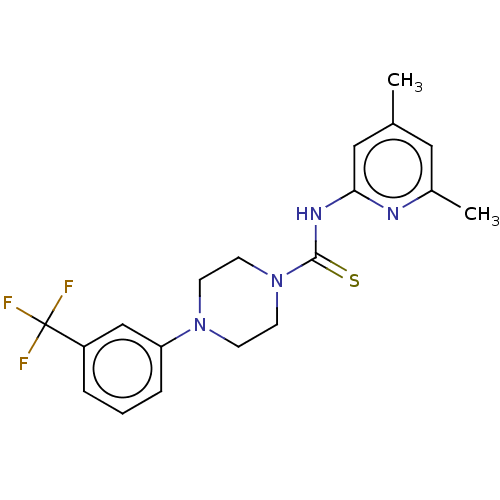
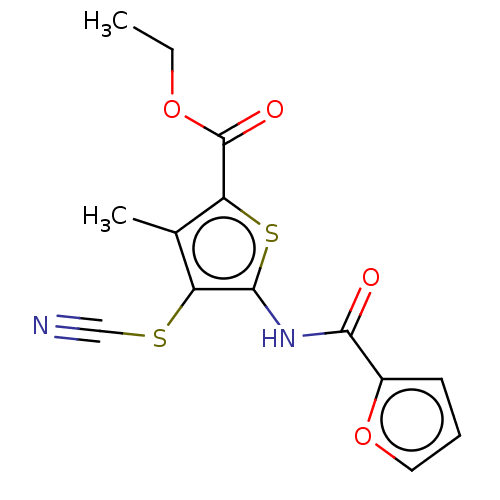
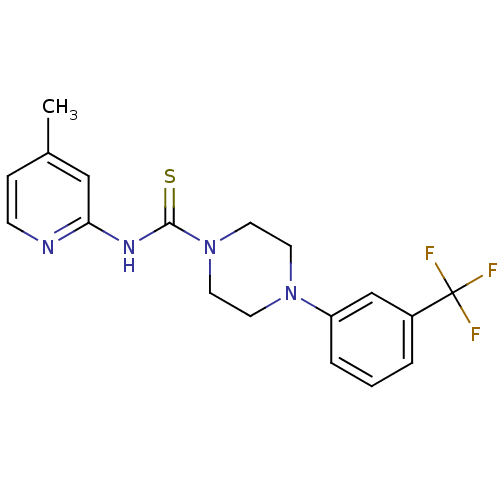
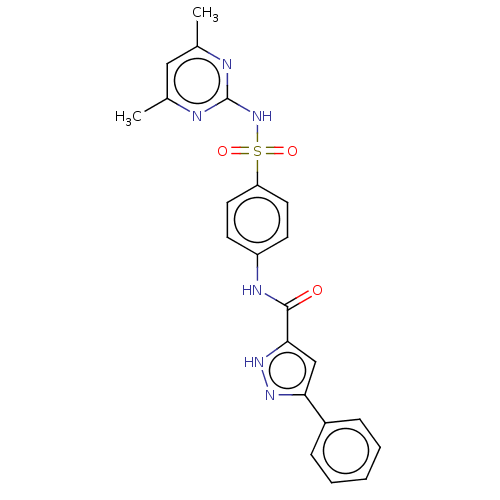
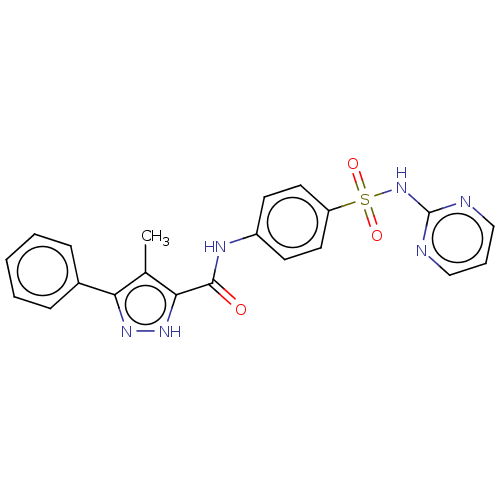
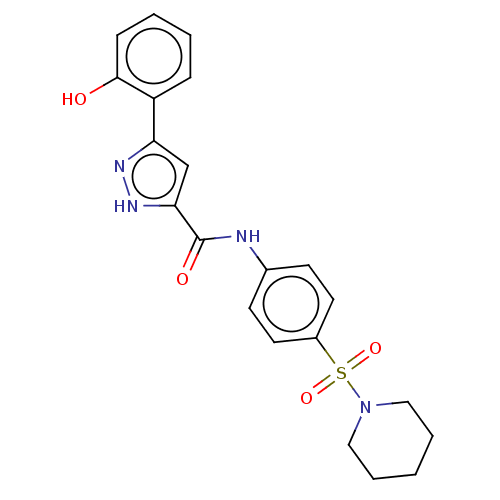
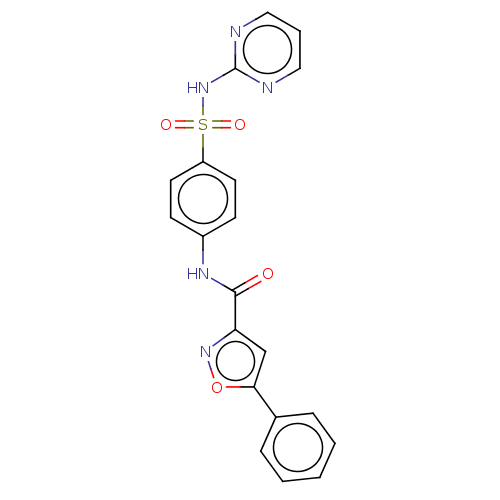
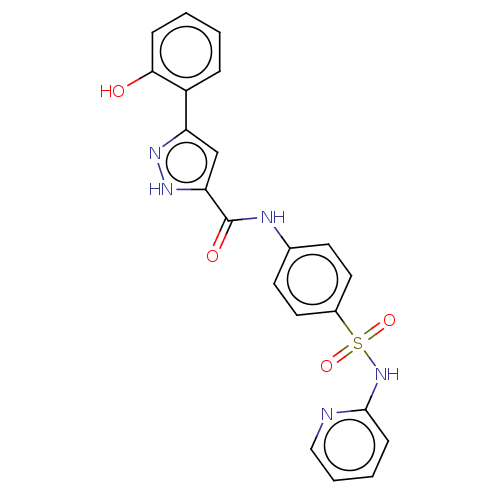
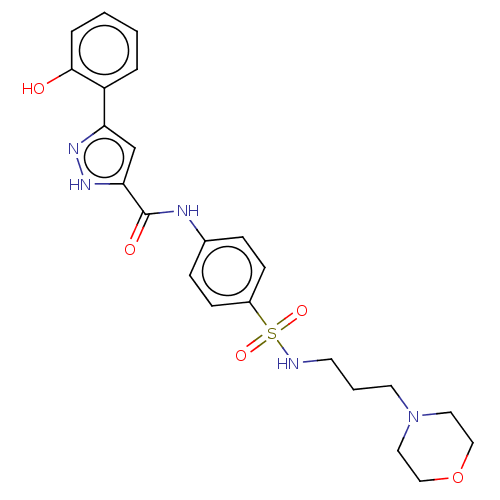
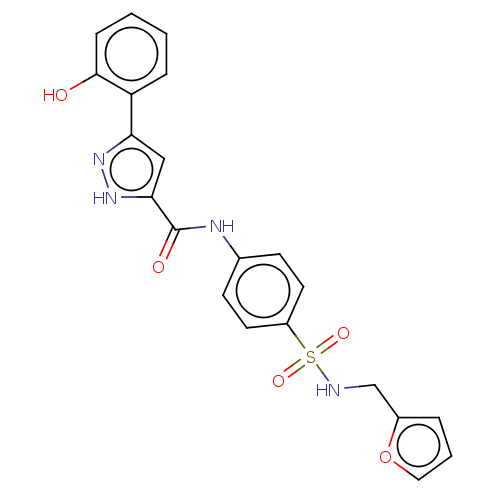
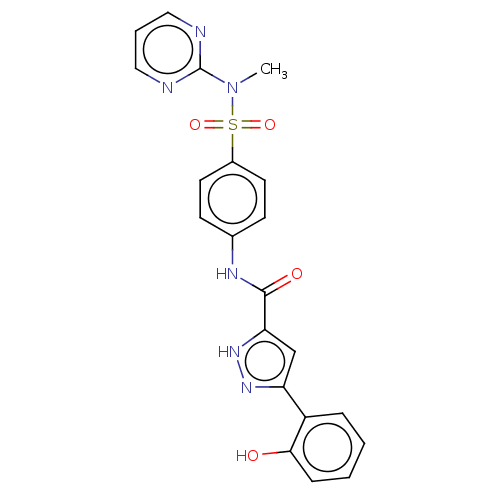
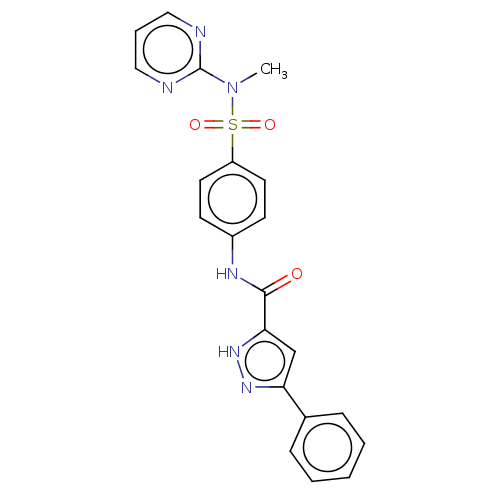
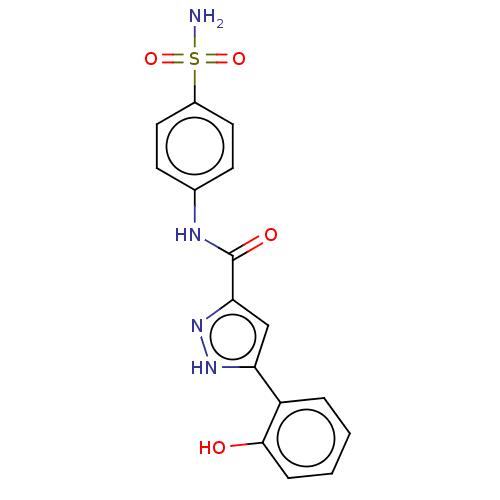
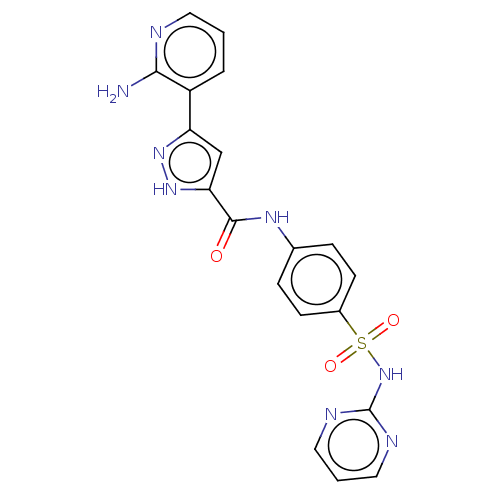
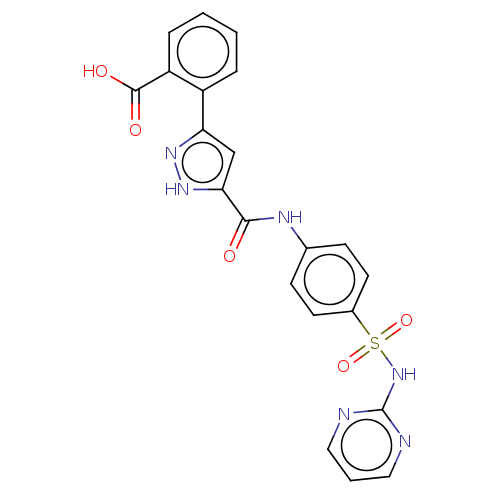
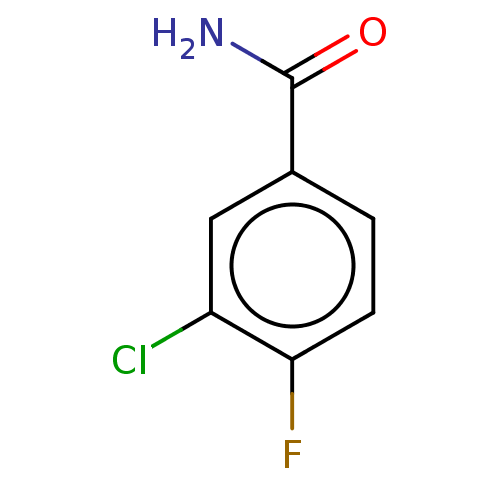
 3D Structure (crystal)
3D Structure (crystal)