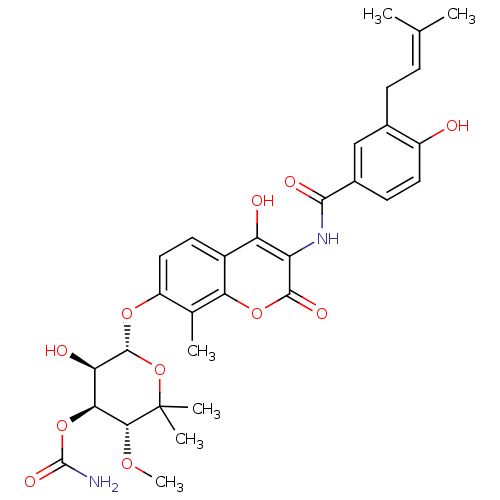
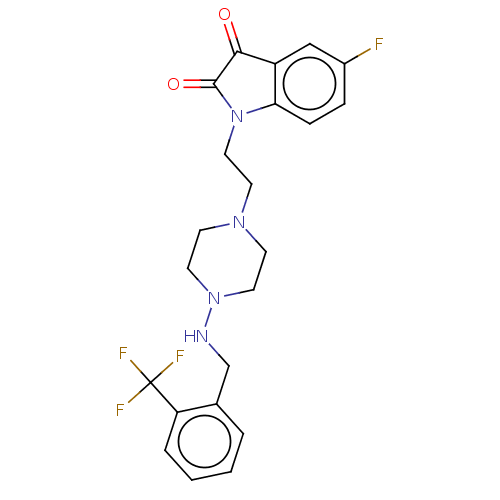
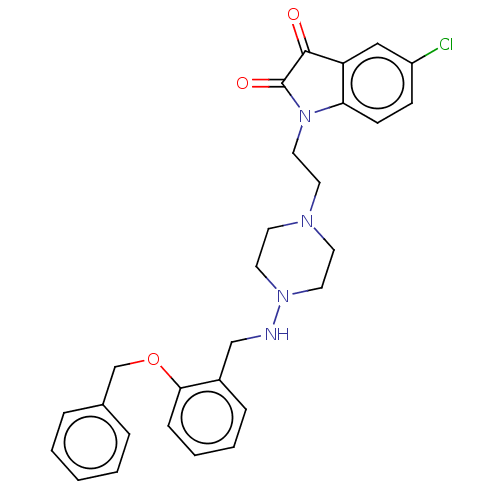
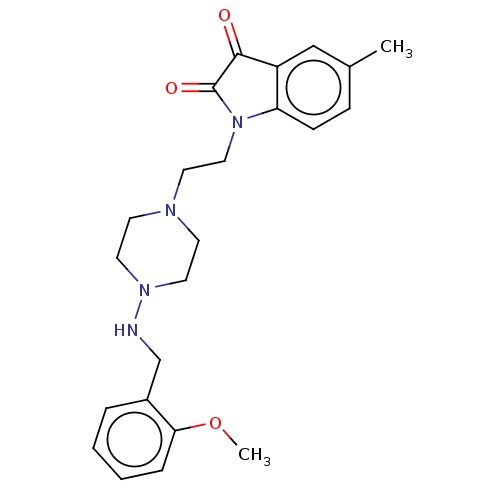
Report error Found 30 Enz. Inhib. hit(s) with all data for entry = 50048209
TargetDNA gyrase subunit B(Escherichia coli (strain K12))
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataKi: 7nMAssay Description:Inhibition of Escherichia coli H560 DNA gyrase B ATPase activity after 30 mins by ammonium molybdate/malachite green-based phosphate detection assayMore data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Escherichia coli (strain K12))
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataKd: 7nMAssay Description:Binding affinity to Escherichia coli H560 DNA gyrase BMore data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 1.06E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 1.38E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 1.41E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 1.58E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 2.02E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 2.03E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 2.17E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 2.21E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 2.66E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 2.83E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 4.30E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 4.32E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 4.75E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 5.10E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 5.36E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 6.15E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 6.80E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 6.97E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 7.01E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 7.30E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 7.60E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 7.60E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 7.90E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 8.12E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 8.30E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 8.40E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 9.20E+4nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair
TargetDNA gyrase subunit B(Mycobacterium smegmatis)
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
R. C. Patel Institute of Pharmaceutical Education and Research
Curated by ChEMBL
Affinity DataIC50: 1.80E+5nMAssay Description:Inhibition of Mycobacterium smegmatis DNA gyrase B expressed in BL21(DE3)pLysS cells assessed as inhibition of DNA supercoiling activity in presence ...More data for this Ligand-Target Pair