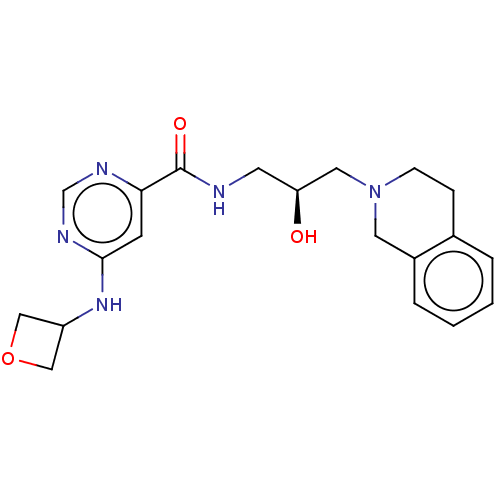
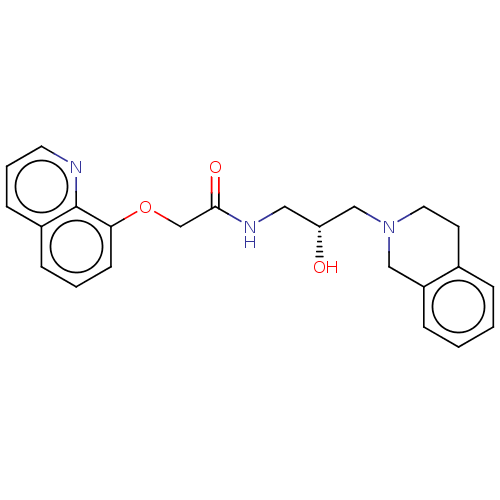
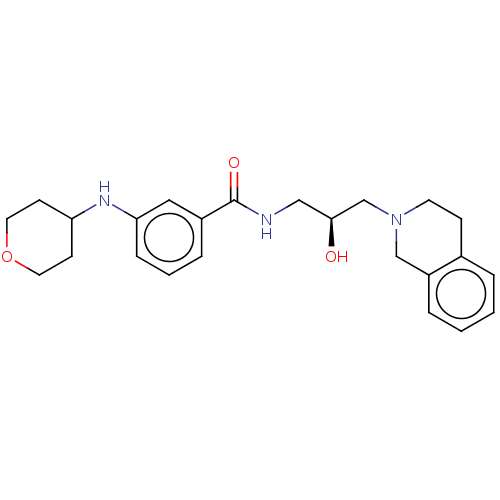
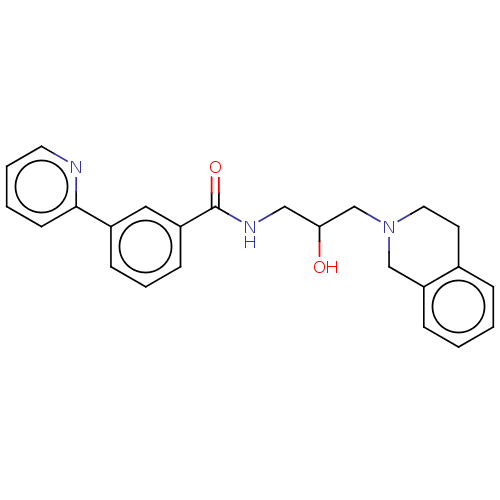
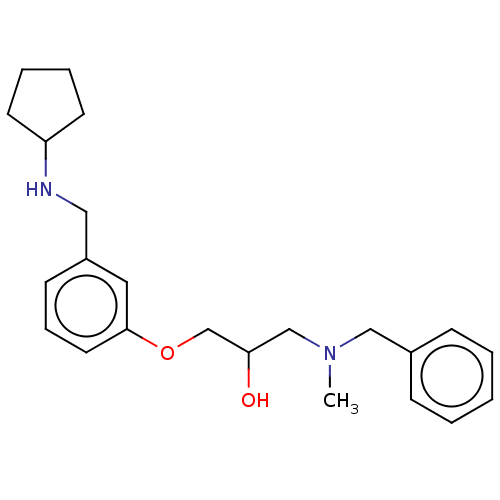
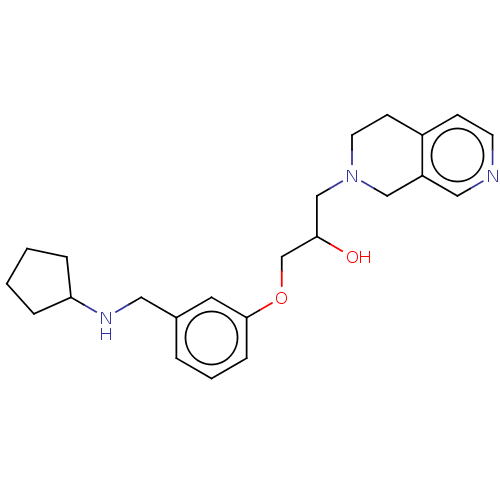
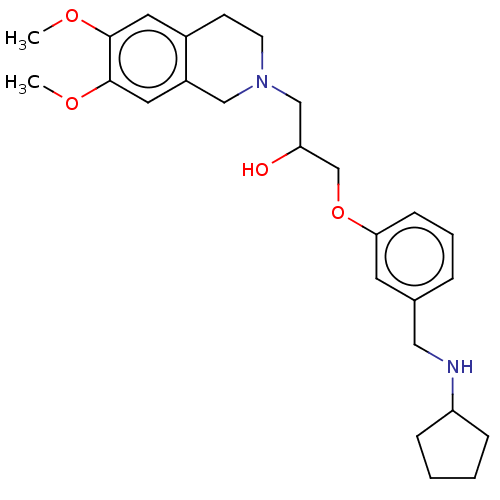
Report error Found 27 Enz. Inhib. hit(s) with all data for entry = 50011699
Affinity DataIC50: 3nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 73nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 95nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 144nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 326nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 327nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 406nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 435nMAssay Description:Inhibition of PRMT5 in human Z138 cells assessed as reduction in symmetrical dimethylation of arginine containing substrate using SmD3 as substrate i...More data for this Ligand-Target Pair
Affinity DataIC50: 7.45E+3nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.68E+4nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of human full length FLAG-tagged PRMT5/human His6-tagged MEP50 expressed in baculovirus-infected Sf9 cells assessed as reduction in tritiu...More data for this Ligand-Target Pair
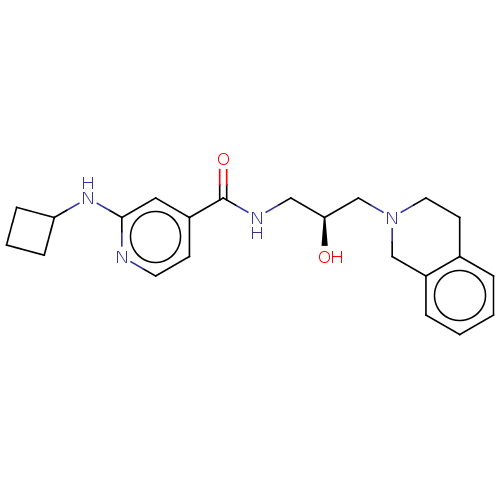
 3D Structure (crystal)
3D Structure (crystal)