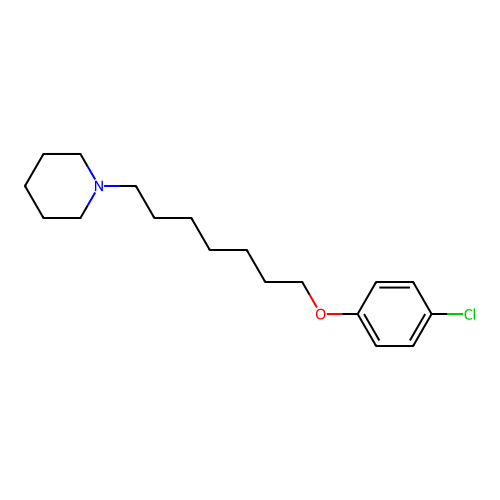
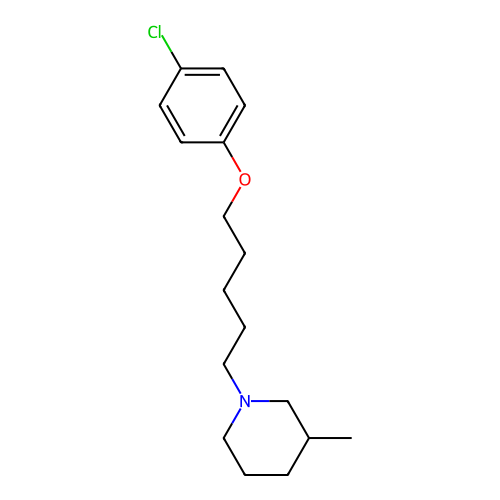
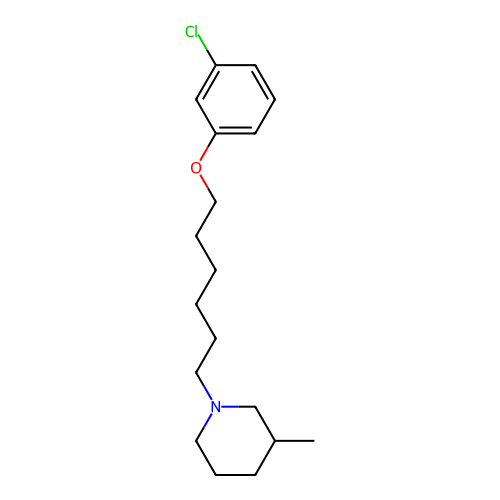
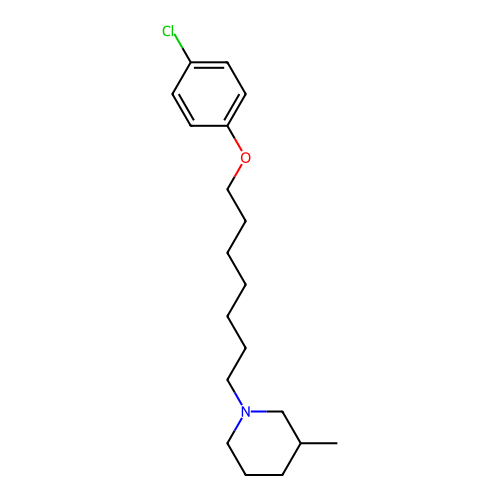
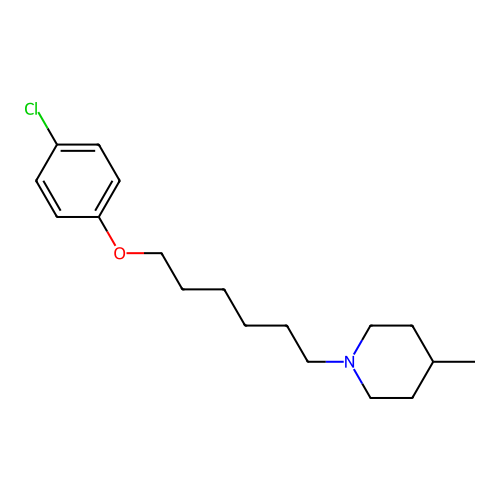
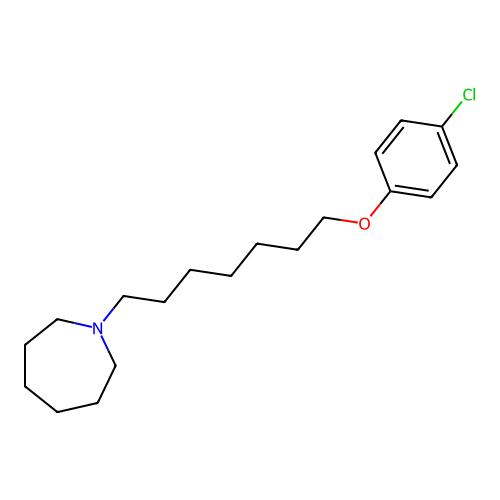
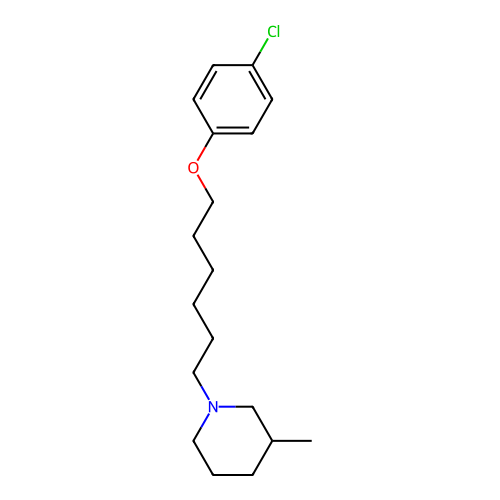
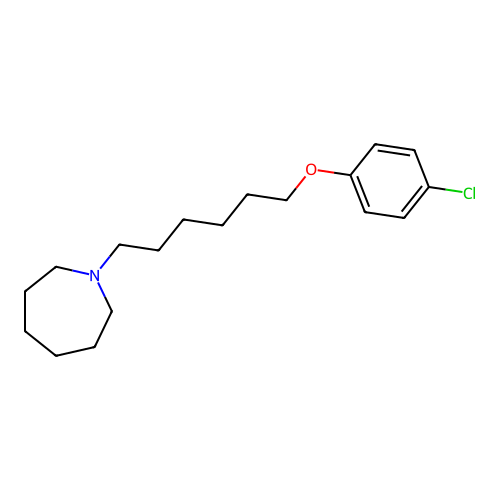
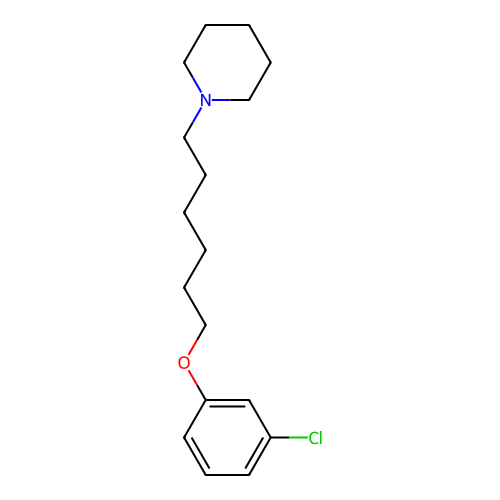
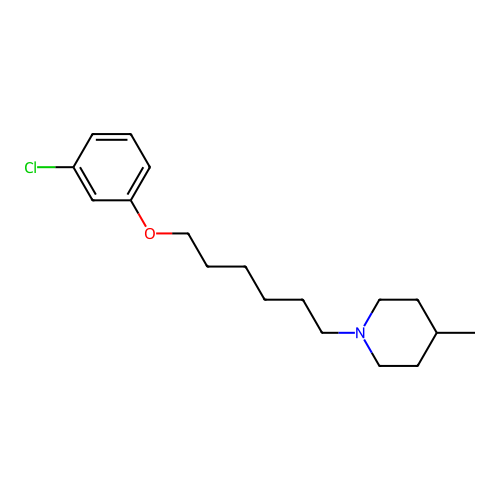
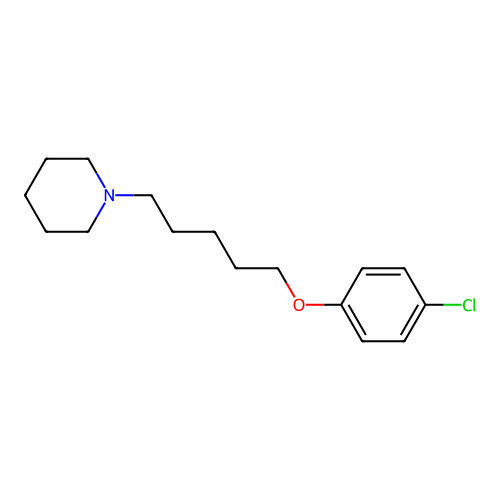
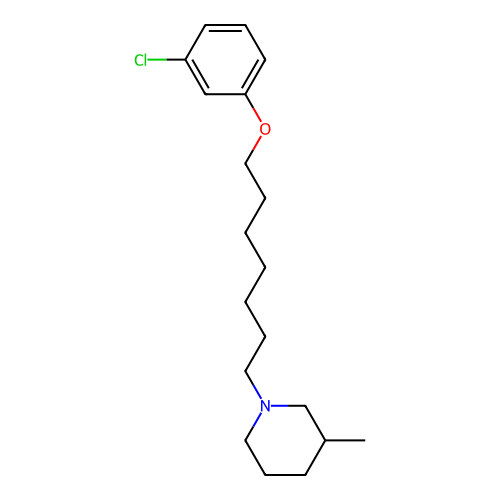
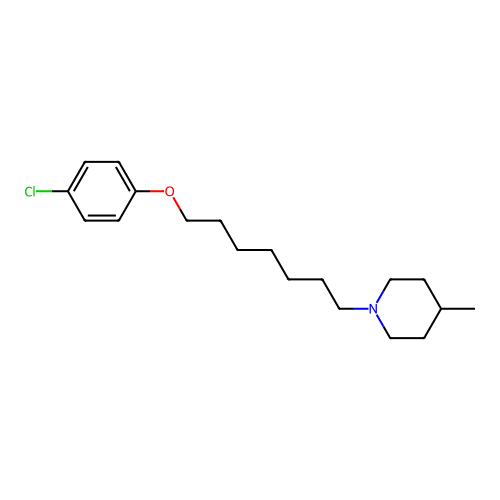
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50046862
Affinity DataEC50: 72nMAssay Description:Antagonist activity at human recombinant histamine H3 receptor expressed in HEK293 cells assessed as inhibition of forskolin/(R)(-)-alpha-methylhista...More data for this Ligand-Target Pair
Affinity DataEC50: 75nMAssay Description:Antagonist activity at human recombinant histamine H3 receptor expressed in HEK293 cells assessed as inhibition of forskolin/(R)(-)-alpha-methylhista...More data for this Ligand-Target Pair
Affinity DataKi: 128nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 133nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 174nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 189nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 203nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 205nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 237nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 255nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 265nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 293nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 333nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 371nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 412nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 427nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 466nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 467nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 800nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 805nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: >1.50E+5nMAssay Description:Displacement of [3H]-Nalpha-methylhistamine from human recombinant histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair