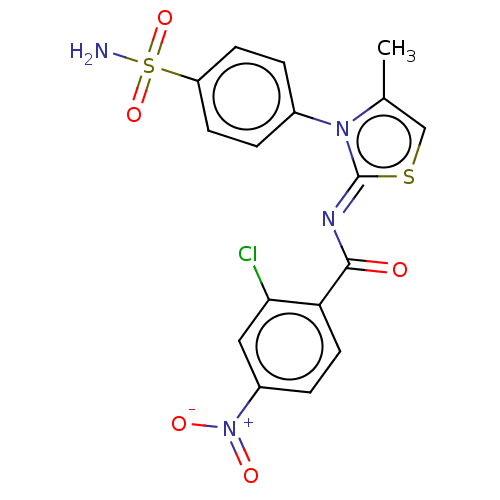
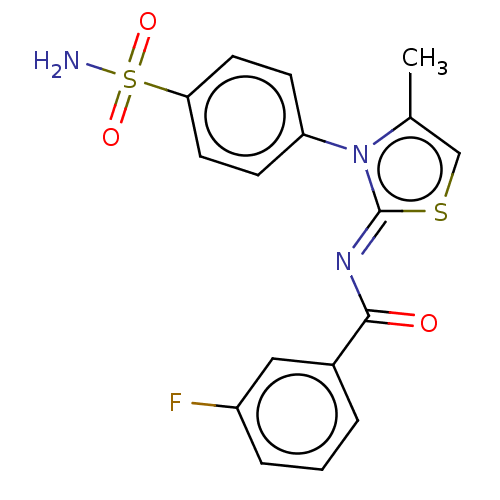
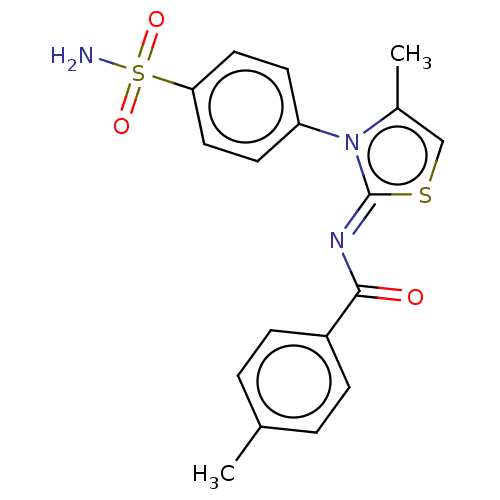
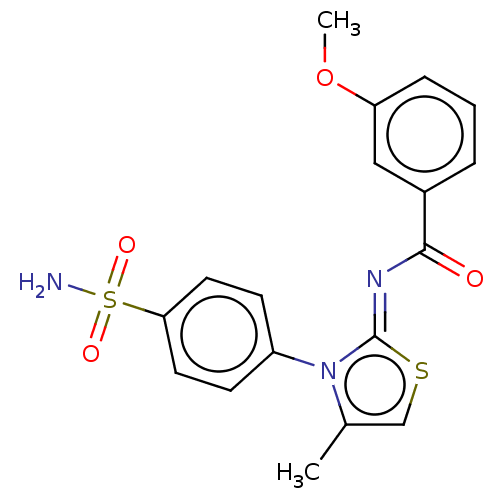
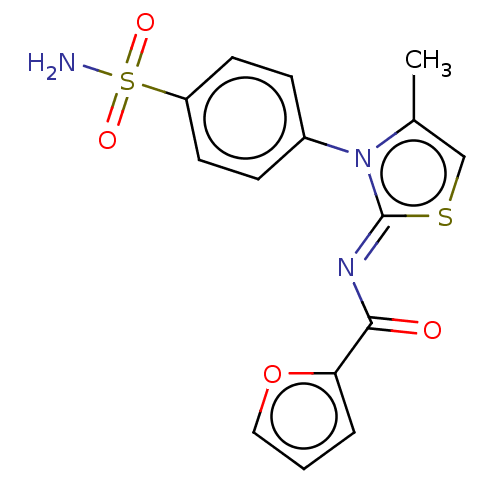
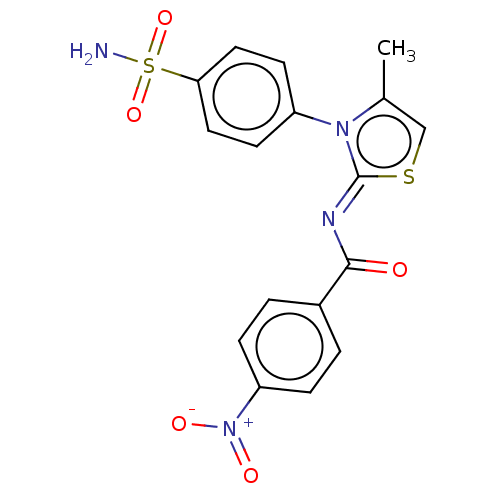
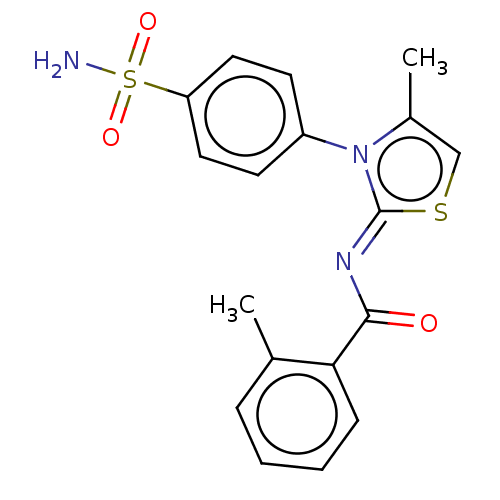
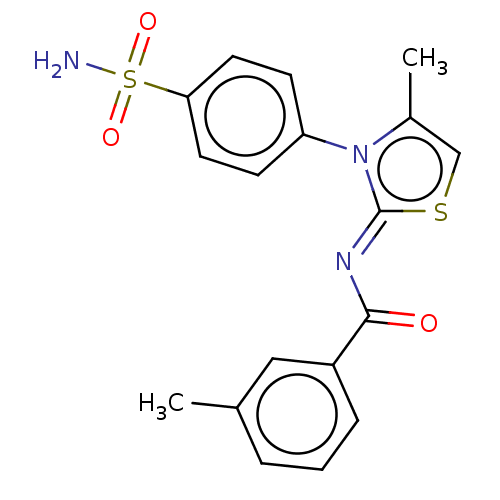
Report error Found 11 Enz. Inhib. hit(s) with all data for entry = 7147
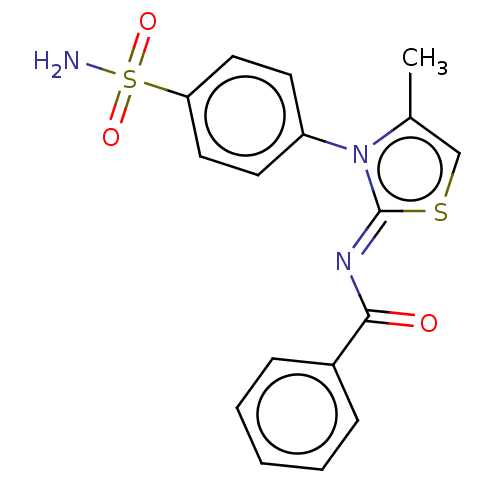
Affinity DataIC50: 58nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
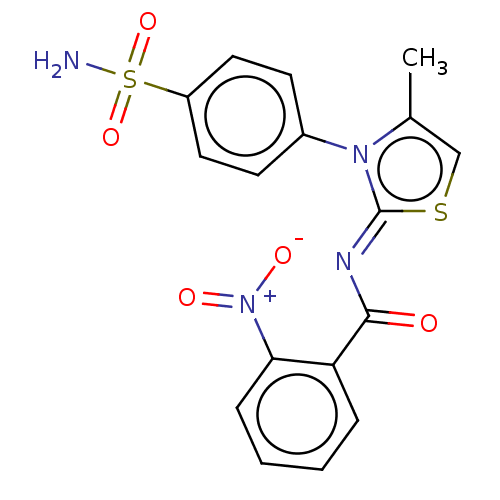
Affinity DataIC50: 64nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 72nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 81nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 96nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair
Affinity DataIC50: 2.09E+4nMpH: 8.2Assay Description:The urease inhibitory activity of synthesized sulfonamides (3a-j) was determined by measuring the amount of ammonia produced by the indophenols metho...More data for this Ligand-Target Pair