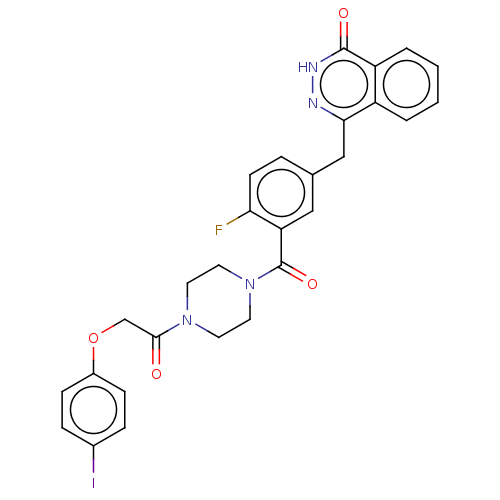
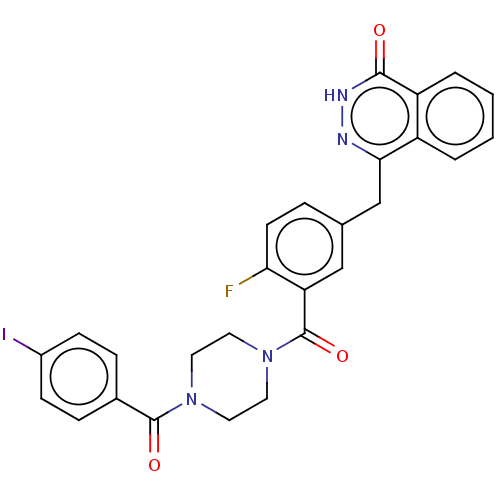
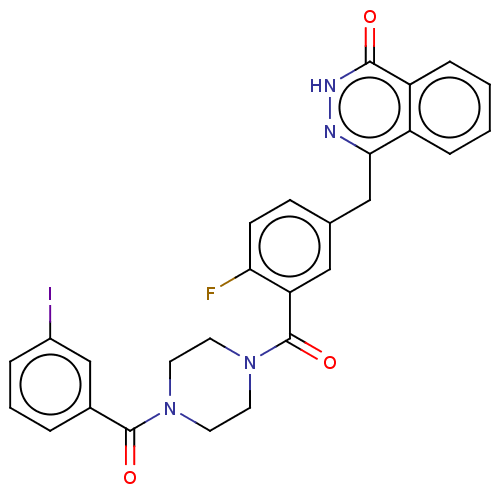
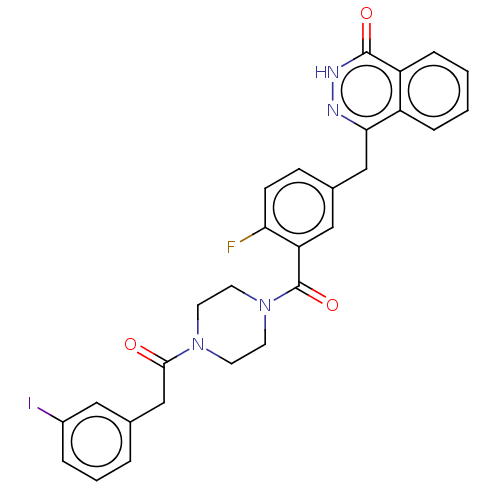
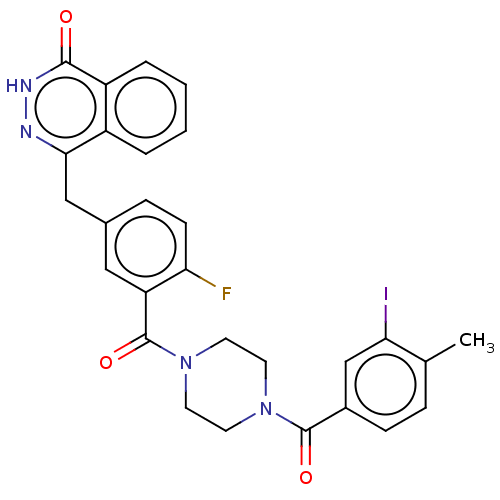
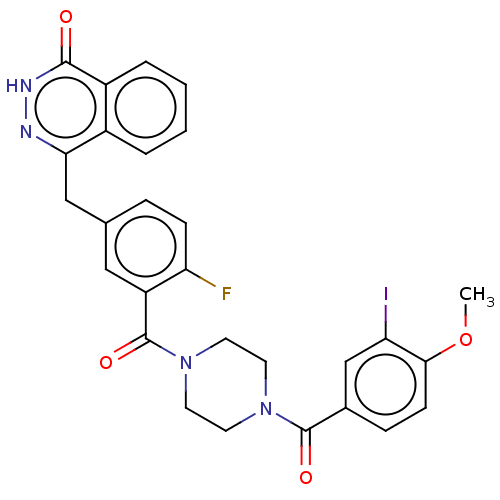
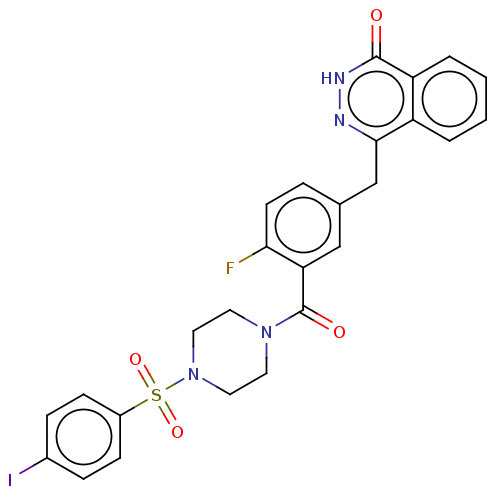
Report error Found 12 Enz. Inhib. hit(s) with all data for entry = 50006349
Affinity DataIC50: 1.60nMAssay Description:Inhibition of PARP1 in human T98G cells incubated for 60 mins by immunofluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of PARP1 in human G7 cells incubated for 60 mins by immunofluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.80nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of PARP1 in human G7 cells incubated for 60 mins by immunofluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.40nMAssay Description:Inhibition of PARP1 in human T98G cells incubated for 60 mins by immunofluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of PARP1 (unknown origin) incubated for 10 mins using biotinylated NAD+ and activated DNA by colorimetric assayMore data for this Ligand-Target Pair
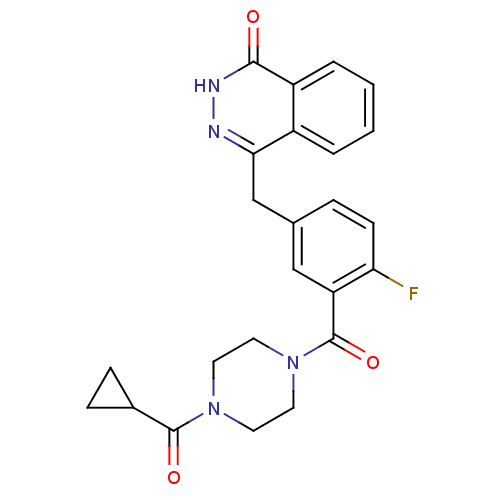
 3D Structure (crystal)
3D Structure (crystal)