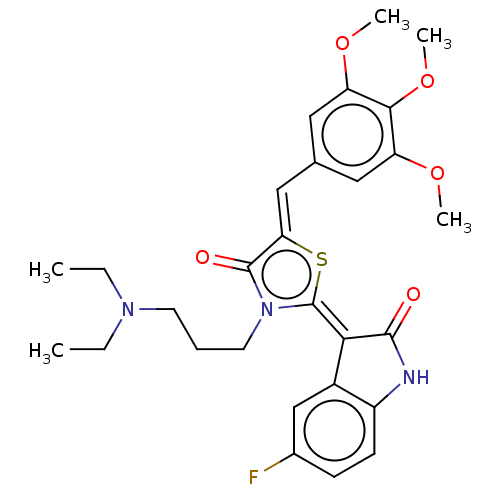
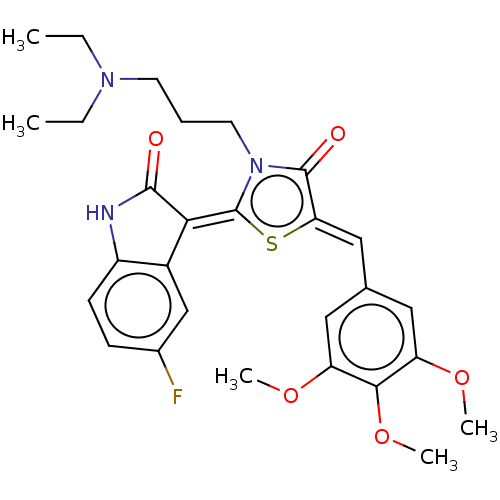
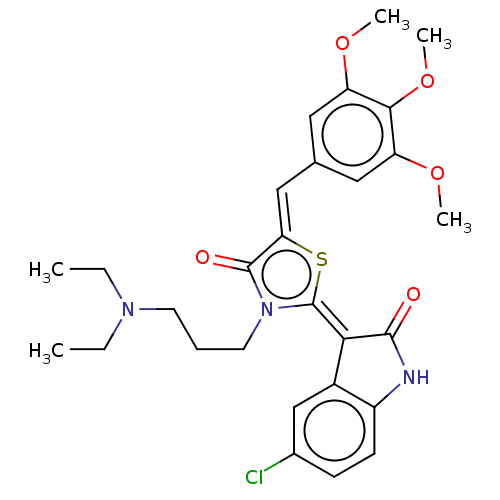
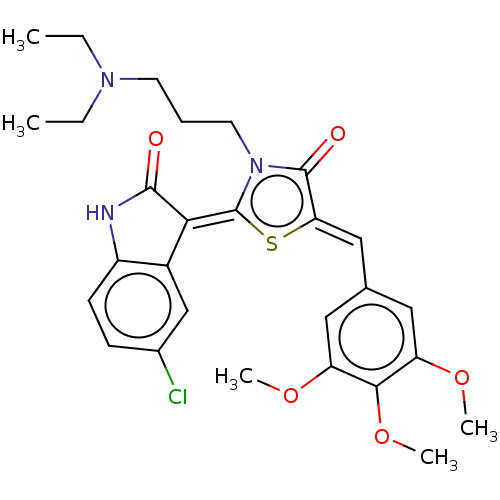
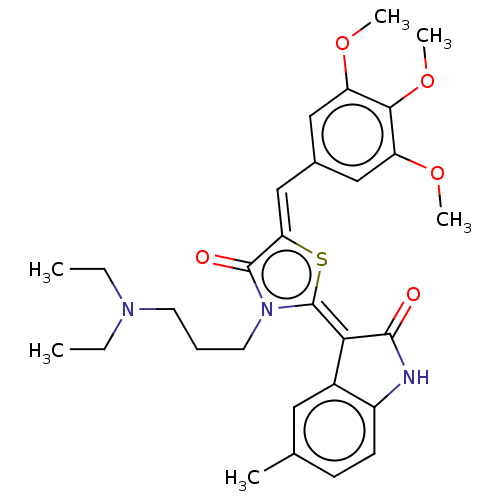
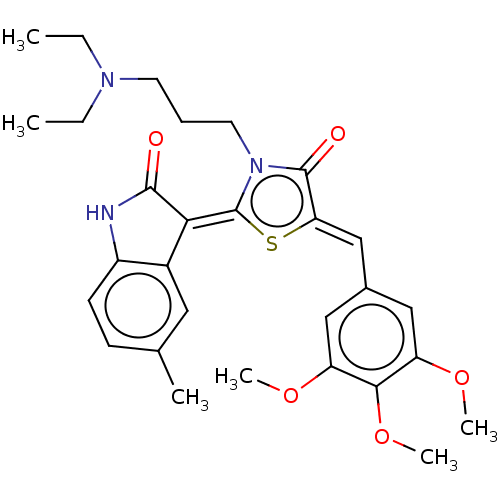
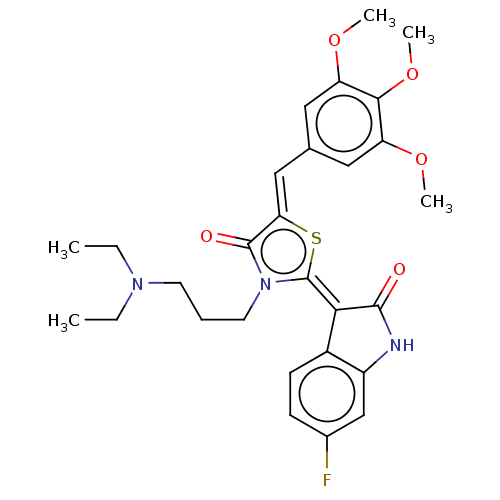
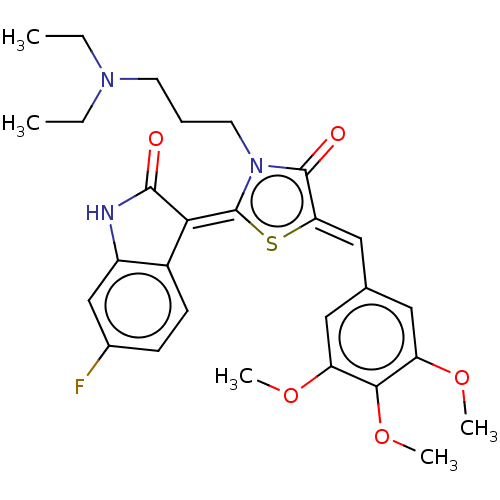
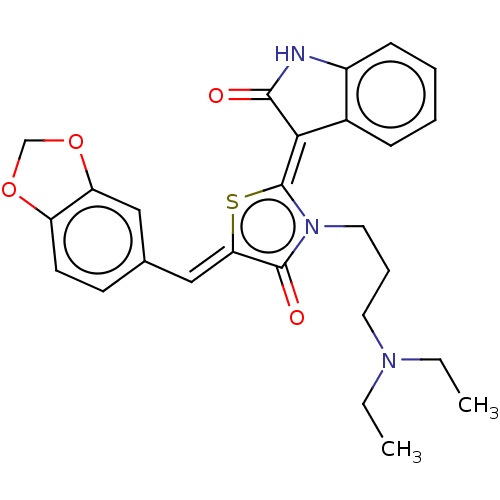
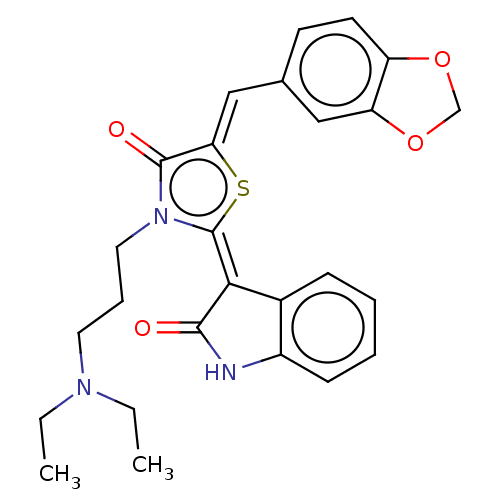
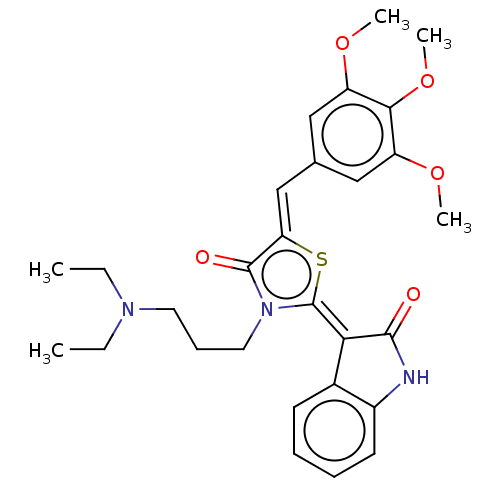
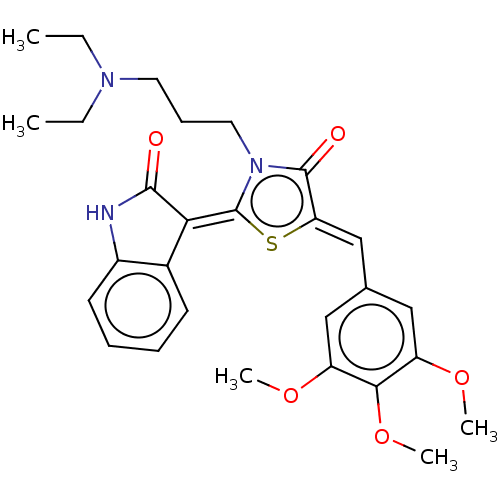
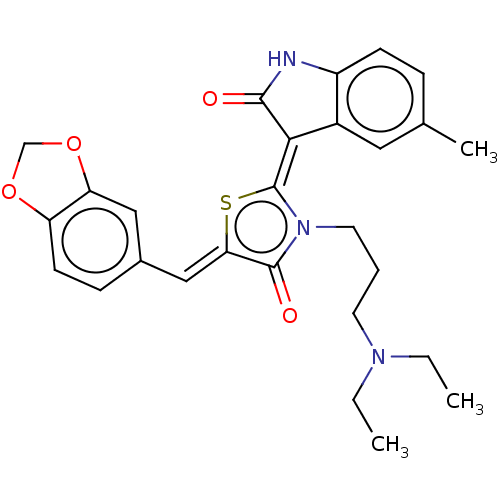
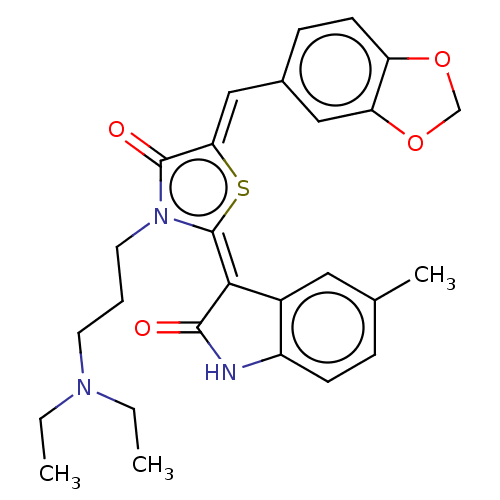
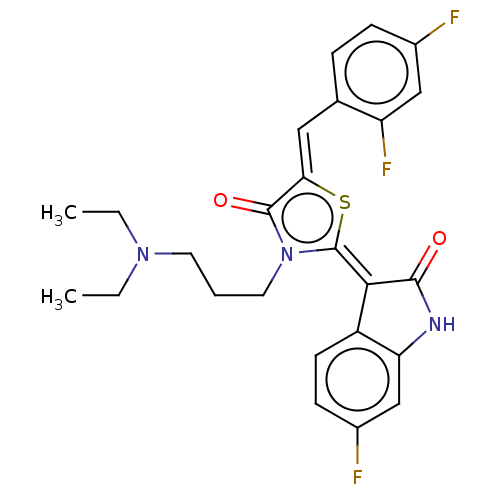
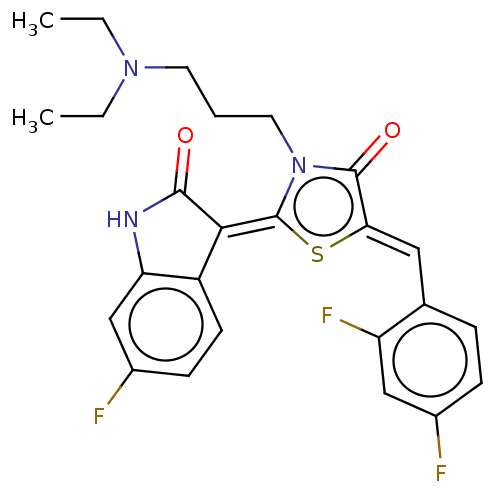
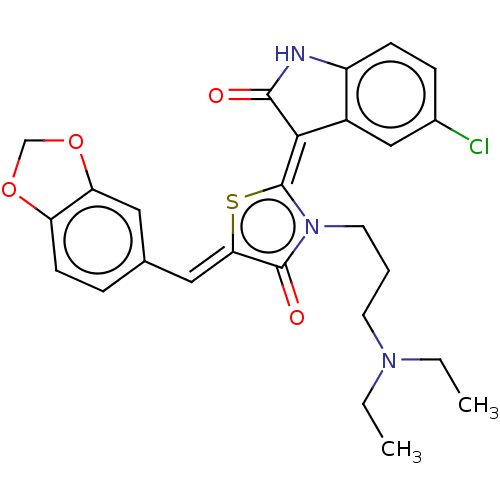
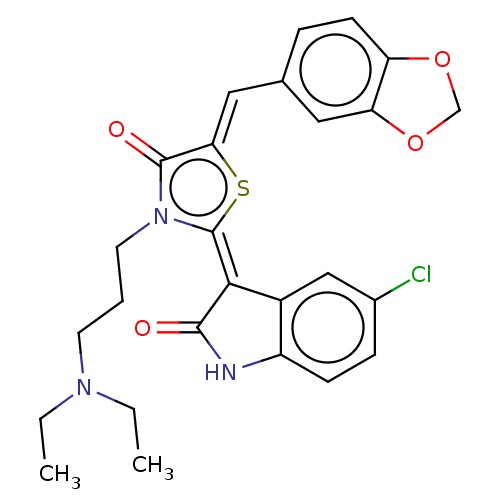
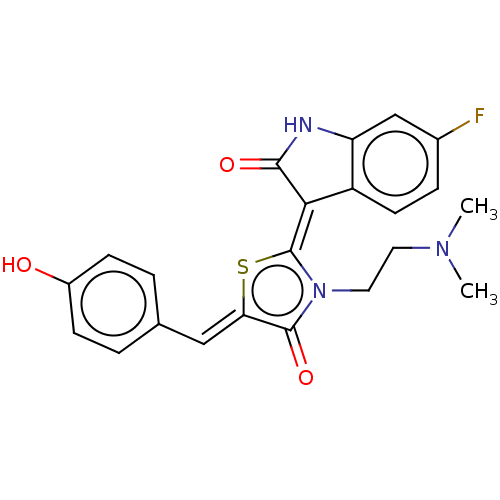
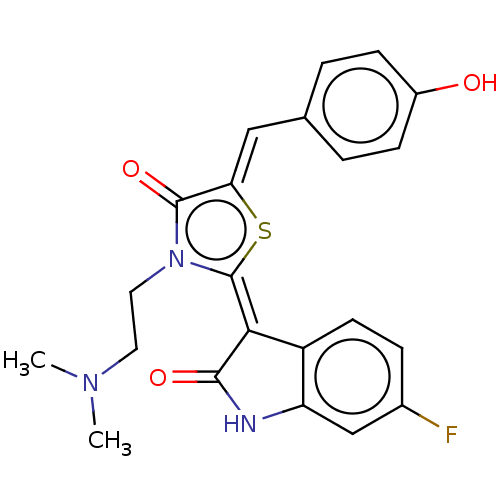
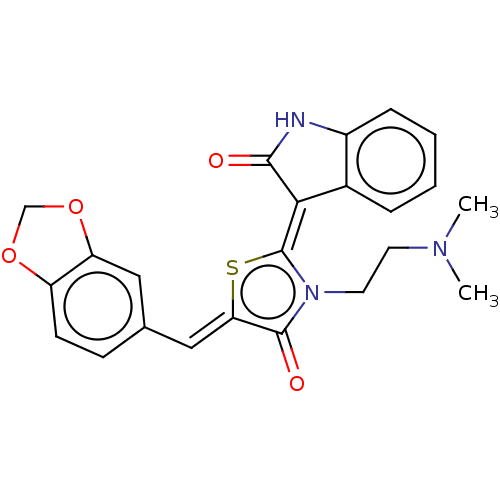
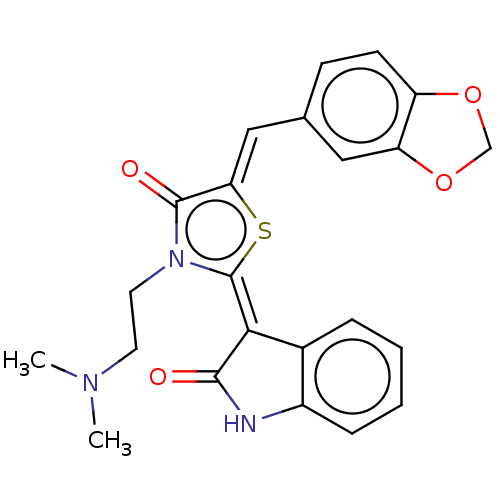
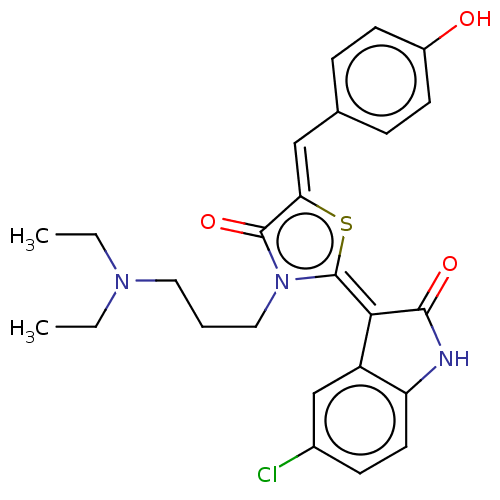
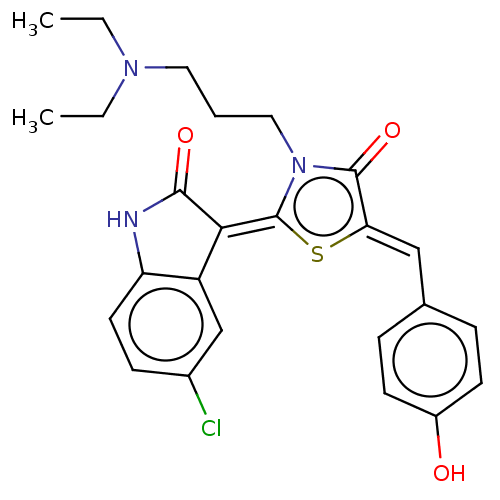
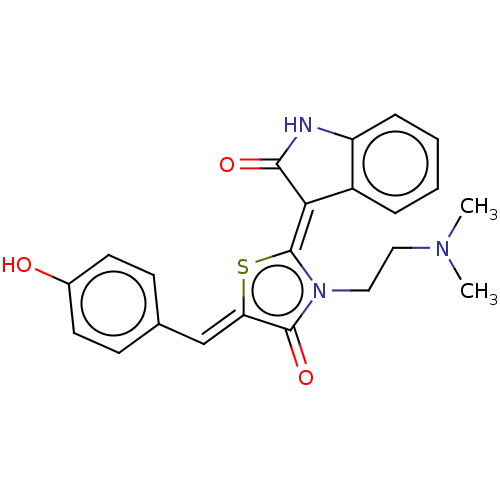
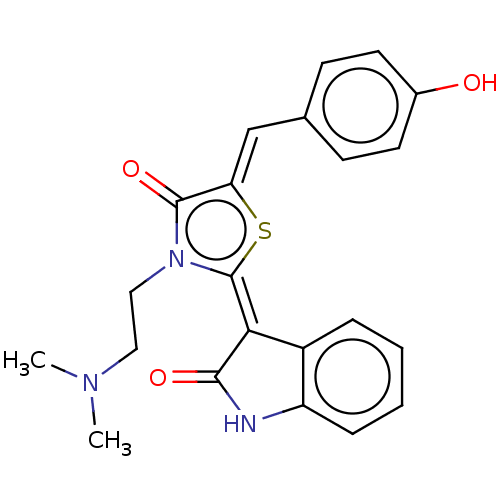
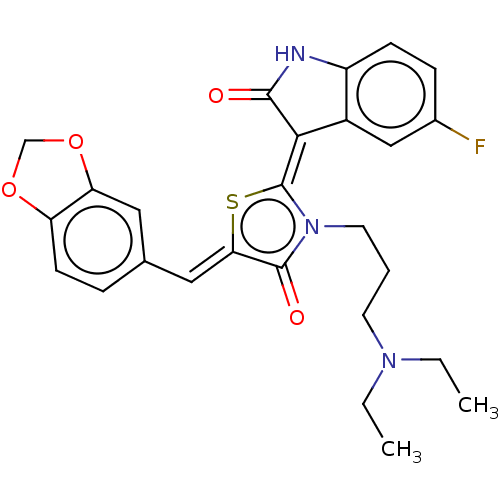
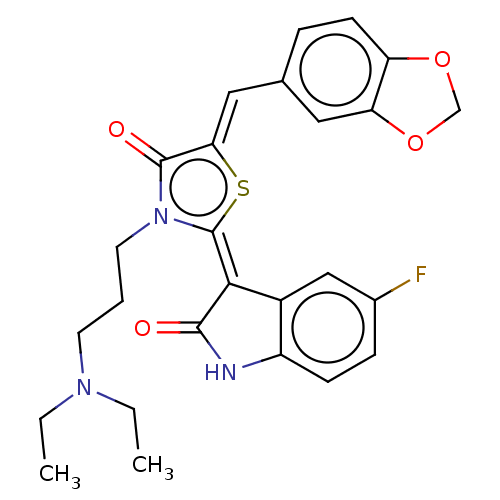
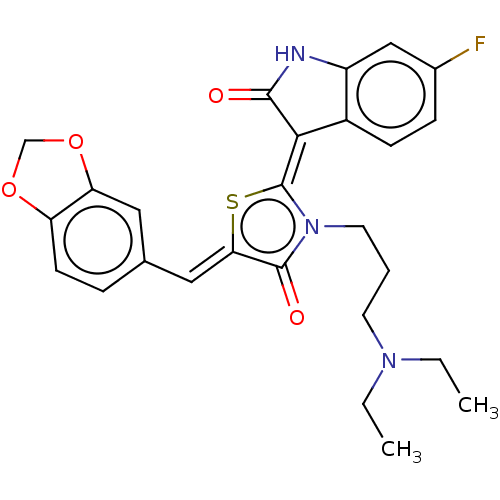
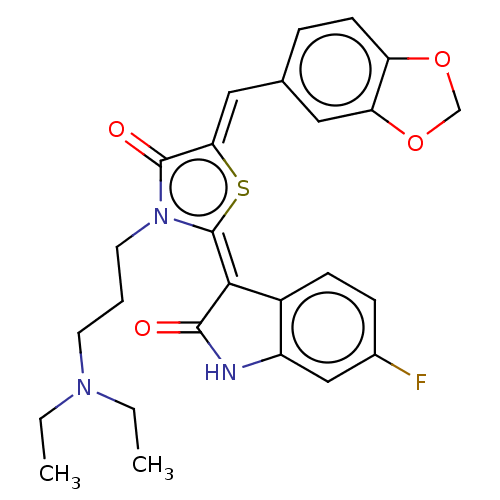
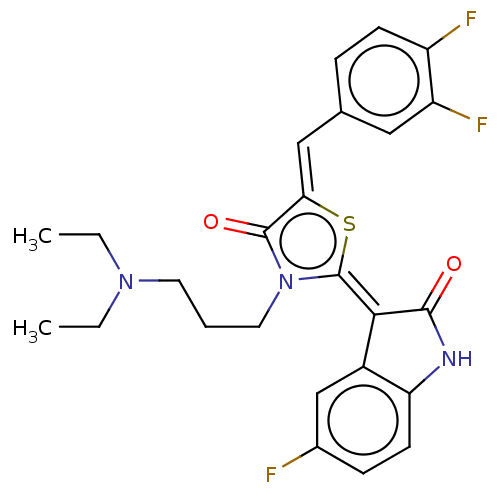
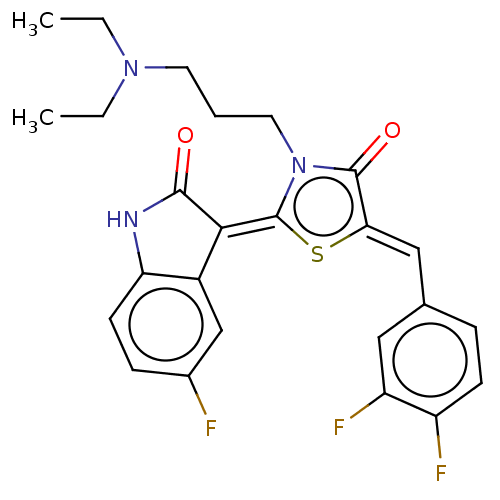
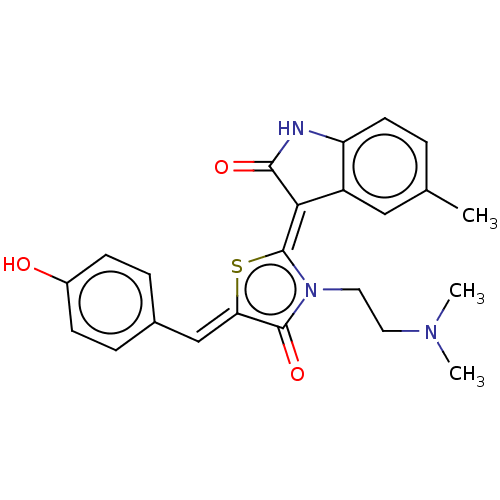
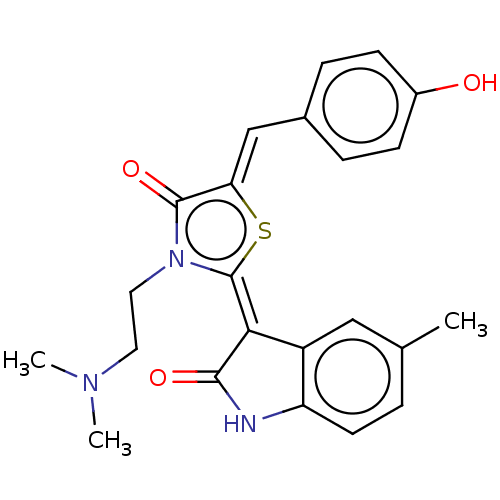
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 50046498
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
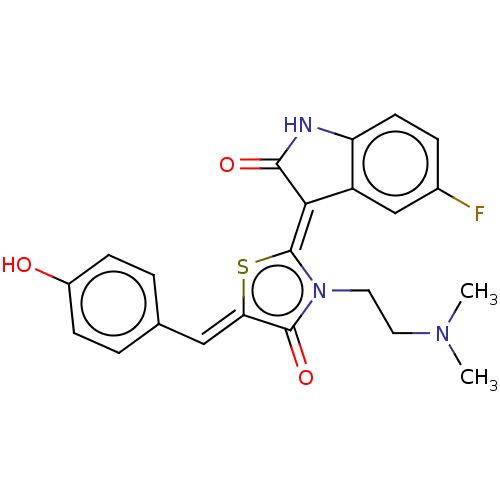
Affinity DataKi: 120nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
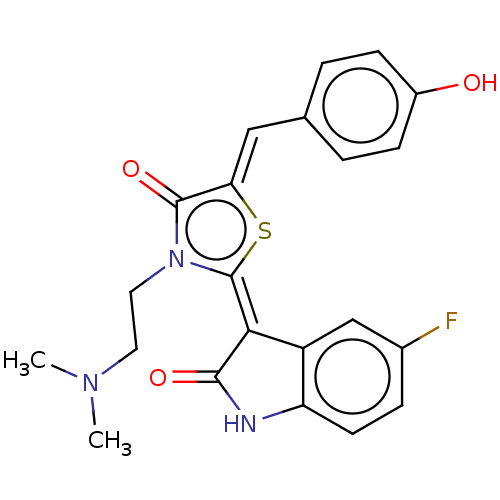
Affinity DataKi: 120nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
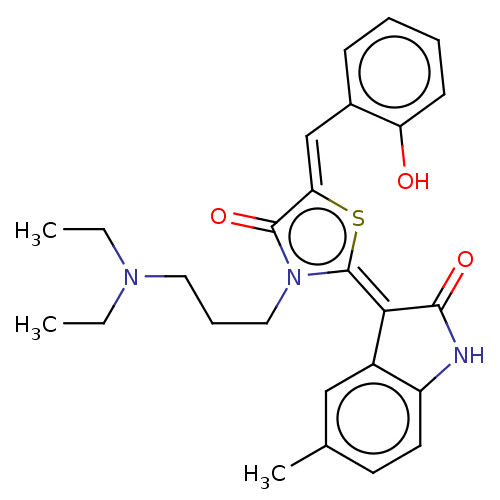
Affinity DataKi: 220nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
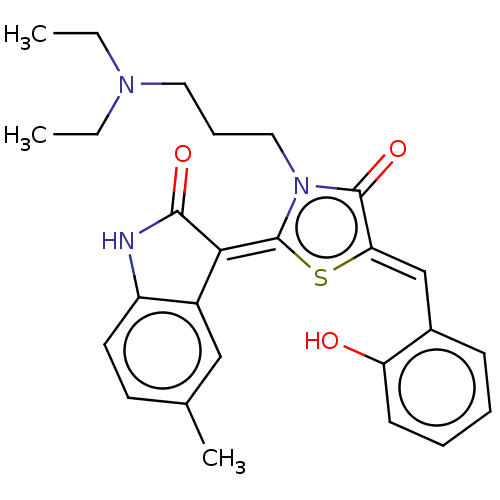
Affinity DataKi: 220nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 320nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 320nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 470nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 470nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 480nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 480nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 730nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 730nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 940nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 940nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.01E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.01E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.34E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.34E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.71E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.71E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.78E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: 1.78E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair
TargetATP-dependent translocase ABCB1(Human)
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
The First Affiliated Hospital of Dalian Medical University
Curated by ChEMBL
Affinity DataKi: >2.23E+3nMAssay Description:Inhibition of P-glycoprotein in human drug-resistant K562/ADR cells assessed as reduction in P-gp mediated rhodamine 123 efflux by spectrofluorometryMore data for this Ligand-Target Pair