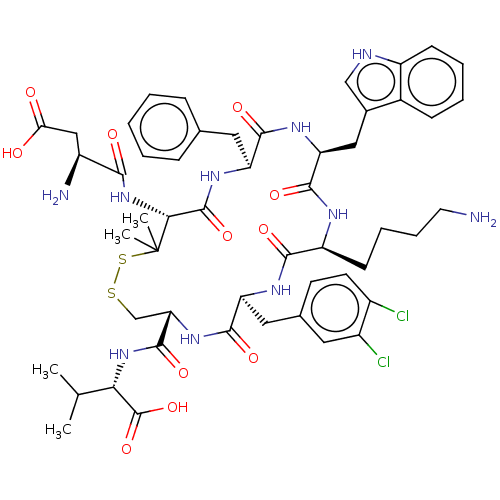
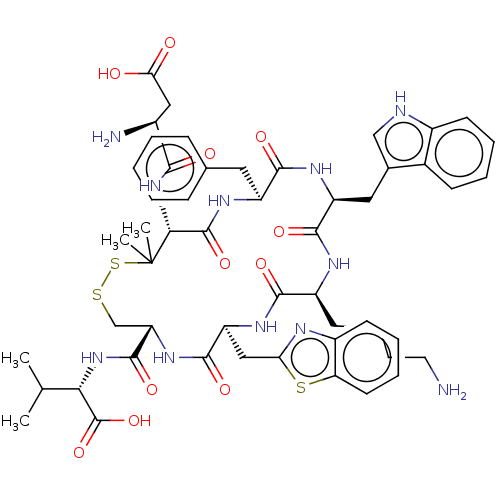
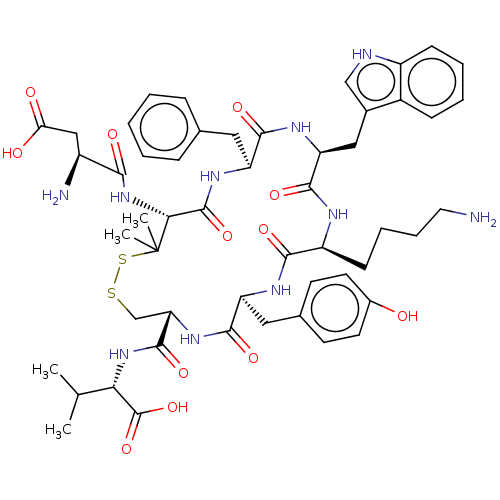
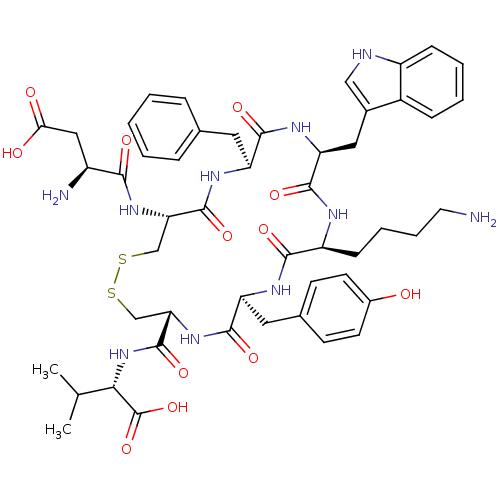
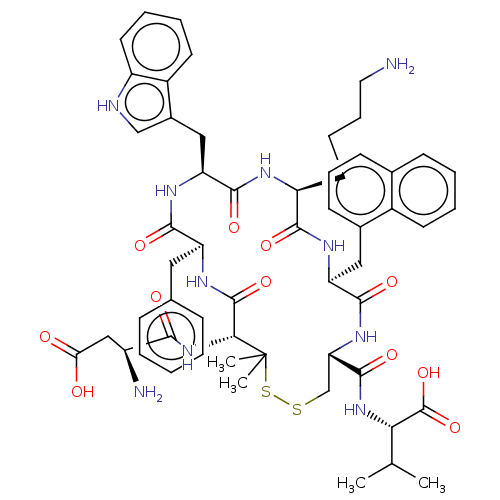
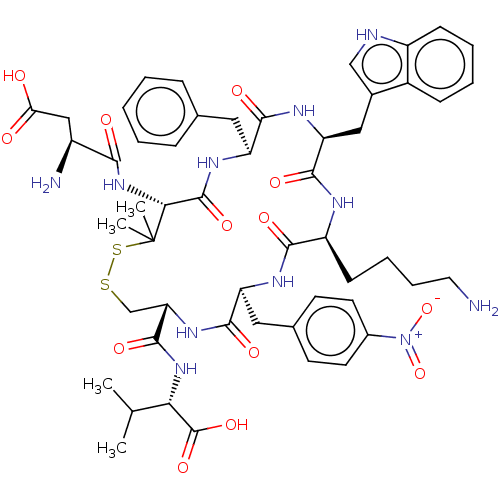
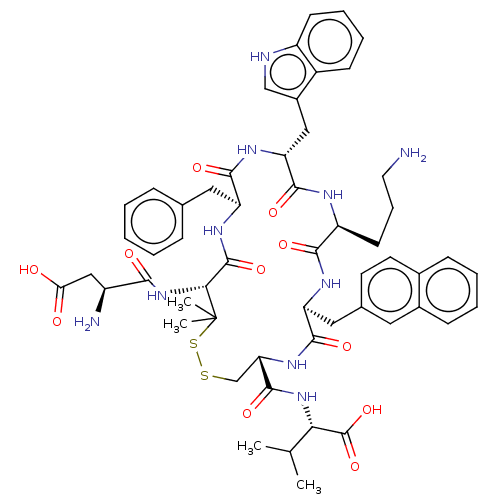
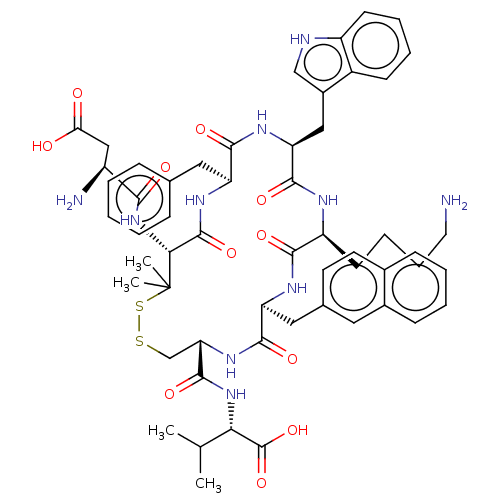
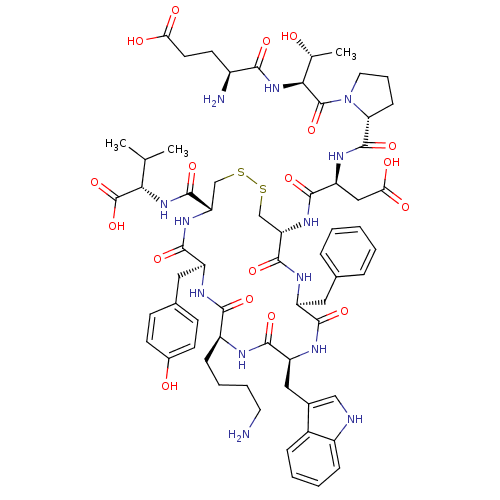
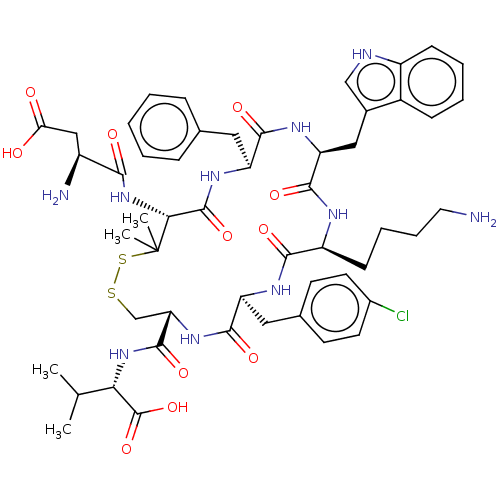
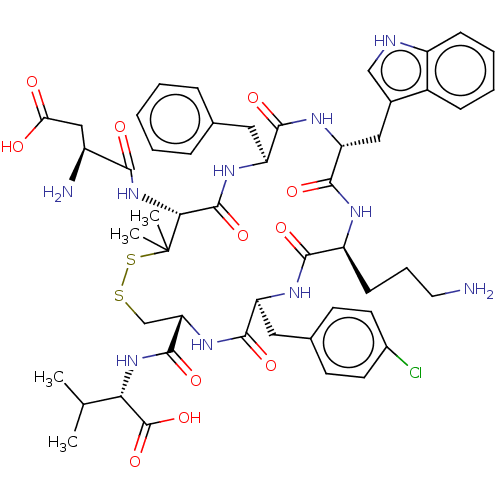
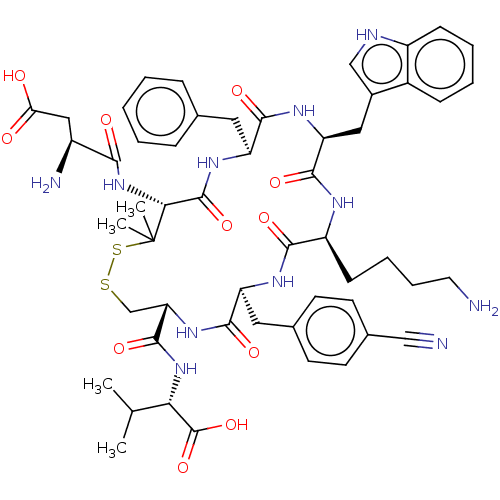
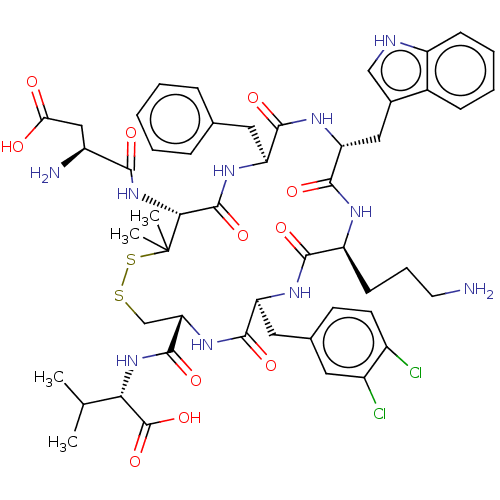
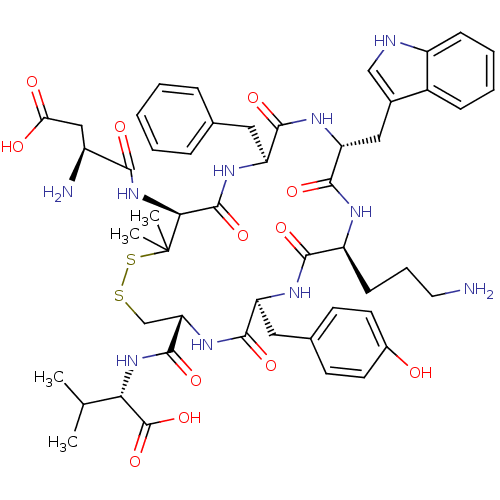
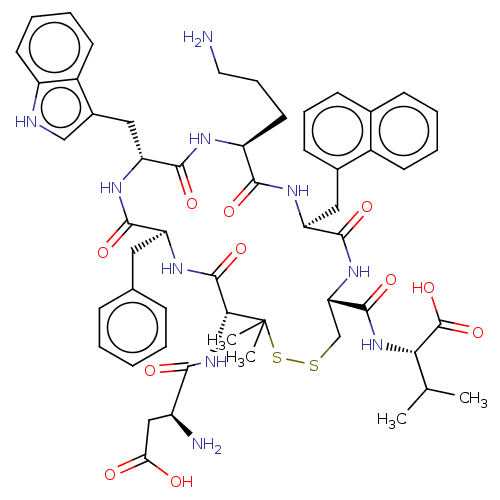
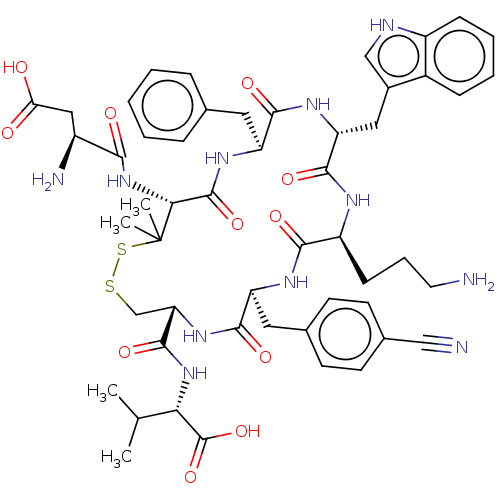
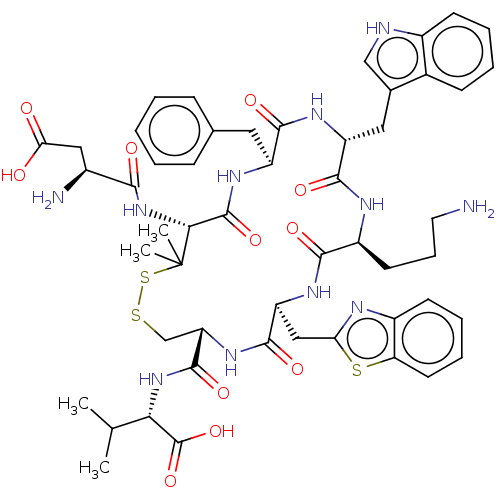
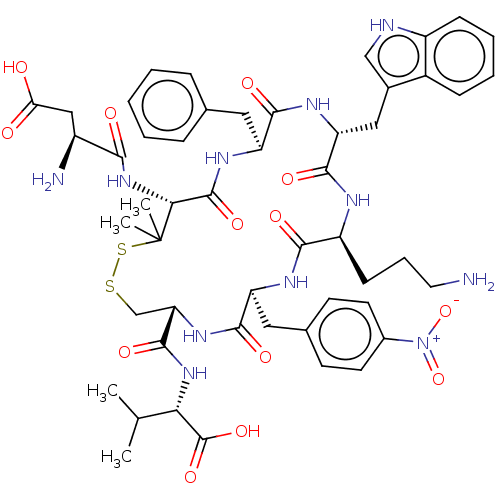
Report error Found 33 Enz. Inhib. hit(s) with all data for entry = 50045185
Affinity DataEC50: 0.0126nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataEC50: 0.0195nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 0.251nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 0.398nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataEC50: 0.398nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataEC50: 0.631nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataKi: 0.724nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataEC50: 0.724nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataKi: 0.776nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 0.794nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataEC50: 0.813nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.10nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataEC50: 2.70nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataEC50: 3.20nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataEC50: 3.60nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataEC50: 3.80nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataEC50: 5nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 6.5nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataEC50: 13nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Displacement of [125I]urotensin-2 from human recombinant UT receptor expressed in CHO-K1 cells by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataEC50: 18nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair
Affinity DataEC50: 34nMAssay Description:Agonist activity at UT receptor in rat aorta assessed as contractionMore data for this Ligand-Target Pair