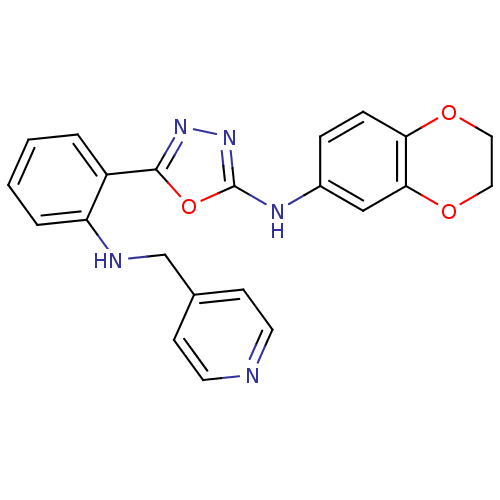
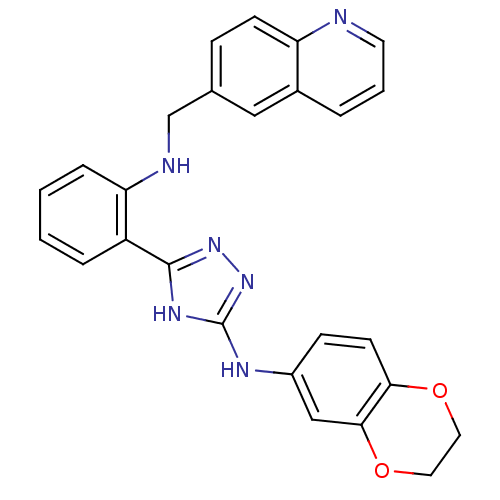
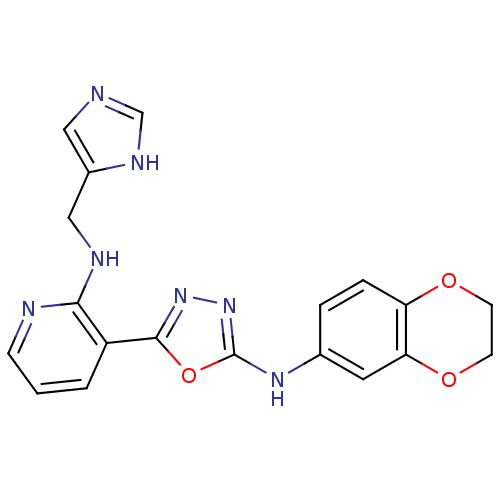
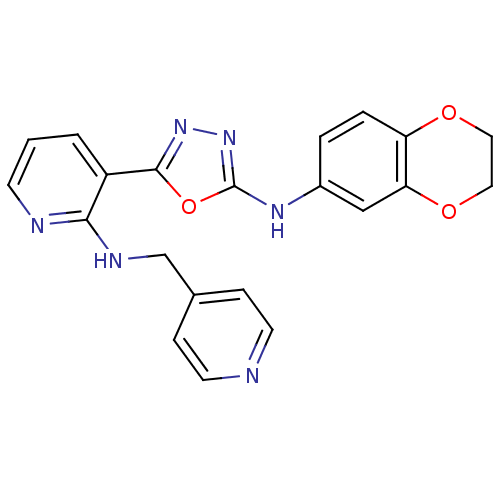
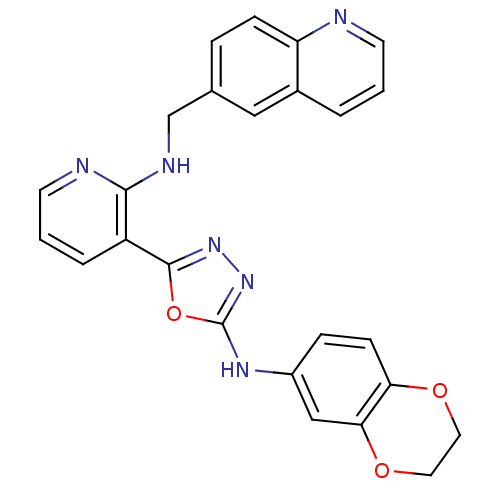
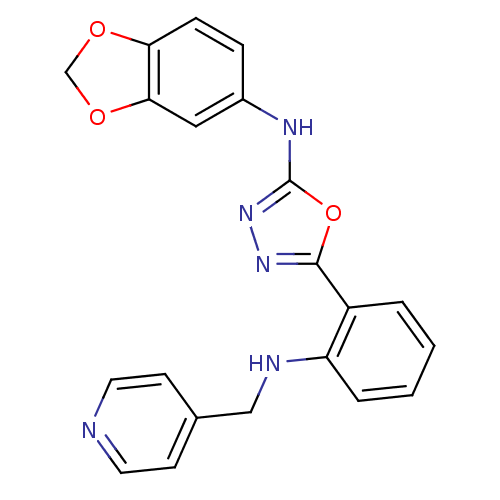
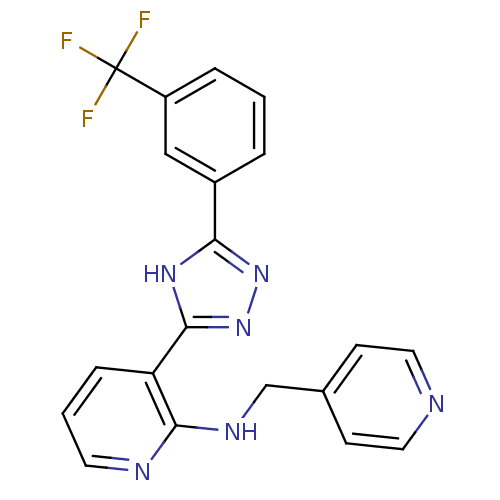
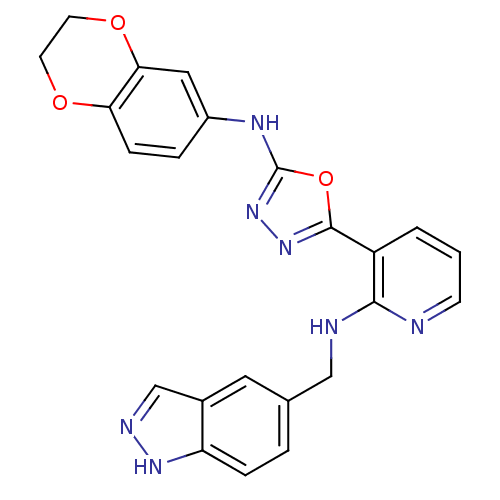
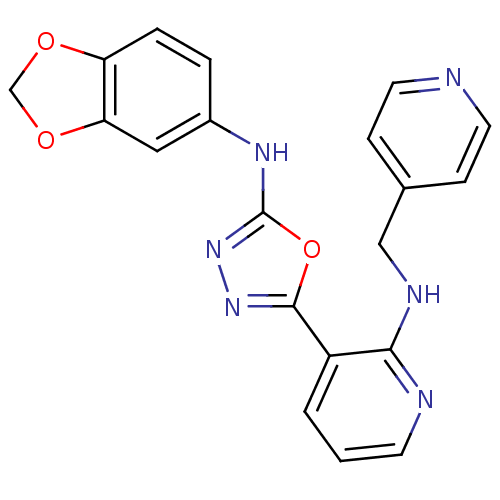
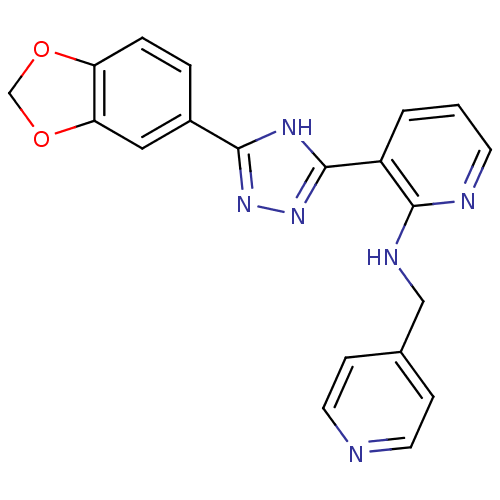
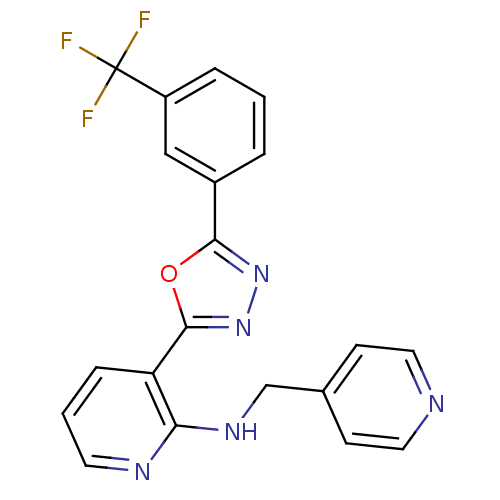
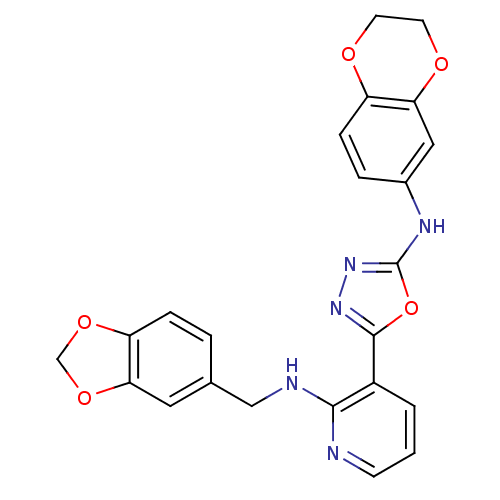
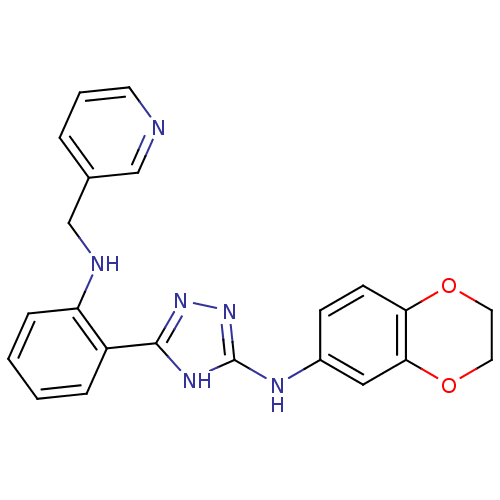
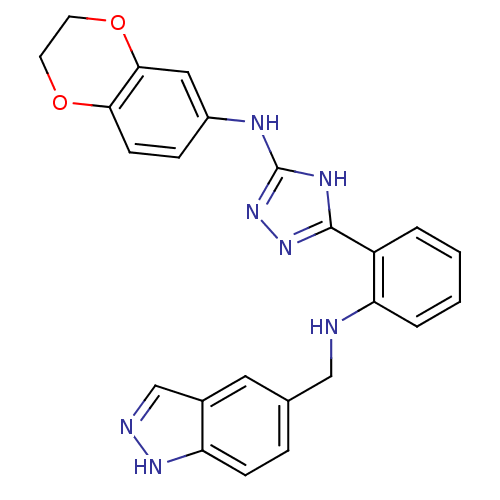
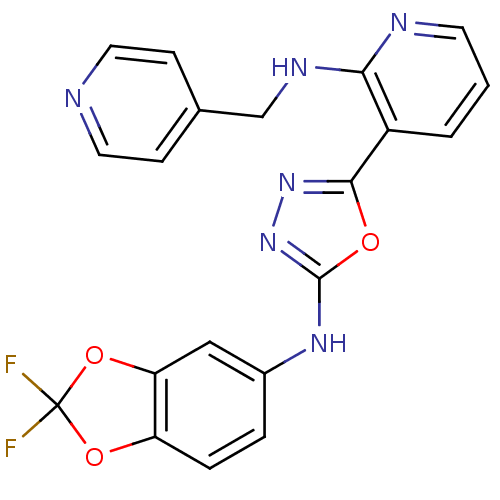
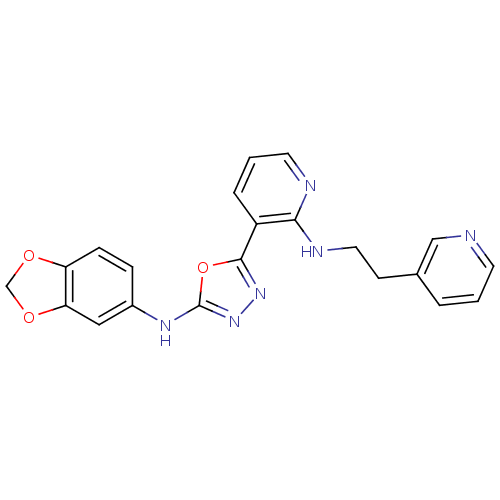
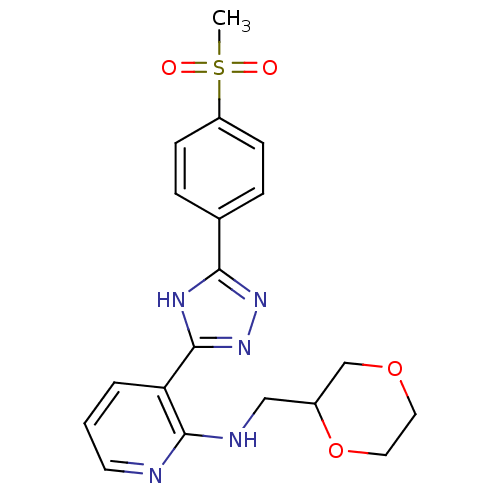
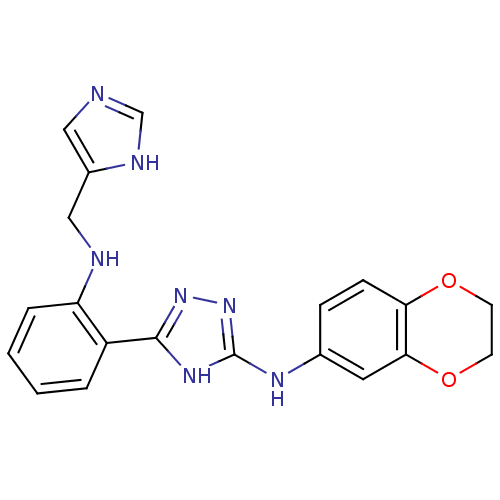
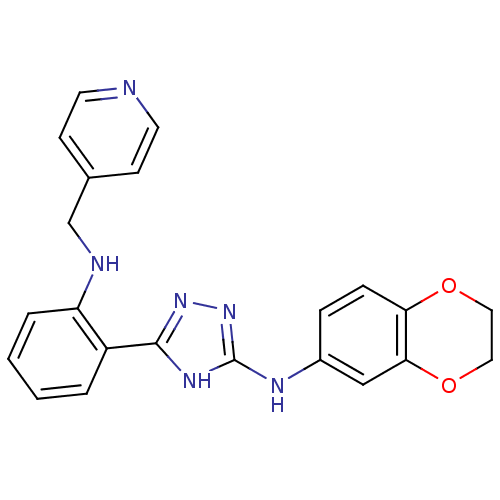
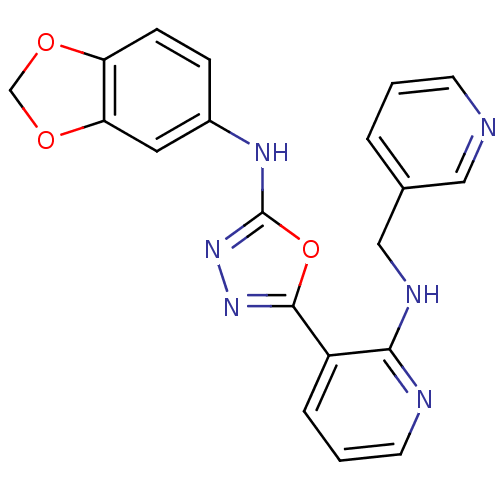
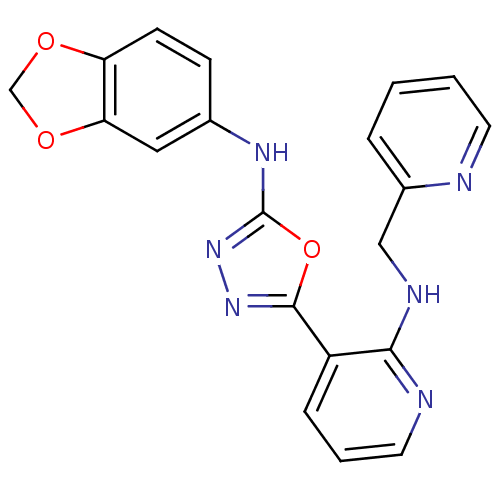
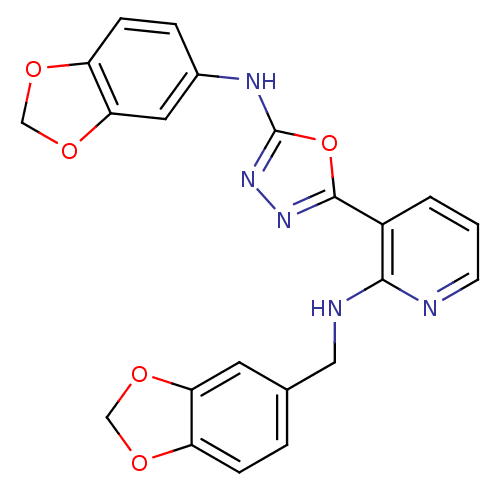
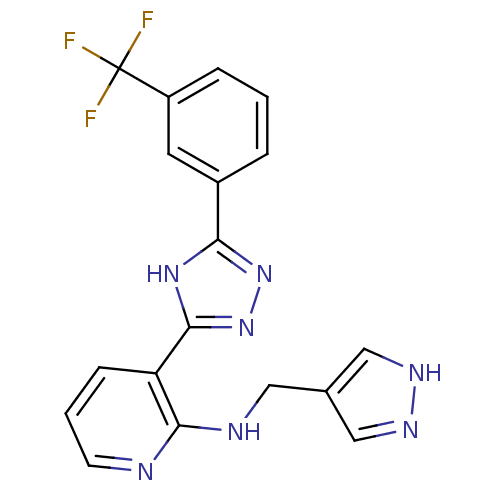
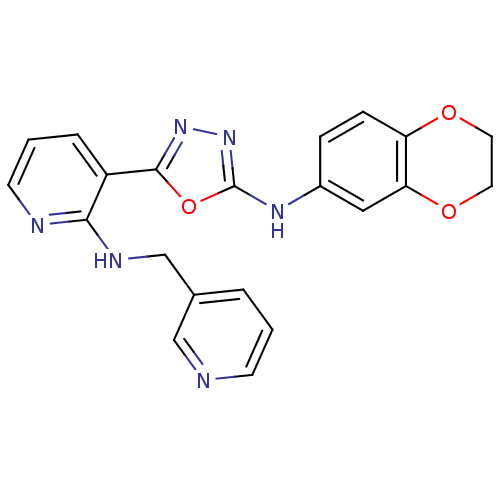
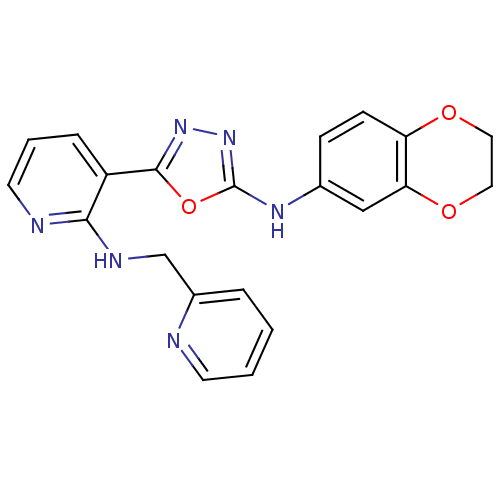
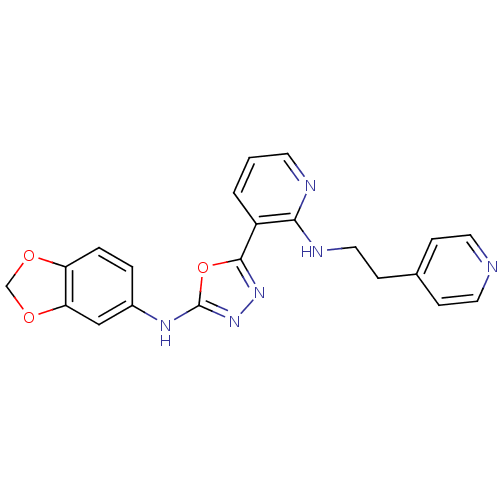
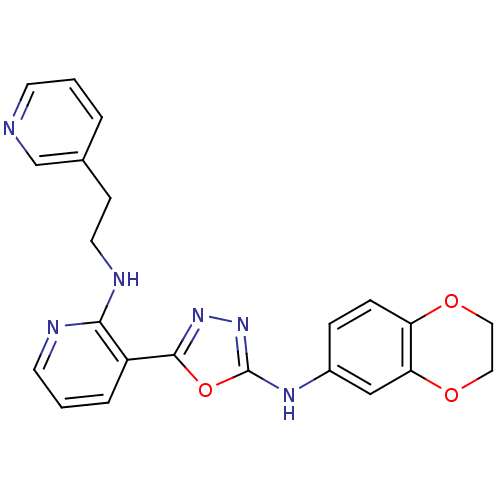
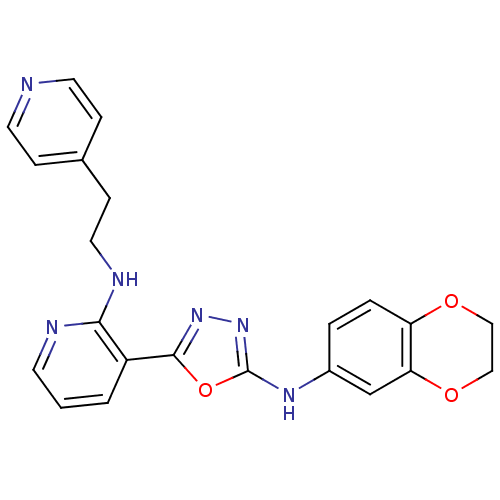
Report error Found 56 Enz. Inhib. hit(s) with all data for entry = 50033065
Affinity DataIC50: 110nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 830nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 830nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.89E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.16E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 3.41E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+3nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30E+3nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR-2 kinase assessed as phosphorylated level of pGAT-biotin peptide preincubated for 5 to 10 mins before addition of substrate measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of VEGFR2 expressed HEK293 cellsMore data for this Ligand-Target Pair