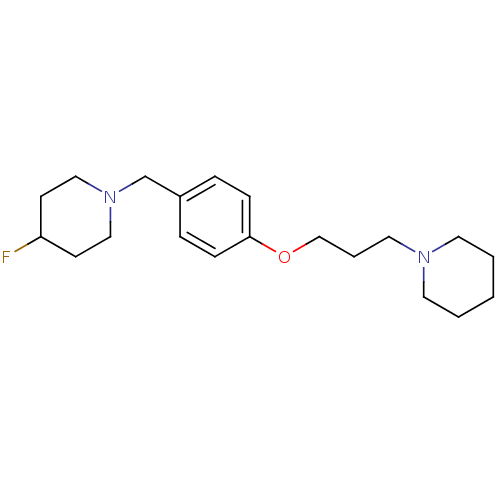
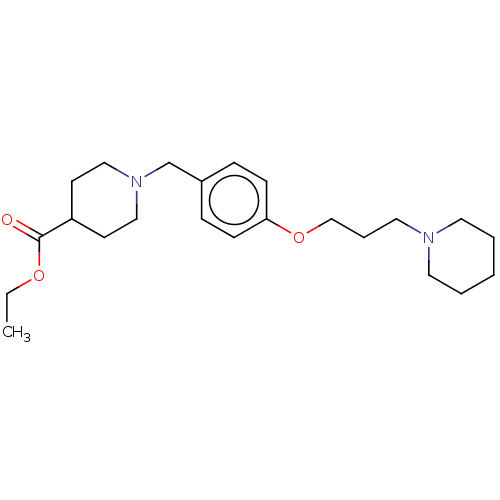
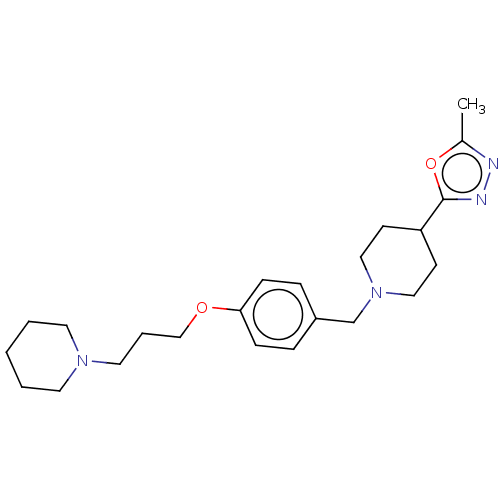
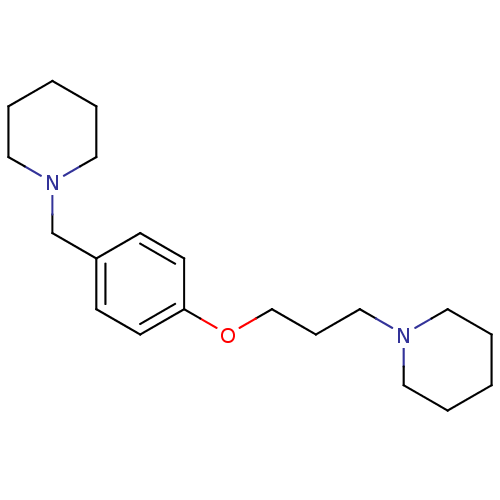
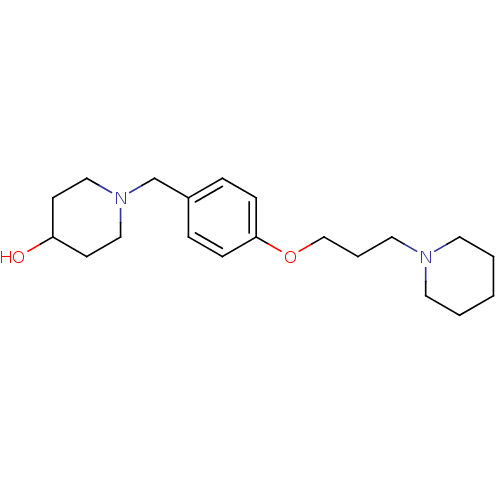
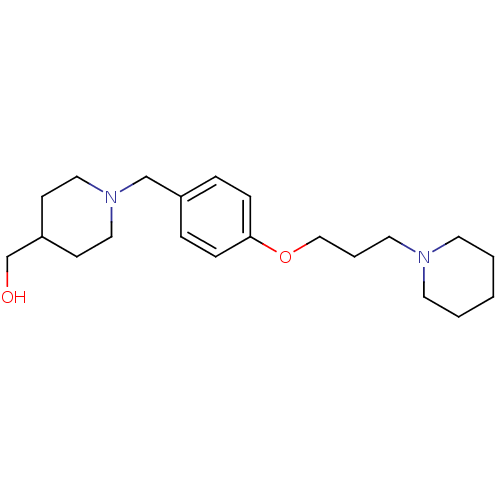
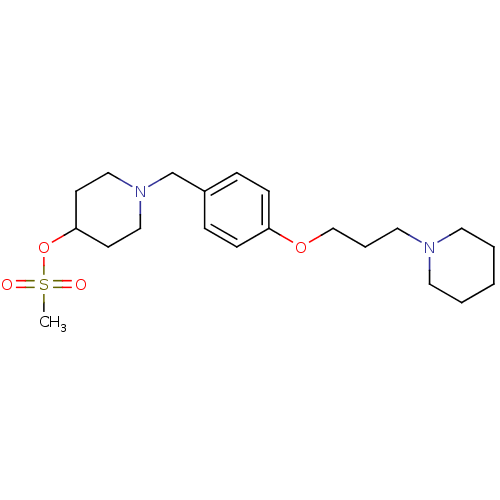
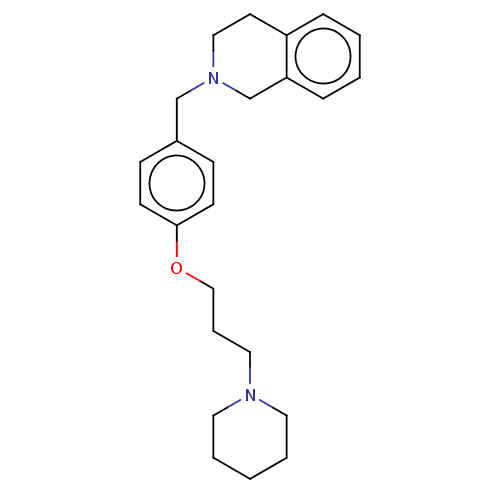
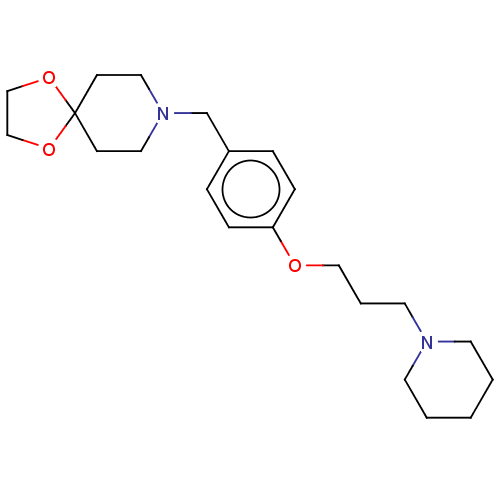
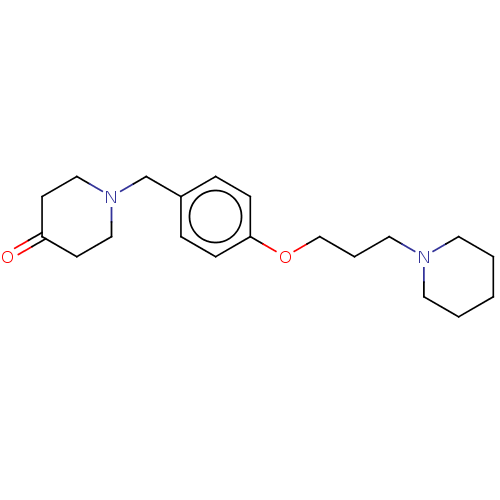
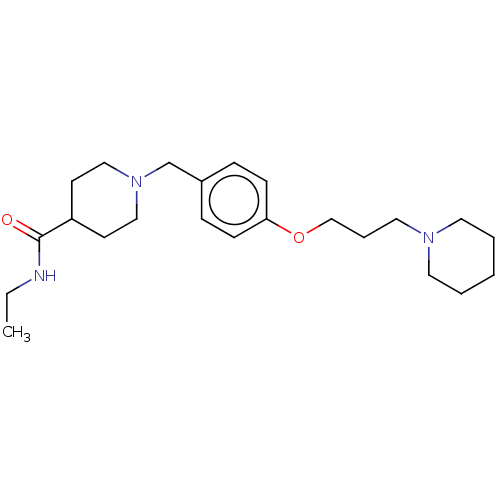
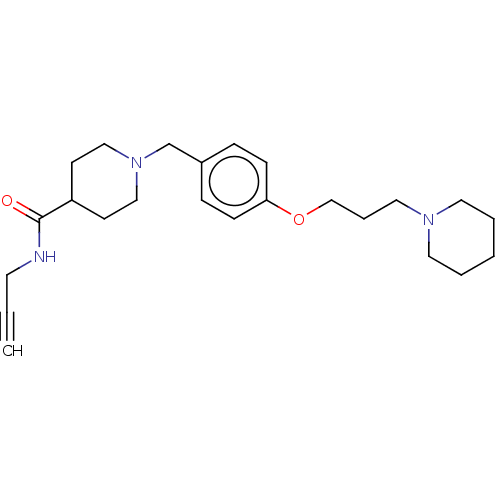
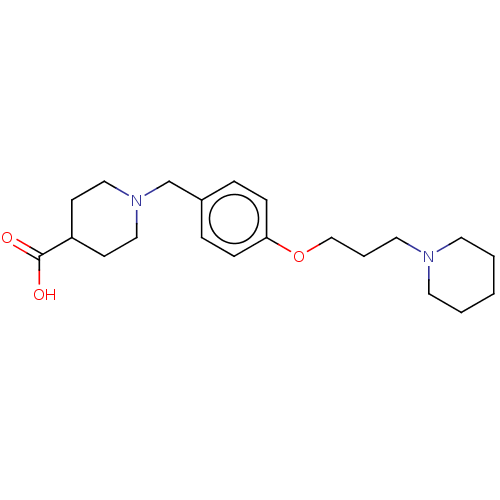
Report error Found 19 Enz. Inhib. hit(s) with all data for entry = 50044347
Affinity DataKi: 0.0933nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0940nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.229nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.479nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.580nMAssay Description:Binding affinity to rat histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.589nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.631nMAssay Description:Binding affinity to rat histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.676nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.741nMAssay Description:Binding affinity to human histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.832nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.840nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity to human histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Binding affinity to human histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [125I]Iodoproxyfan from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Displacement of [125I]Iodoproxyfan from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [125I]Iodoproxyfan from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [125I]Iodoproxyfan from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [125I]Iodoproxyfan from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from recombinant human histamine H3 receptor expressed in HEK293 cells after 90 minsMore data for this Ligand-Target Pair