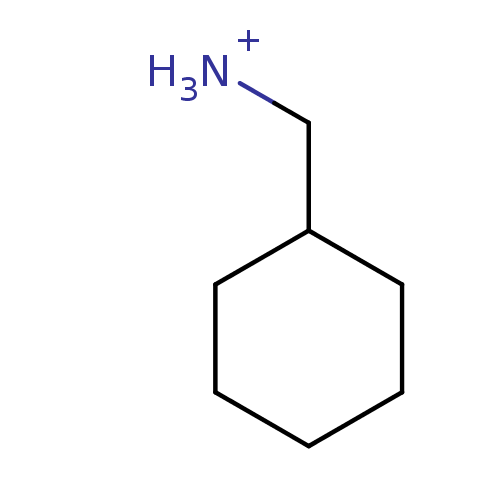 Found 23 hits with all data for this entryid
Found 23 hits with all data for this entryid
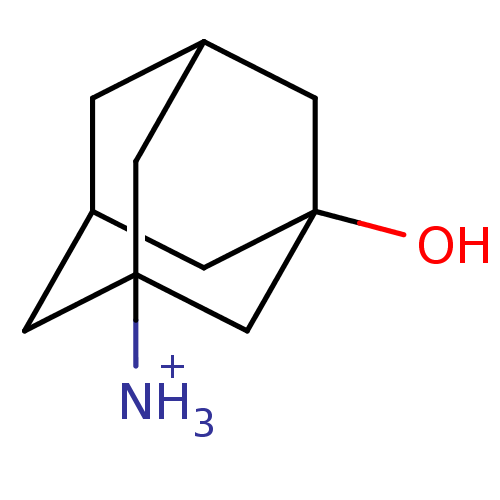
ITC DataΔG°: -59kJ/mole
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
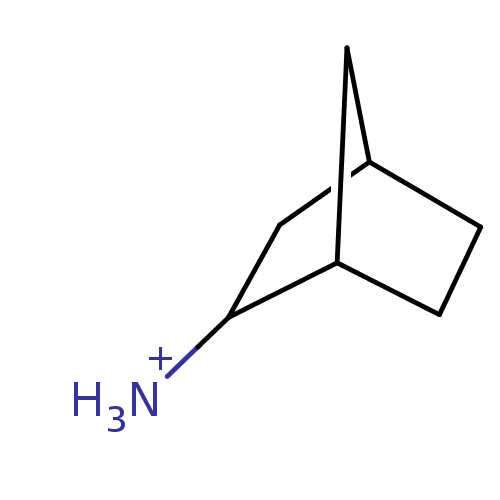
ITC DataΔG°: -55.7kJ/mole
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -52.8kJ/mole logk: 1.80E+8
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
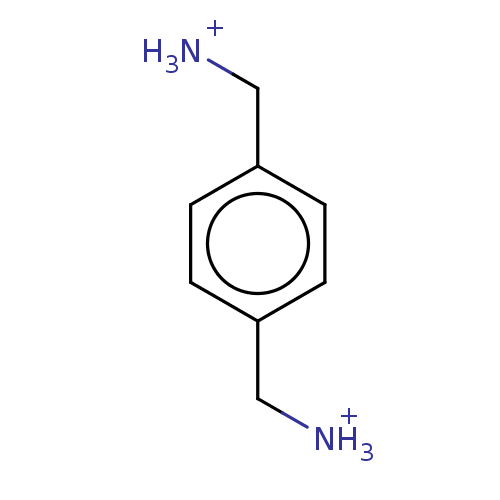
ITC DataΔG°: -49.4kJ/mole logk: 4.60E+8
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -48.6kJ/mole logk: 3.10E+8
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -46.5kJ/mole logk: 1.30E+8
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -42.3kJ/mole logk: 2.40E+7
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -41kJ/mole logk: 1.70E+7
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -40kJ/mole logk: 1.10E+7
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -36kJ/mole logk: 1.80E+6
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -35kJ/mole logk: 1.50E+6
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -33kJ/mole logk: 5.80E+5
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -33kJ/mole logk: 6.50E+5
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -32kJ/mole logk: 3.80E+5
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -28.1kJ/mole logk: 8.50E+4
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -28kJ/mole logk: 7.10E+4
pH: 7.4 T: 24.85°C
pH: 7.4 T: 24.85°C
ITC DataΔG°: -27.7kJ/mole logk: 7.00E+4
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -26.3kJ/mole logk: 4.00E+4
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -25kJ/mole logk: 2.00E+4
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -23kJ/mole logk: 1.30E+4
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -22kJ/mole logk: 7.10E+3
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -15.6kJ/mole logk: 540
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
ITC DataΔG°: -15.6kJ/mole logk: 540
pH: 9.2 T: 24.85°C
pH: 9.2 T: 24.85°C
- Reorder this table by ΔH°
- Reorder this table by ΔS°
- Reorder this table by pH
- Reorder this table by T
- Change energy unit to: kcal/mol