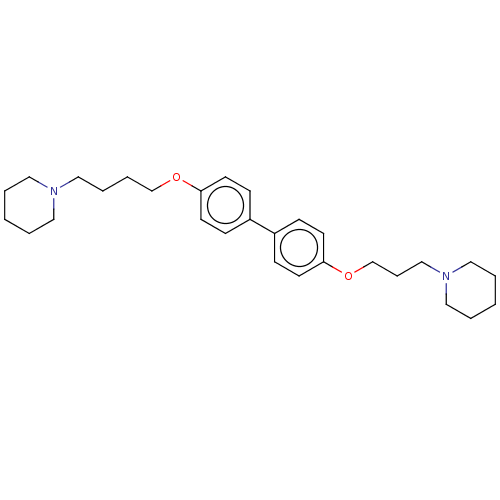
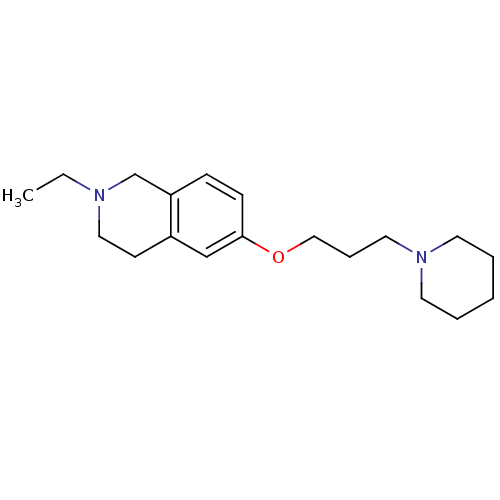
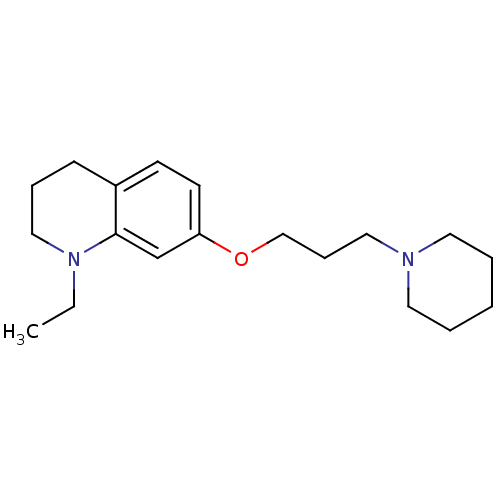
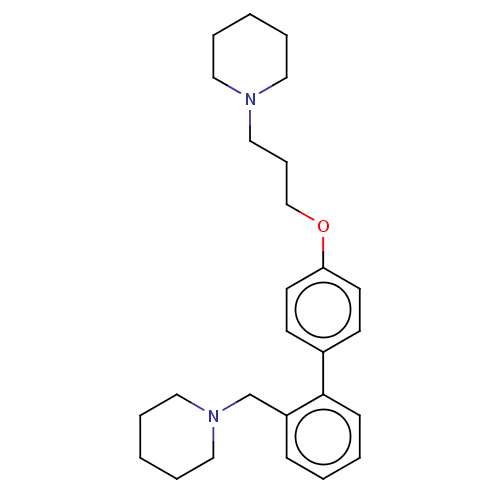
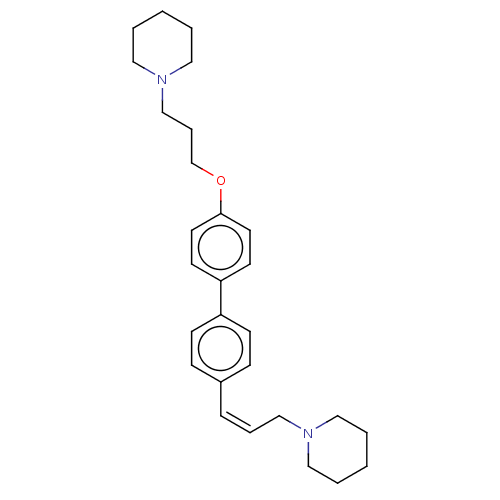
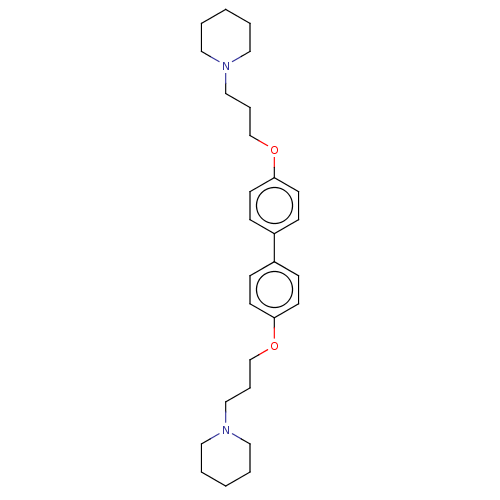
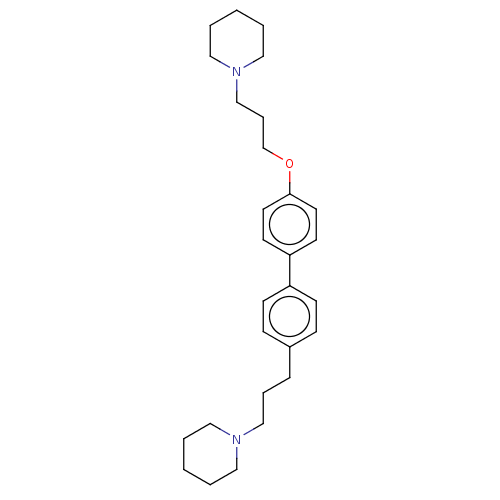
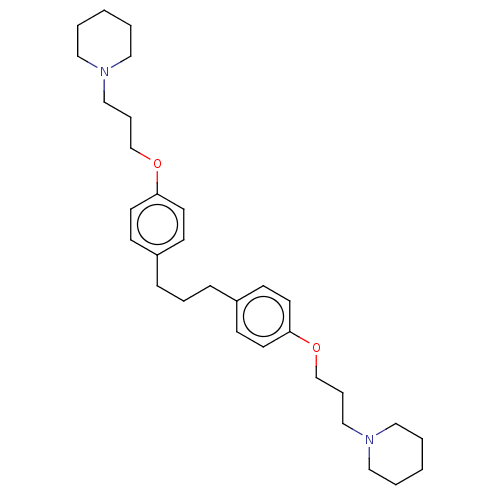
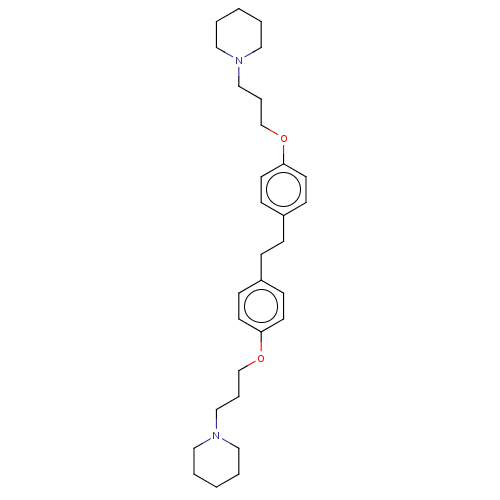
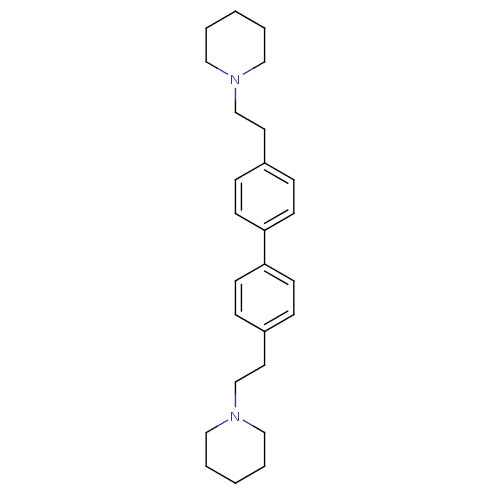
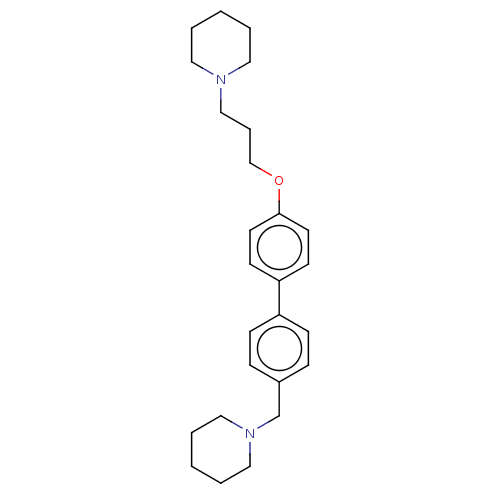
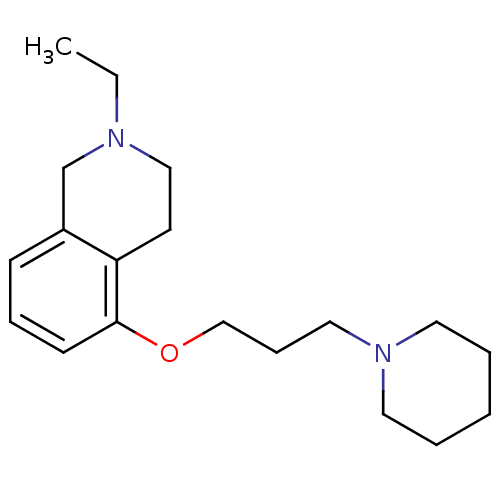
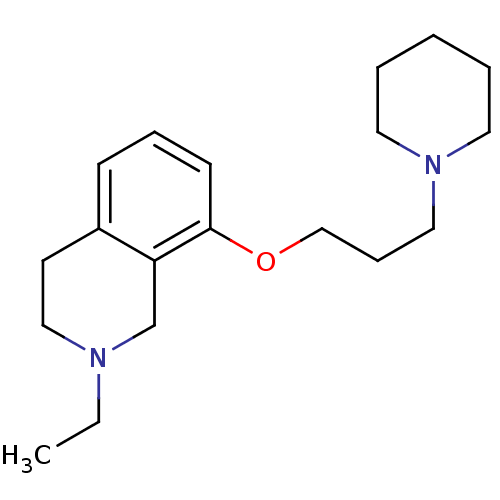
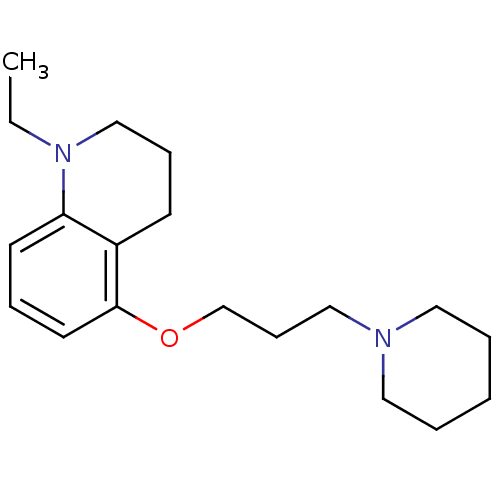
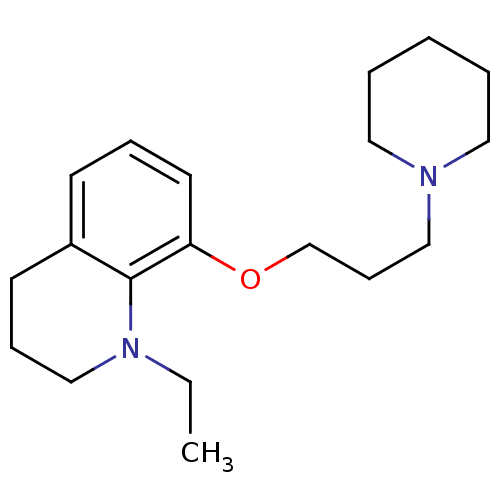
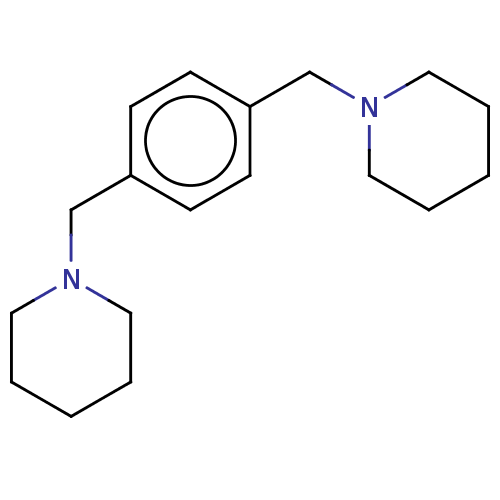
Report error Found 37 Enz. Inhib. hit(s) with all data for entry = 50005555
Affinity DataKi: 0.0450nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0570nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.130nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.170nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.340nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.340nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.490nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.510nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.530nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.540nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.549nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.640nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.655nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.670nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 6.60nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 9.30nMAssay Description:Displacement of [125I]iodoproxyfan from histamine H3 receptor (unknown origin) expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 84nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 95nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 156nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.23E+3nMAssay Description:Binding affinity to histamine H3 receptor (unknown origin)More data for this Ligand-Target Pair