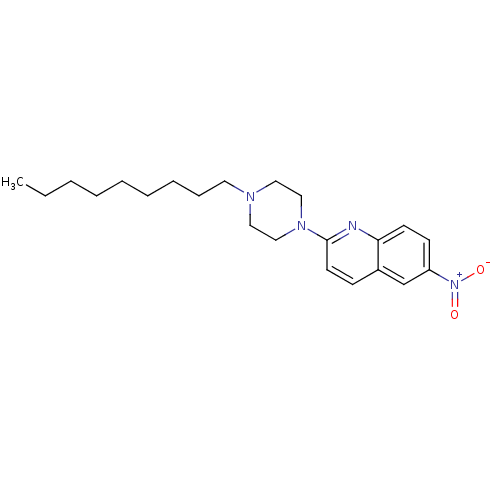
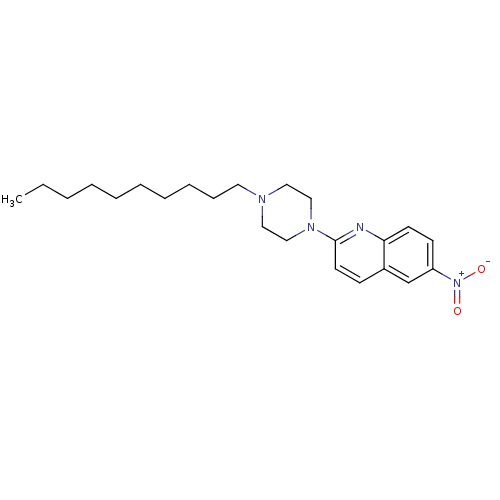
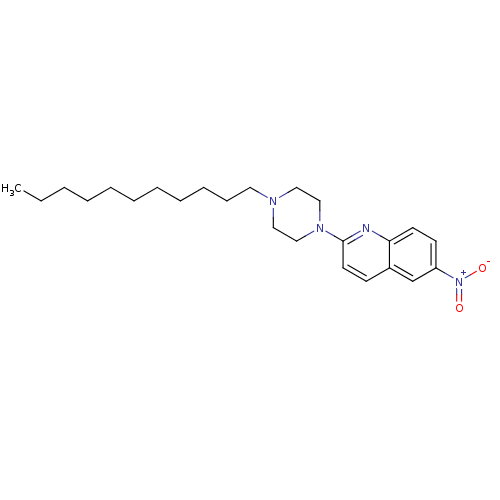
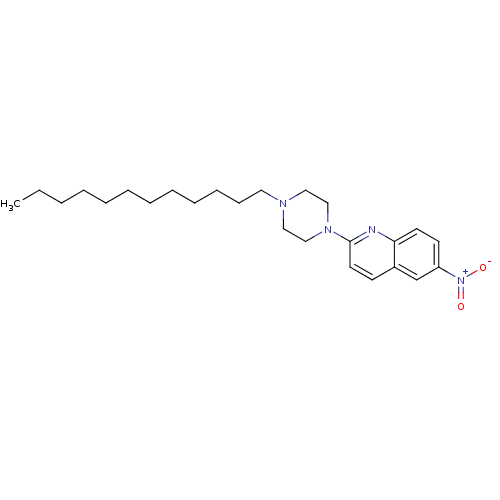
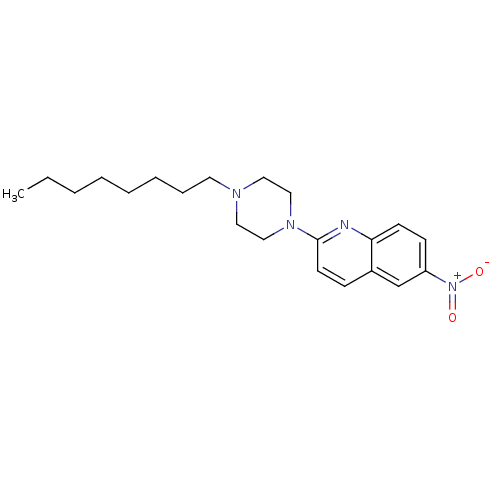
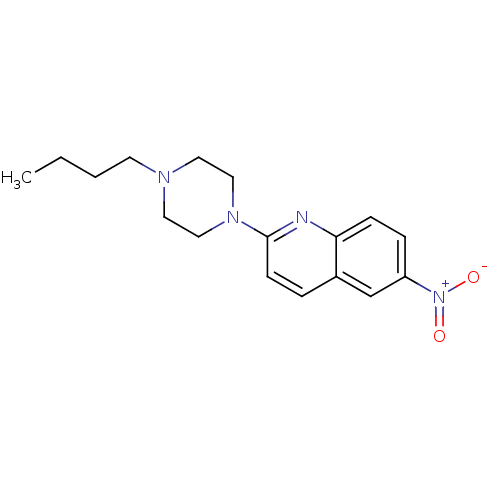
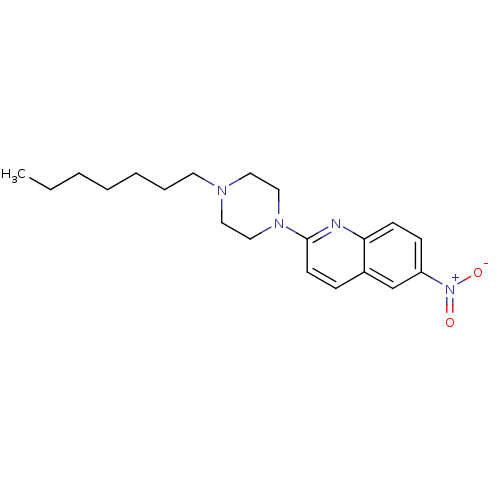
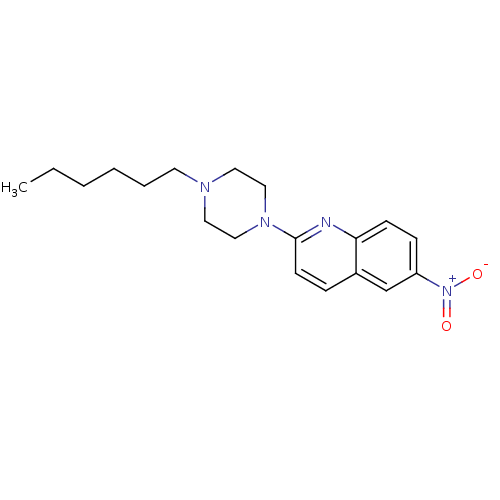
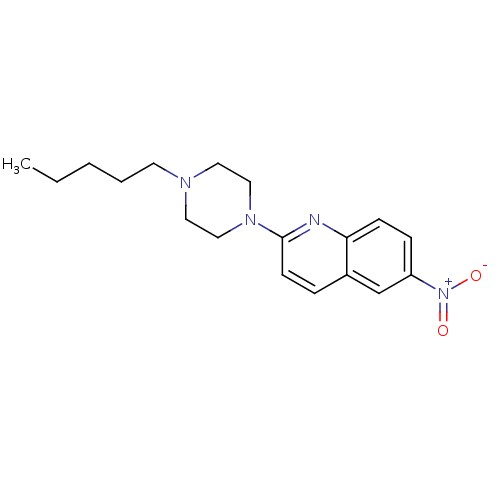
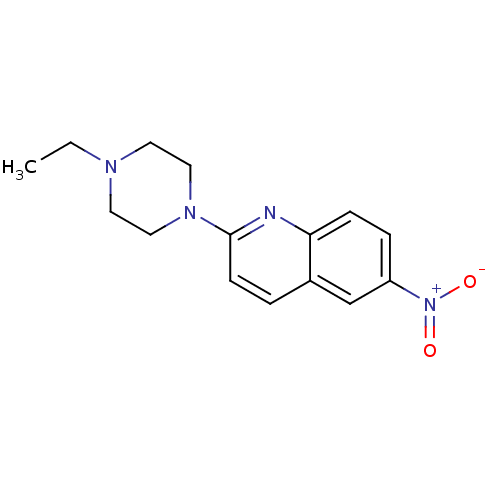
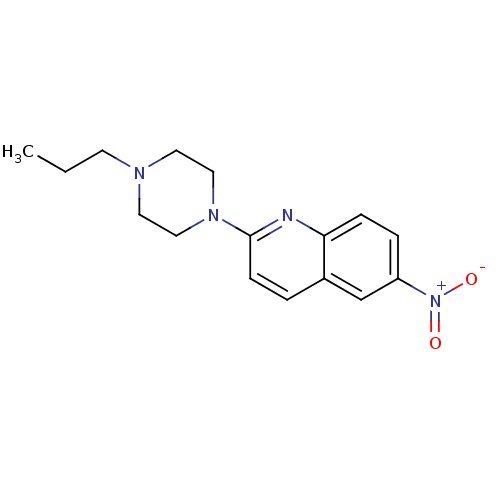
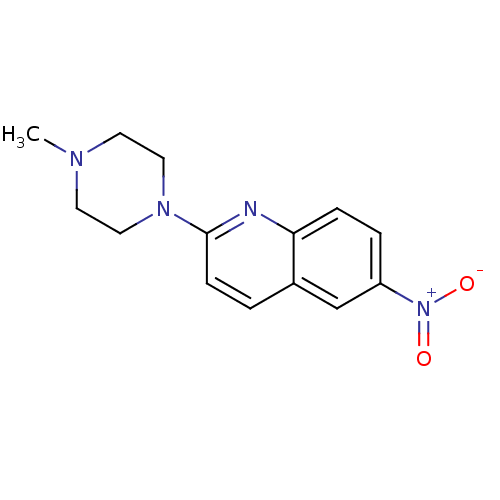
Report error Found 12 Enz. Inhib. hit(s) with all data for entry = 6038
Affinity DataKi: 1.80nM ΔG°: -49.9kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nM ΔG°: -47.6kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 12.6nM ΔG°: -45.1kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 16.1nM ΔG°: -44.5kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 20.8nM ΔG°: -43.8kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 24nM ΔG°: -43.5kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 54.1nM ΔG°: -41.5kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 72.9nM ΔG°: -40.7kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 80.5nM ΔG°: -40.5kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 86.1nM ΔG°: -40.3kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 496nM ΔG°: -36.0kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nM ΔG°: -33.6kJ/molepH: 7.7 T: 2°CAssay Description:The assay was performed in accordance with the method described by Owens et al. with slight modifications. Rat cerebral cortex was homogenized in 30...More data for this Ligand-Target Pair