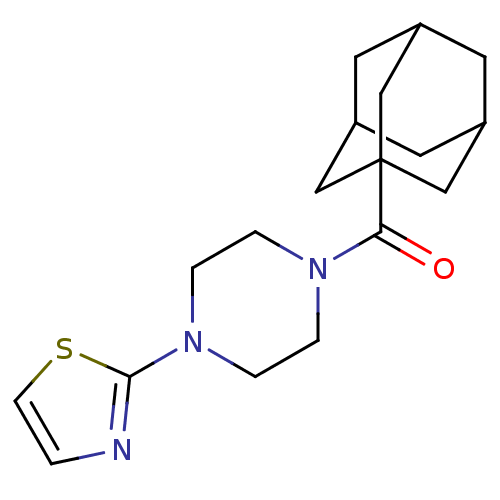
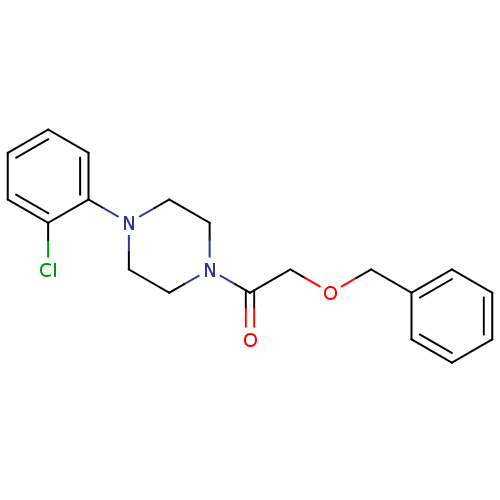
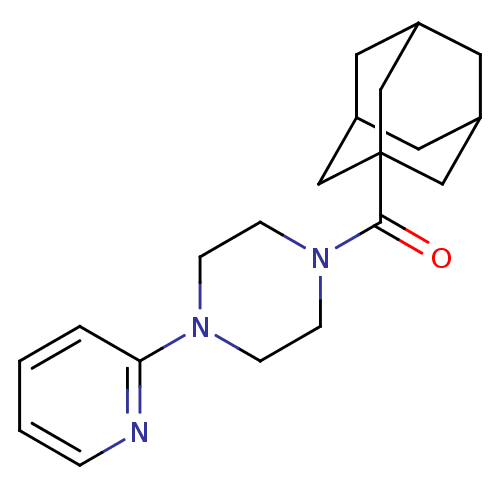
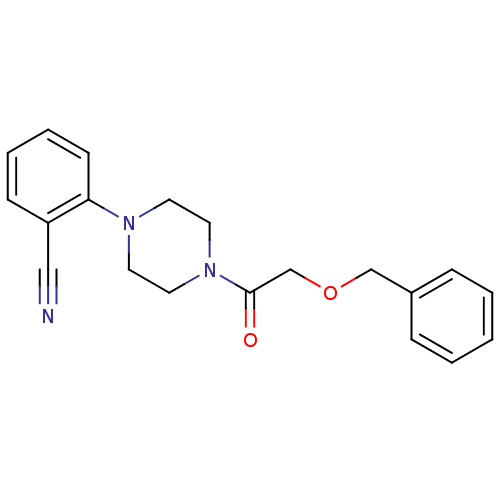
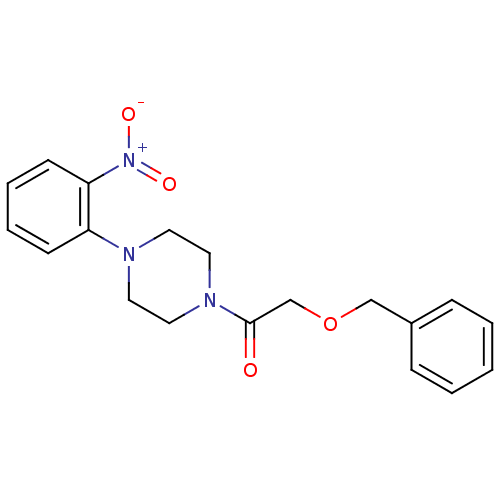
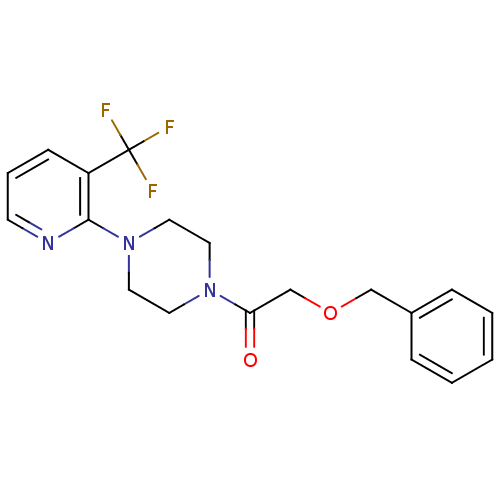
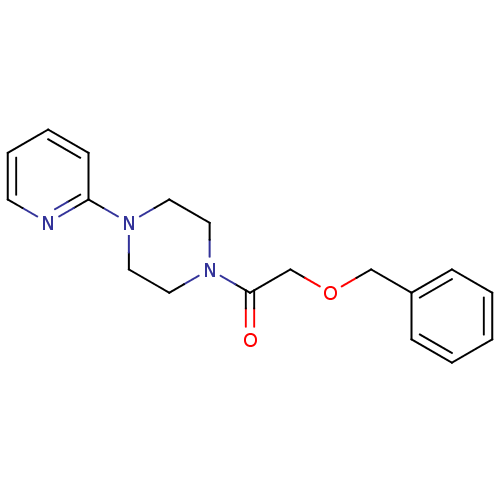
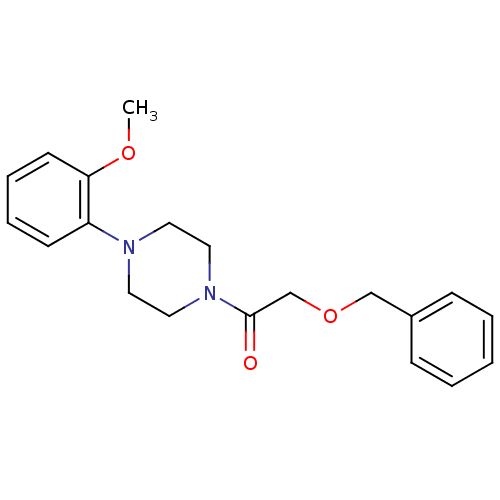
Report error Found 13 Enz. Inhib. hit(s) with all data for entry = 50033067
Affinity DataKi: 440nMAssay Description:Negative allosteric modulation at rat mGluR5 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 540nMAssay Description:Negative allosteric modulation at rat mGluR5 receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 820nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataIC50: 990nMAssay Description:Negative allosteric modulation at rat mGluR5 receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 1.60E+3nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: 1.80E+3nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: 4.30E+3nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: 5.40E+3nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: 9.20E+3nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Positive allosteric modulation at rat mGluR5 receptor expressed in HEK293 cells assessed as glutamate-induced calcium fluorescence by Fluo-4/AM dye-b...More data for this Ligand-Target Pair