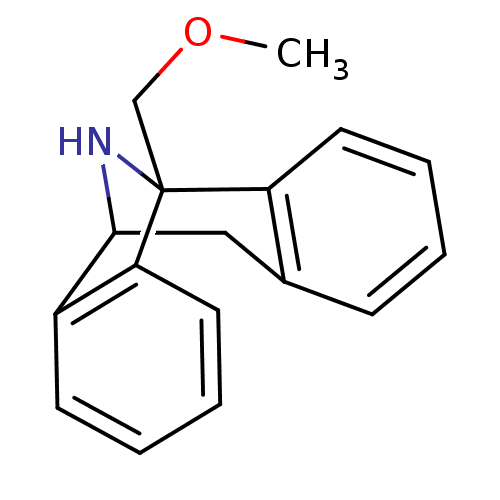
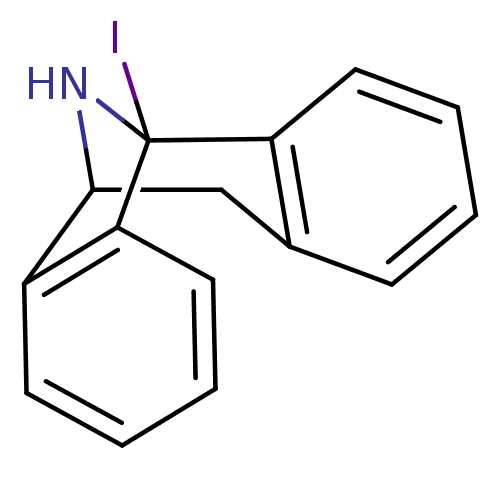
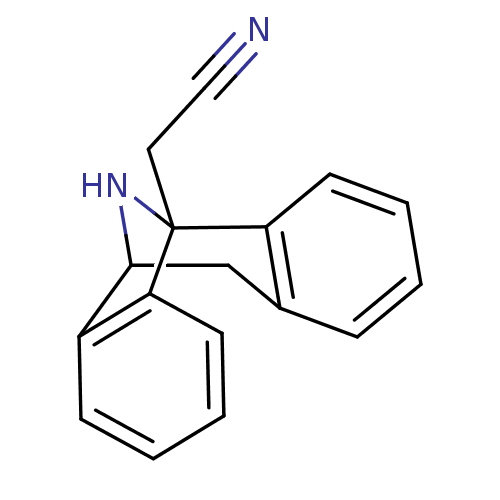
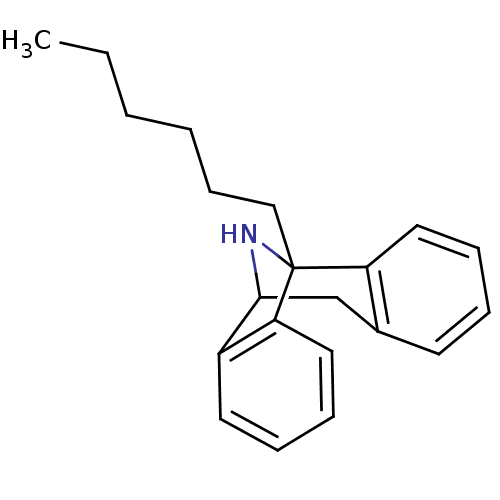
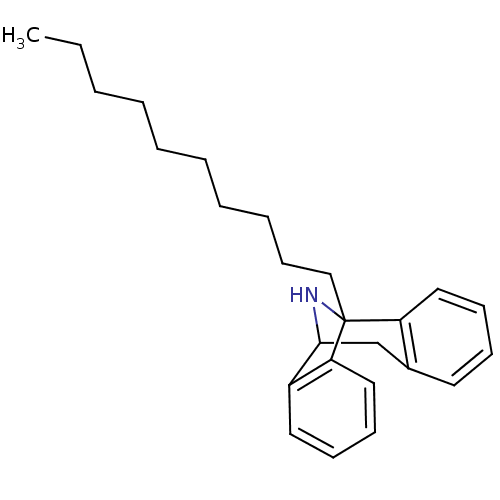
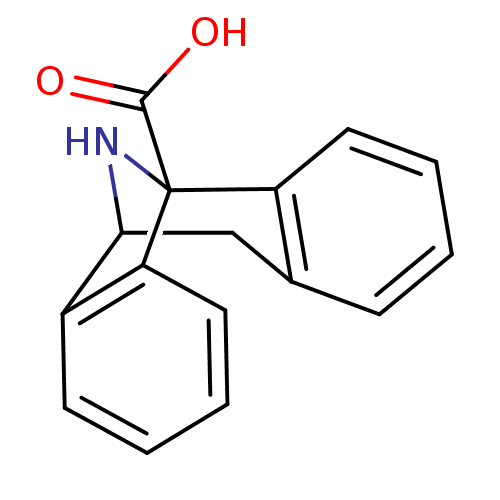
Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 50035864
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
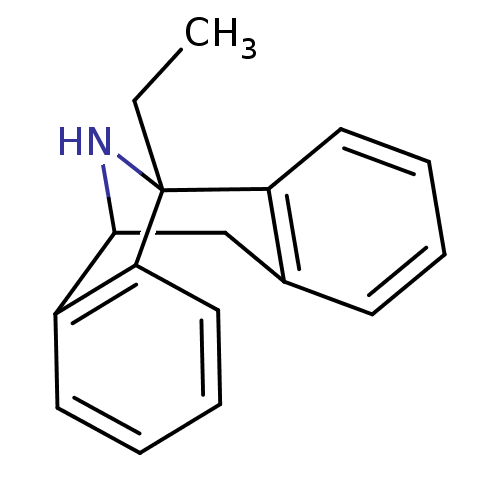
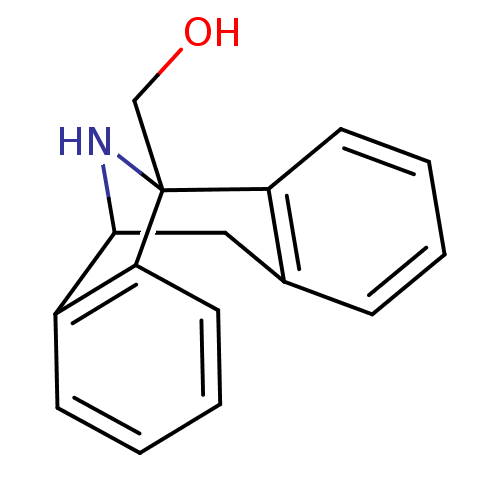
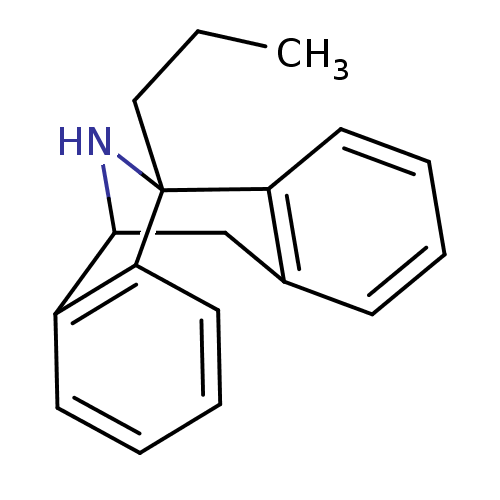
Affinity DataKi: 9nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
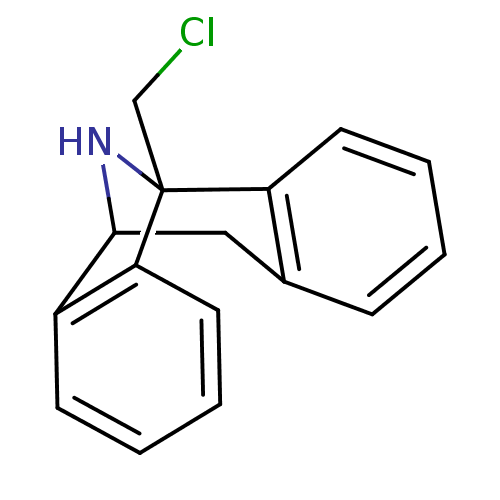
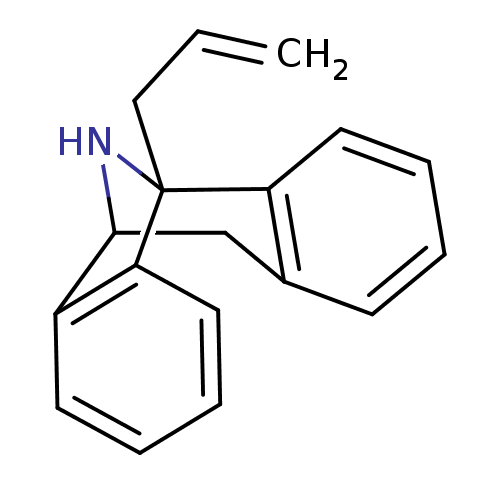
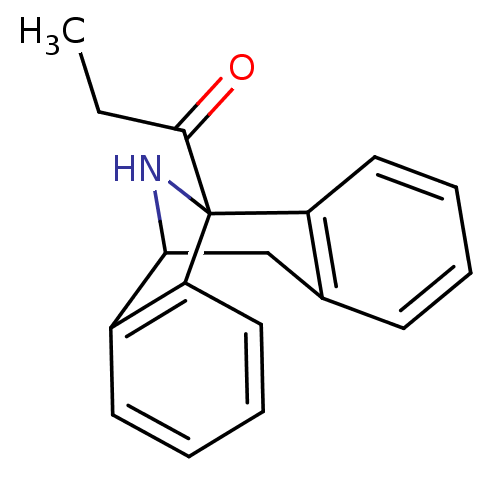
Affinity DataKi: 12nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
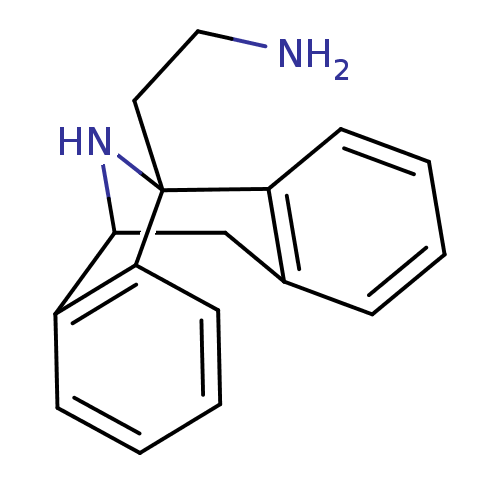
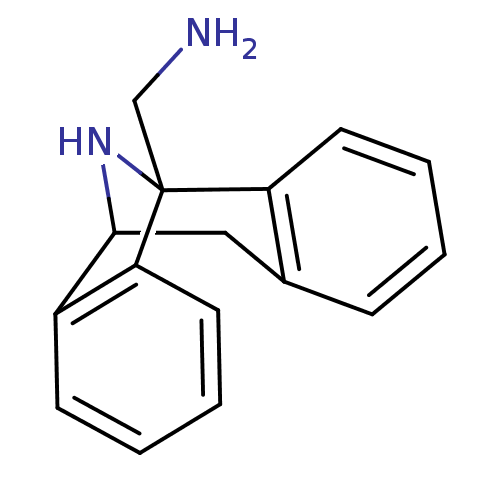
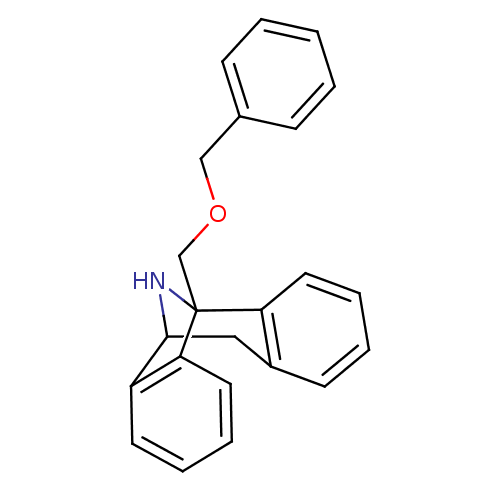
Affinity DataKi: 12nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
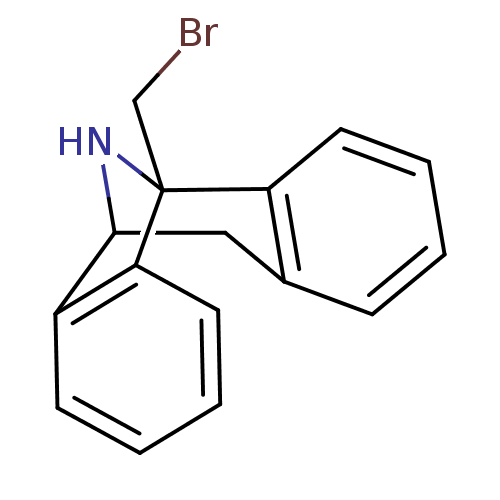
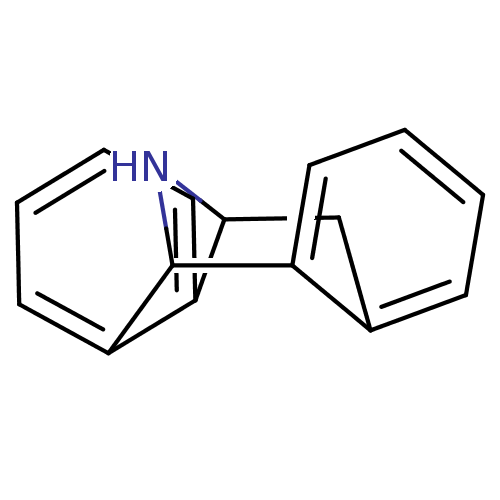
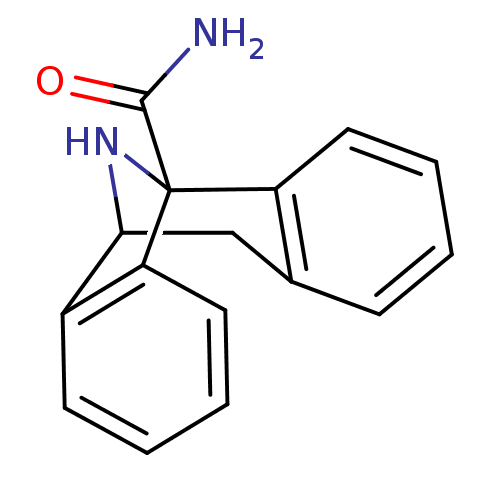
Affinity DataKi: 20nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 31nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 31nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 33nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 40nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 70nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 117nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 120nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 300nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 430nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 967nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 980nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 1.10E+3nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 7.50E+3nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 1.00E+4nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 1.13E+4nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 2.16E+4nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1(Rat)
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
National Institute of Diabetes and Digestive and Kidney Diseases
Curated by ChEMBL
Affinity DataKi: 3.23E+4nMAssay Description:Ability to displace [3H]TCP from high affinity PCP N-methyl-D-aspartate glutamate receptor in rat brain homogenates in vitro was determinedMore data for this Ligand-Target Pair
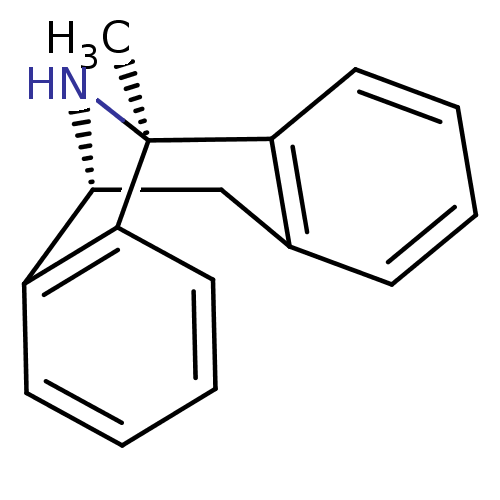
 3D Structure (crystal)
3D Structure (crystal)