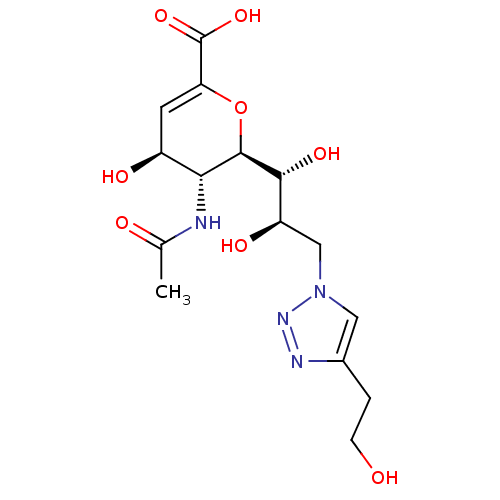
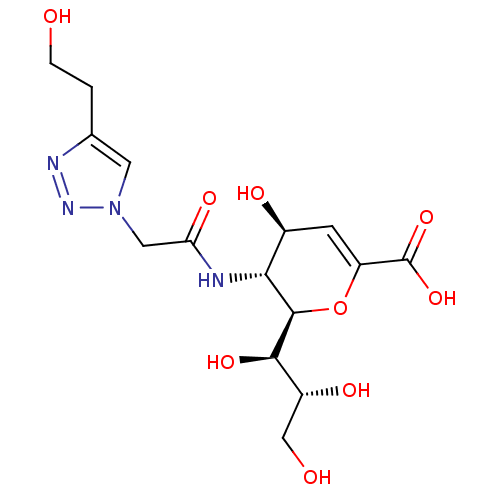
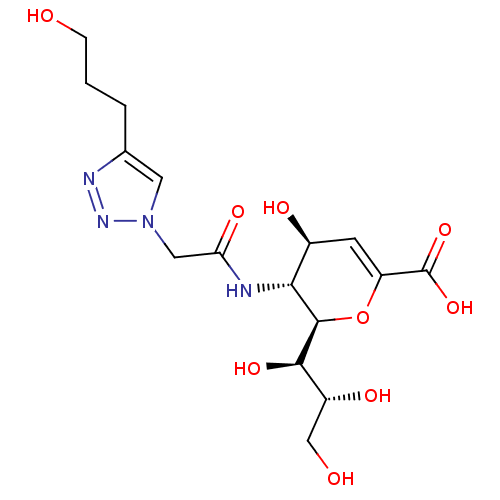
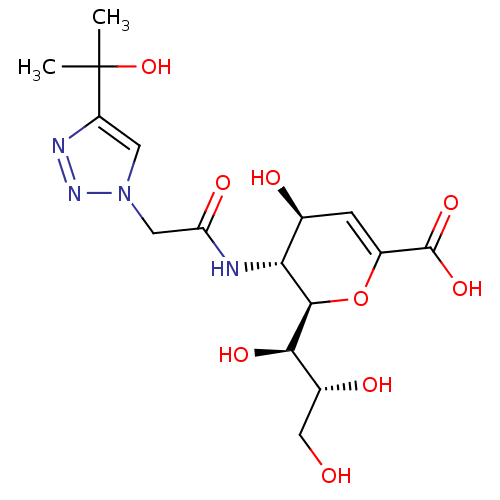
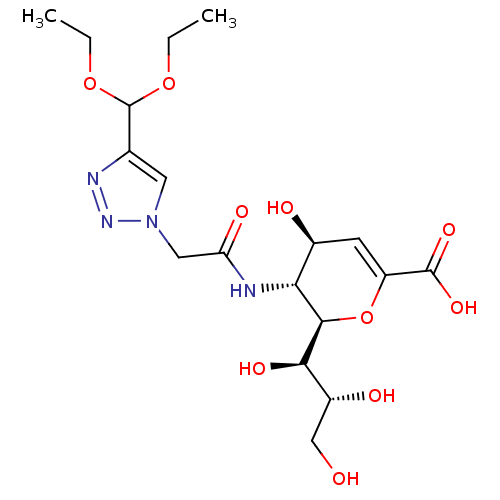
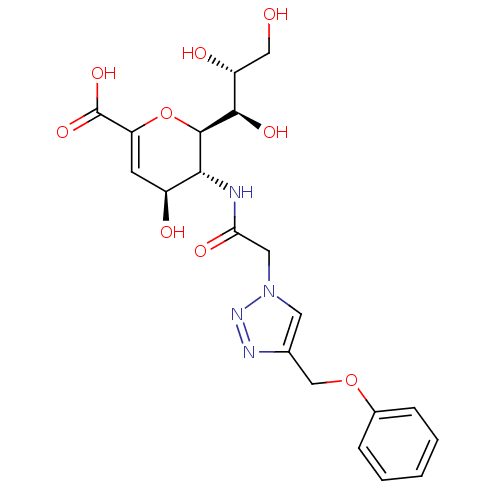
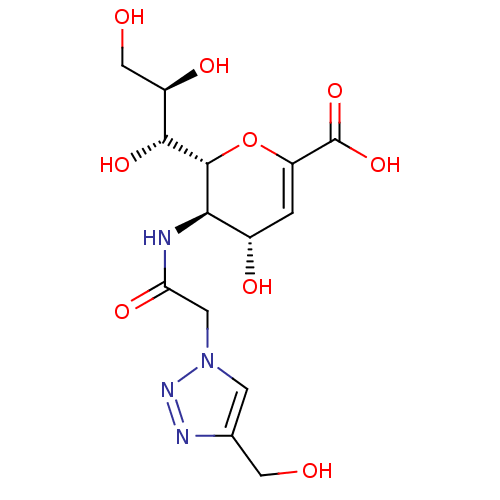
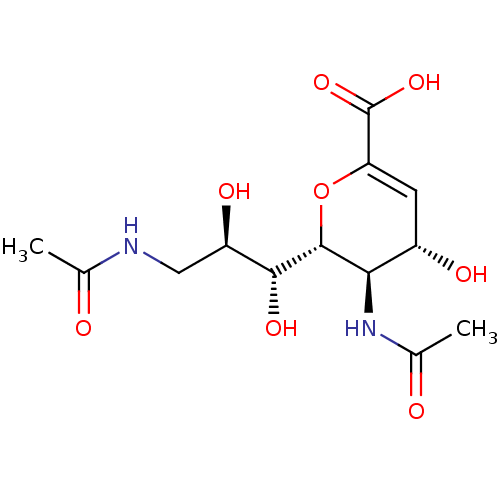
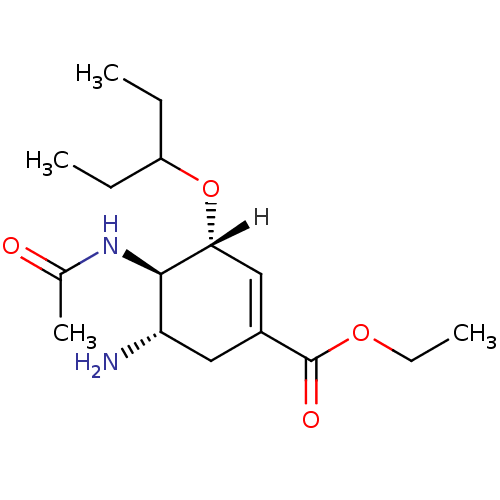
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 50039317
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of GM3 hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+4nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of GM3 hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysis by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of 4-methylumbelliferyl-alpha-D-glucopyranoside hydrolysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+7nMAssay Description:Inhibition of human neuraminidase 3 assessed as inhibition of GM3 hydrolysisMore data for this Ligand-Target Pair