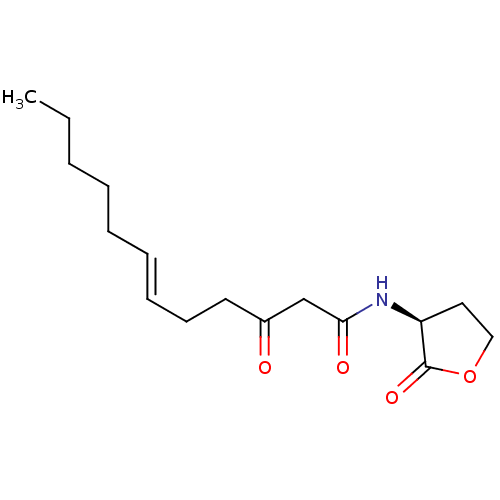
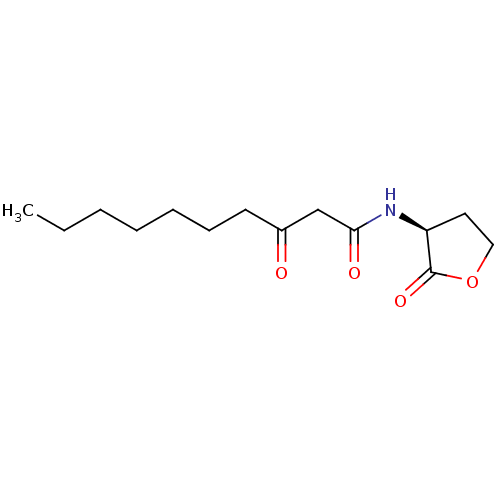
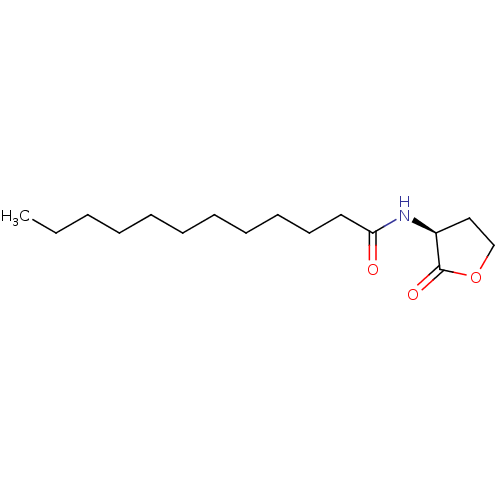
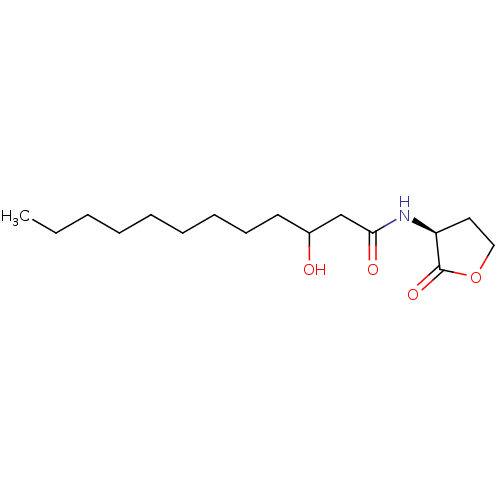
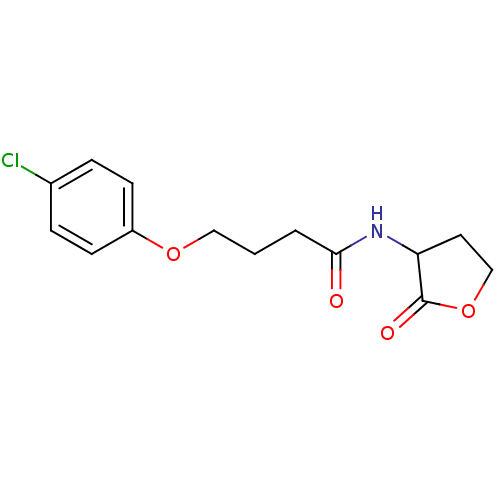
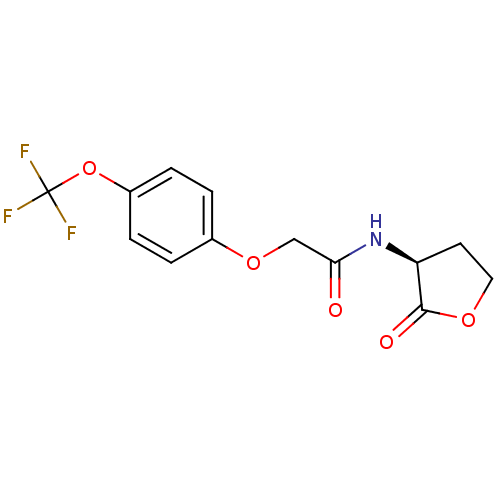
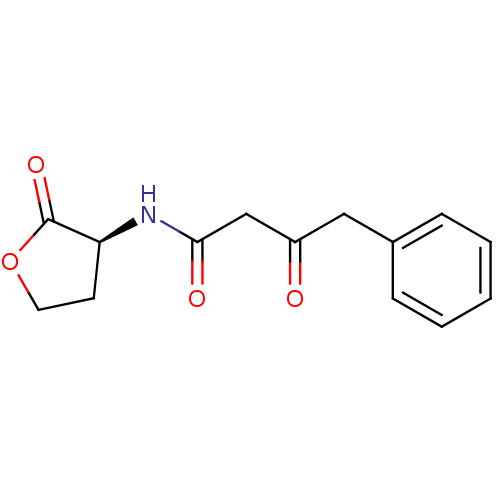
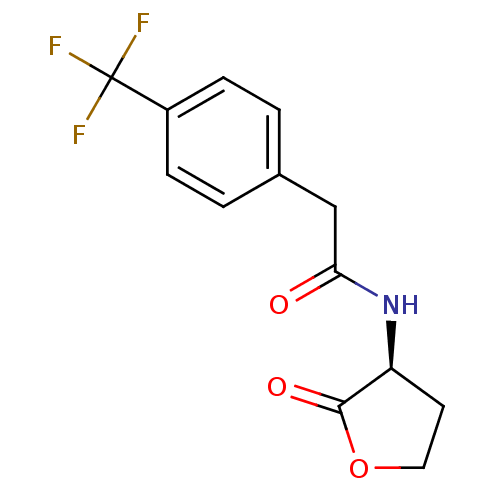
Report error Found 41 Enz. Inhib. hit(s) with all data for entry = 50033801
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
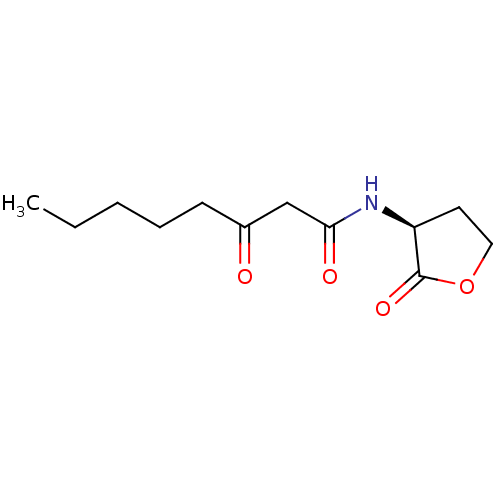
Affinity DataEC50: 7nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
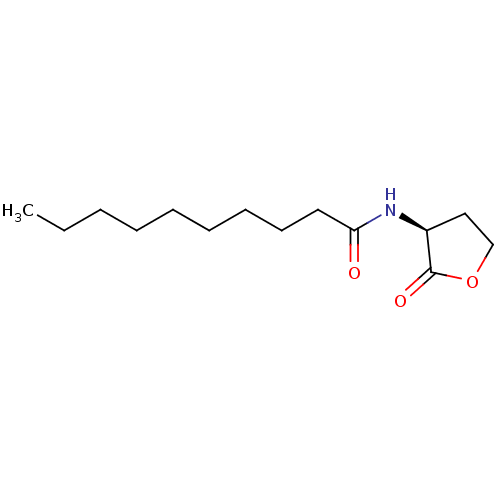
Affinity DataEC50: 7nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
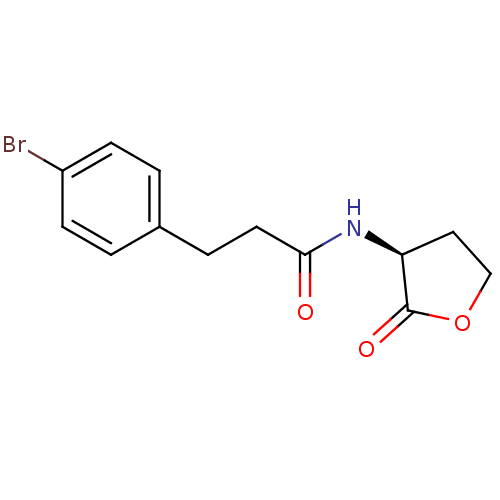
Affinity DataEC50: 10nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in absence of natural autoinducerMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
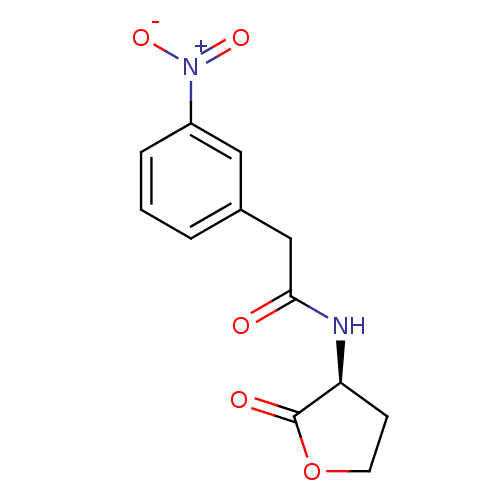
Affinity DataEC50: 10nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 15nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 15nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 33nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 40nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in absence of natural autoinducerMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 54nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in presence of 7.5 nM natural auto-inducer 3-oxo-C12-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 150nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR cognate receptor expressed in Escherichia coli harboring pKDT17 assessed as activation of lasB expres...More data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in presence of 7.5 nM natural auto-inducer 3-oxo-C12-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 340nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in presence of 7.5 nM natural auto-inducer 3-oxo-C12-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LuxR(Vibrio fischeri)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 350nMAssay Description:Agonist activity at Vibrio fischeri luxR receptor in absence of natural autoinducerMore data for this Ligand-Target Pair
TargetCviR transcriptional regulator(Chromobacterium violaceum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 377nMAssay Description:Inhibition of Chromobacterium violaceum CviR-dependent GFP fluorescence in Escherichia coliMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LuxR(Vibrio fischeri)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Antagonist activity at Vibrio fischeri luxR receptor in presence of 5 uM 3-oxo-C6-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein TraR(Agrobacterium tumefaciens)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 460nMAssay Description:Antagonist activity at Agrobacterium tumefaciens TraR receptor in presence of 100 nM 3-oxo-C8-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataEC50: 540nMAssay Description:Agonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in absence of natural autoinducerMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LuxR(Vibrio fischeri)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 610nMAssay Description:Antagonist activity at Vibrio fischeri luxR receptor in presence of 5 uM 3-oxo-C6-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein TraR(Agrobacterium tumefaciens)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 690nMAssay Description:Antagonist activity at Agrobacterium tumefaciens TraR receptor in presence of 100 nM 3-oxo-C8-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein TraR(Agrobacterium tumefaciens)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 770nMAssay Description:Antagonist activity at Agrobacterium tumefaciens TraR receptor in presence of 100 nM 3-oxo-C8-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LuxR(Vibrio fischeri)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 770nMAssay Description:Antagonist activity at Vibrio fischeri luxR receptor in presence of 5 uM 3-oxo-C6-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein TraR(Agrobacterium tumefaciens)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 810nMAssay Description:Antagonist activity at Agrobacterium tumefaciens TraR receptor in presence of 100 nM 3-oxo-C8-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LuxR(Vibrio fischeri)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 860nMAssay Description:Antagonist activity at Vibrio fischeri luxR receptor in presence of 5 uM 3-oxo-C6-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein TraR(Agrobacterium tumefaciens)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 920nMAssay Description:Antagonist activity at Agrobacterium tumefaciens TraR receptor in presence of 100 nM 3-oxo-C8-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein TraR(Agrobacterium tumefaciens)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.05E+3nMAssay Description:Antagonist activity at Agrobacterium tumefaciens TraR receptor in presence of 100 nM 3-oxo-C8-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein TraR(Agrobacterium tumefaciens)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.25E+3nMAssay Description:Antagonist activity at Agrobacterium tumefaciens TraR receptor in presence of 100 nM 3-oxo-C8-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LuxR(Vibrio fischeri)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.35E+3nMAssay Description:Antagonist activity at Vibrio fischeri luxR receptor in presence of 5 uM 3-oxo-C6-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LuxR(Vibrio fischeri)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.36E+3nMAssay Description:Antagonist activity at Vibrio fischeri luxR receptor in presence of 5 uM 3-oxo-C6-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.72E+3nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in presence of 7.5 nM natural auto-inducer 3-oxo-C12-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.75E+3nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in presence of 7.5 nM natural auto-inducer 3-oxo-C12-HSLMore data for this Ligand-Target Pair
TargetAutoinducer 1 sensor kinase/phosphatase LuxN(Vibrio harveyi)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of LuxN-dependent bioluminescence in Vibrio harveyiMore data for this Ligand-Target Pair
TargetAutoinducer 1 sensor kinase/phosphatase LuxN(Vibrio harveyi)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of LuxN-dependent bioluminescence in Vibrio harveyiMore data for this Ligand-Target Pair
TargetAutoinducer 1 sensor kinase/phosphatase LuxN(Vibrio harveyi)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibition of LuxN-dependent bioluminescence in Vibrio harveyiMore data for this Ligand-Target Pair
TargetAutoinducer 1 sensor kinase/phosphatase LuxN(Vibrio harveyi)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of LuxN-dependent bioluminescence in Vibrio harveyiMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.67E+3nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor expressed in Escherichia coli in presence of 7.5 nM natural auto-inducer 3-oxo-C12-HSLMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.52E+4nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor assessed as inhibition of bioluminescence by reporter gene assayMore data for this Ligand-Target Pair
TargetAutoinducer 1 sensor kinase/phosphatase LuxN(Vibrio harveyi)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 6.23E+4nMAssay Description:Inhibition of LuxN-dependent bioluminescence in Vibrio harveyiMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor assessed as inhibition of bioluminescence by reporter gene assayMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.13E+5nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor assessed as inhibition of bioluminescence by reporter gene assayMore data for this Ligand-Target Pair
TargetTranscriptional activator protein LasR(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.00E+5nMAssay Description:Antagonist activity at Pseudomonas aeruginosa LasR receptor assessed as inhibition of bioluminescence by reporter gene assayMore data for this Ligand-Target Pair
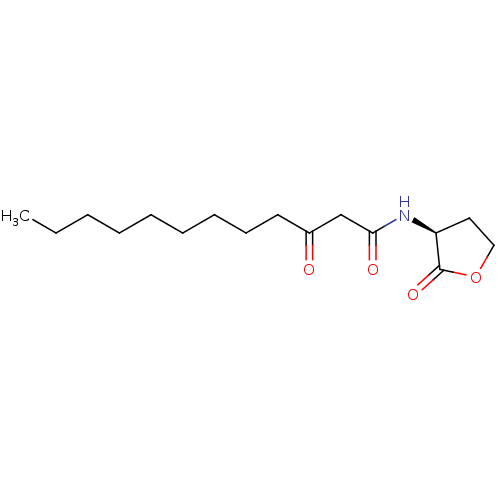
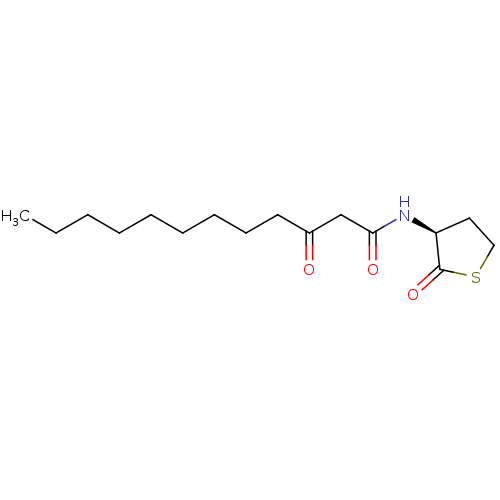
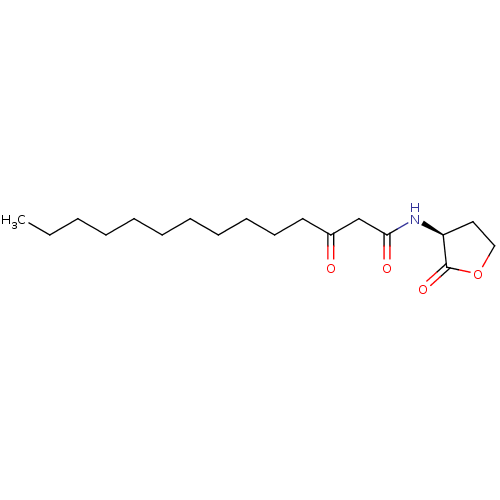
 3D Structure (crystal)
3D Structure (crystal)