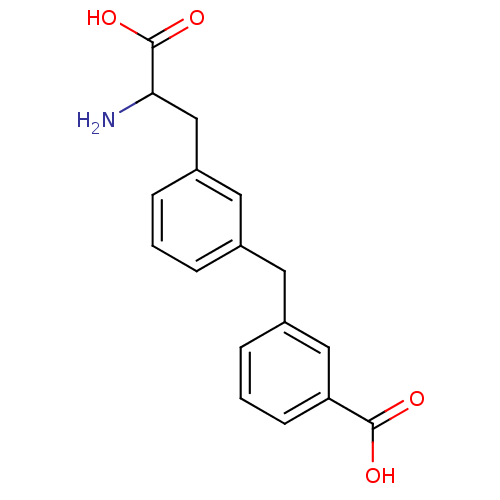
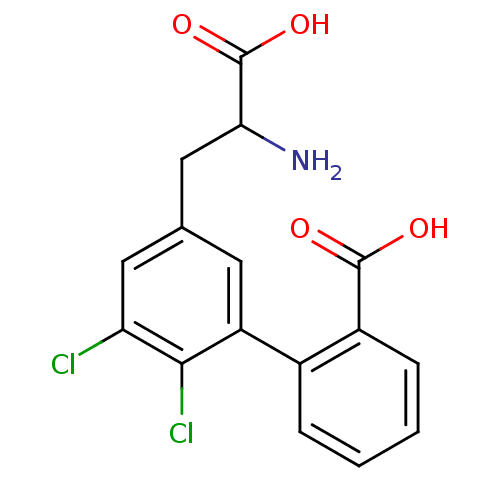
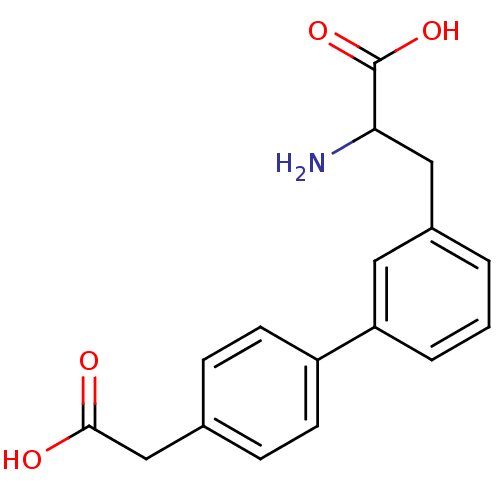
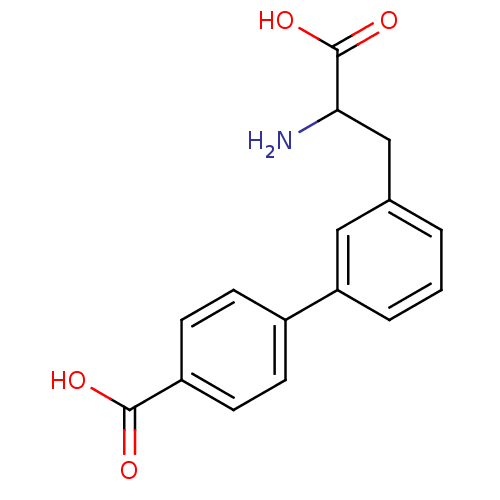
Report error Found 20 Enz. Inhib. hit(s) with all data for entry = 50030924
Affinity DataKi: 2.80E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.90E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKd: 3.94E+3nMAssay Description:Displacement of [3H]S-glutamate from rat GluR5 S1S2 ligand binding core expressed in Escherichia coli (DE3)More data for this Ligand-Target Pair
Affinity DataKi: 4.08E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 7.90E+3nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+4nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair
Affinity DataKd: 1.46E+5nMAssay Description:Displacement of [3H]AMPA from wild type rat GluR2 S1S2 ligand binding coreMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+6nMAssay Description:Displacement of [3H]SYM2081 from rat recombinant iGluR5-1b expressed in baculovirus-infected insect Sf9 cellsMore data for this Ligand-Target Pair