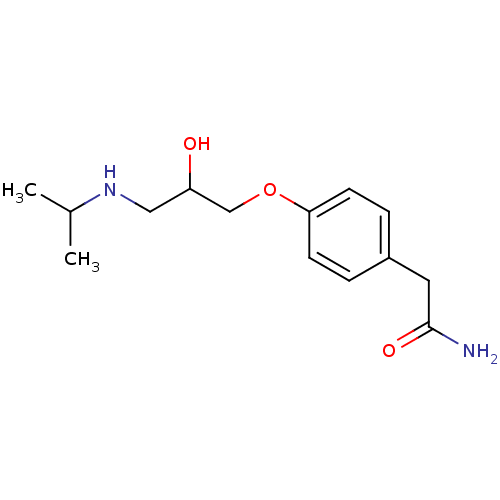
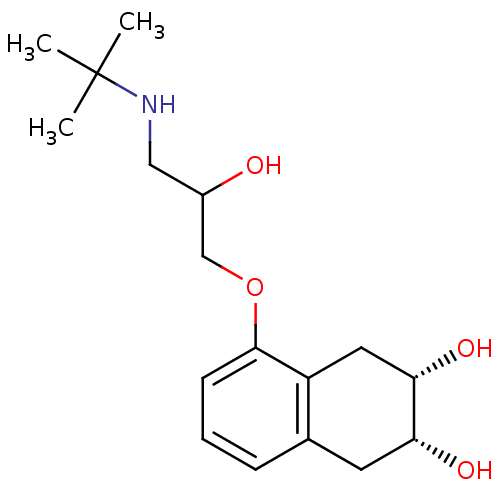
Report error Found 45 Enz. Inhib. hit(s) with all data for entry = 50038667
Affinity DataKi: 24nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 27nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKd: 200nMAssay Description:Binding affinity to L-FABP high binding affinity site by titration calorimetry methodMore data for this Ligand-Target Pair
Affinity DataKi: 256nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 334nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 379nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 405nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 531nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKd: 900nMAssay Description:Binding affinity to L-FABP low binding affinity site by titration calorimetry methodMore data for this Ligand-Target Pair
TargetFatty acid-binding protein, intestinal(Human)
Monash University (Parkville Campus)
Curated by ChEMBL
Monash University (Parkville Campus)
Curated by ChEMBL
Affinity DataKi: 1.00E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from I-FABPMore data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 1.27E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 1.86E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKd: 2.00E+3nMAssay Description:Binding affinity to human L-FABPMore data for this Ligand-Target Pair
Affinity DataKi: 2.66E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 2.90E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 3.22E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 6.92E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
TargetFatty acid-binding protein, intestinal(Human)
Monash University (Parkville Campus)
Curated by ChEMBL
Monash University (Parkville Campus)
Curated by ChEMBL
Affinity DataKi: 8.90E+3nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from I-FABPMore data for this Ligand-Target Pair
Affinity DataKi: 1.16E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 1.29E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
TargetFatty acid-binding protein, intestinal(Human)
Monash University (Parkville Campus)
Curated by ChEMBL
Monash University (Parkville Campus)
Curated by ChEMBL
Affinity DataKi: 2.00E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from I-FABPMore data for this Ligand-Target Pair
Affinity DataKi: 2.21E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
TargetFatty acid-binding protein, intestinal(Human)
Monash University (Parkville Campus)
Curated by ChEMBL
Monash University (Parkville Campus)
Curated by ChEMBL
Affinity DataKi: 2.60E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from I-FABPMore data for this Ligand-Target Pair
Affinity DataKi: 2.75E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
TargetFatty acid-binding protein, intestinal(Human)
Monash University (Parkville Campus)
Curated by ChEMBL
Monash University (Parkville Campus)
Curated by ChEMBL
Affinity DataKi: 3.30E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from I-FABPMore data for this Ligand-Target Pair
Affinity DataKi: 3.54E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 4.13E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 4.44E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 4.76E+4nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 1.01E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 1.15E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 1.19E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 1.61E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 1.79E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 2.22E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 3.48E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 4.48E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
Affinity DataKi: 7.17E+5nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 2.31E+6nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP high binding affinity site expressed in Escherichia coli BL21 by co...More data for this Ligand-Target Pair
Affinity DataKi: 3.78E+6nMAssay Description:Displacement of 1-anilinonaphthalene-8-sulphonic acid from rat recombinant L-FABP low binding affinity site expressed in Escherichia coli BL21 by com...More data for this Ligand-Target Pair
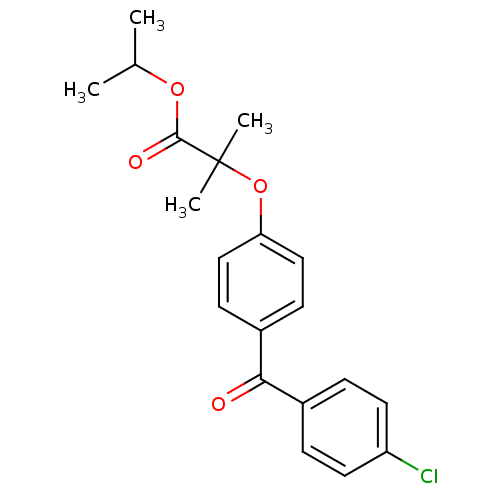
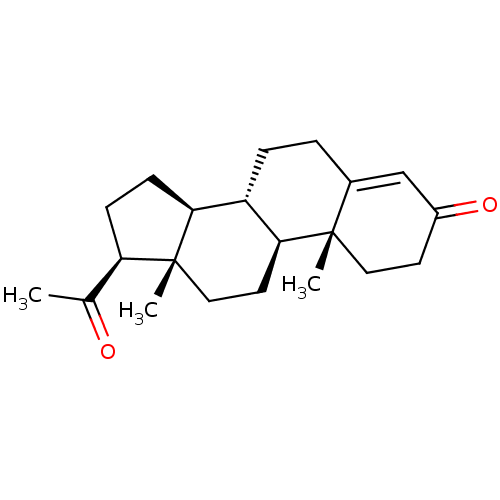
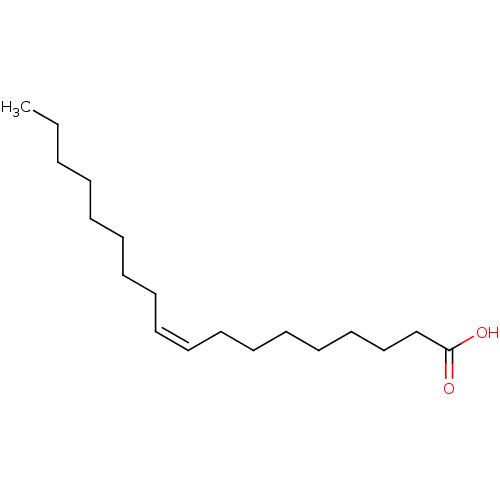
 3D Structure (crystal)
3D Structure (crystal)