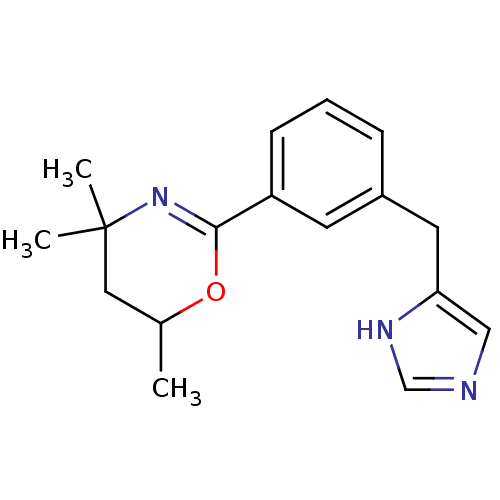
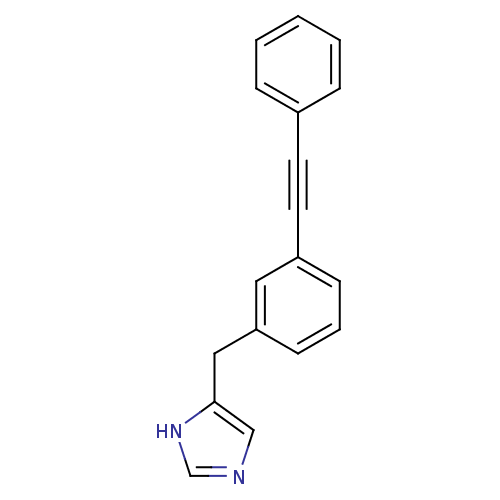
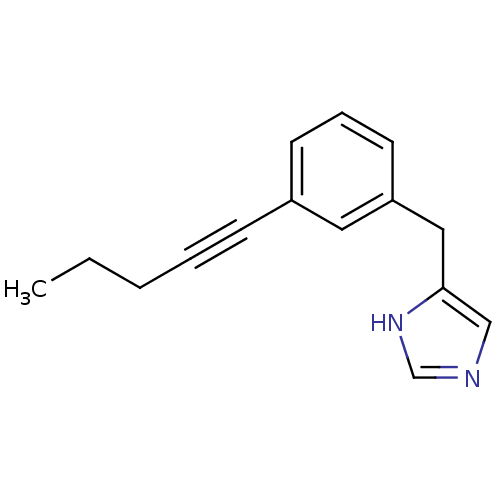
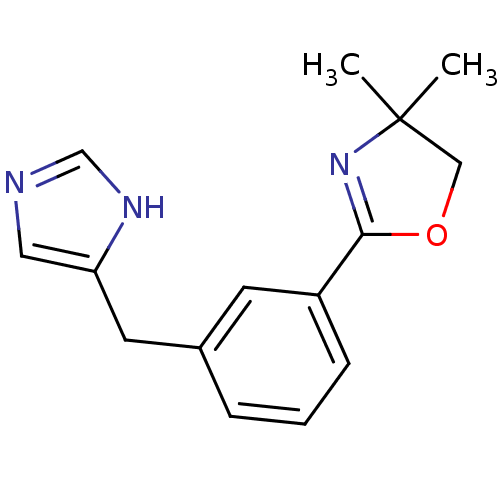
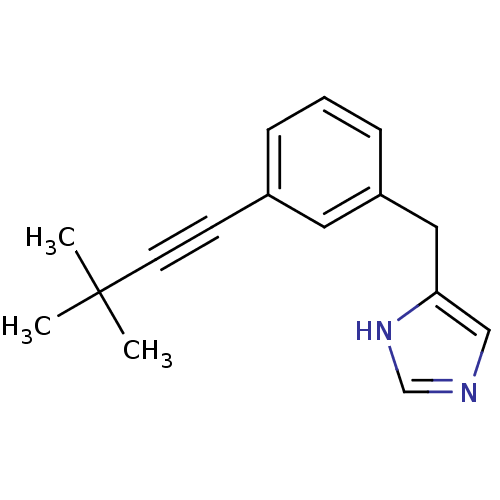
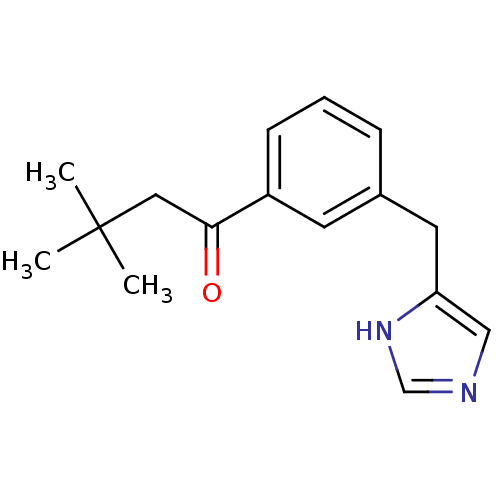
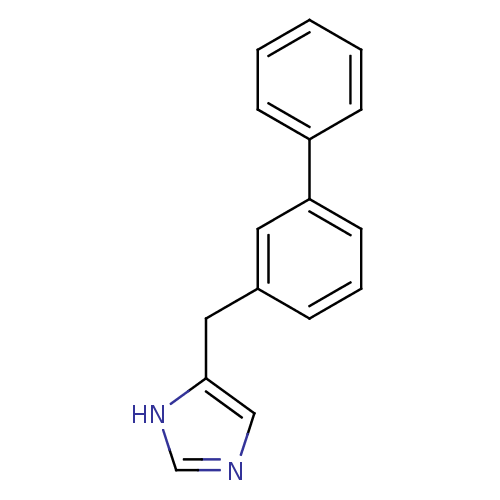
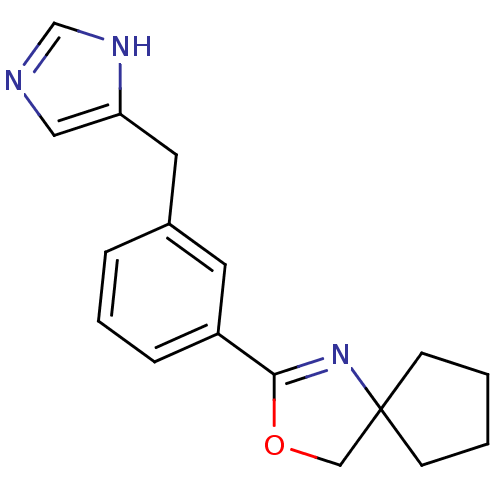
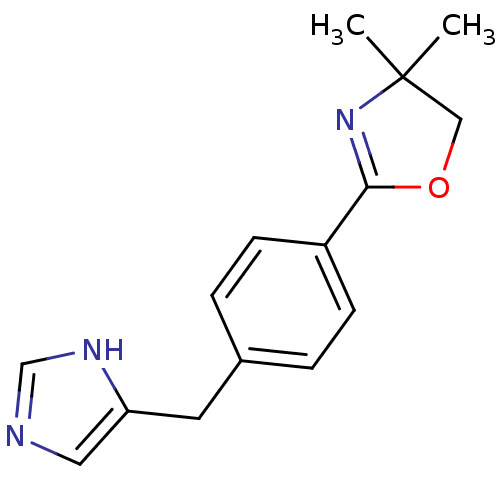
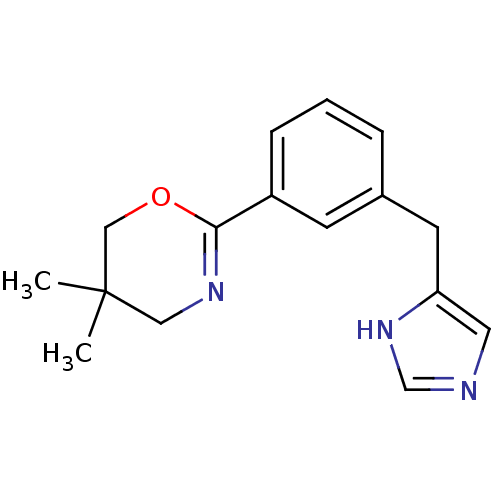
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 2605
Affinity DataKi: 0.339nM ΔG°: -54.1kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 0.407nM ΔG°: -53.6kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/mole EC50: 2nMpH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.7kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -48.6kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.9kJ/mole EC50: 16nMpH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.9kJ/mole EC50: 8nMpH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.9kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH4R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 6nM ΔG°: -46.9kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH4R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -46.2kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH4R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 32nM ΔG°: -42.8kJ/mole EC50: 79nMpH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 33nM ΔG°: -42.7kJ/mole EC50: 200nMpH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 40nM ΔG°: -42.2kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 41nM ΔG°: -42.2kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 56nM ΔG°: -41.4kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 89nM ΔG°: -40.2kJ/mole EC50: 79nMpH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 155nM ΔG°: -38.9kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 162nM ΔG°: -38.8kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 257nM ΔG°: -37.6kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 417nM ΔG°: -36.4kJ/mole EC50: 631nMpH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 427nM ΔG°: -36.4kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 537nM ΔG°: -35.8kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 708nM ΔG°: -35.1kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 813nM ΔG°: -34.8kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 1.74E+3nM ΔG°: -32.9kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 2.51E+3nM ΔG°: -32.0kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 2.51E+3nM ΔG°: -32.0kJ/molepH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH3R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:Ligand displacement assays were performed on CHO cells membranes expressing hH4R. Retained radioactivity was determined by liquid scintillation count...More data for this Ligand-Target Pair
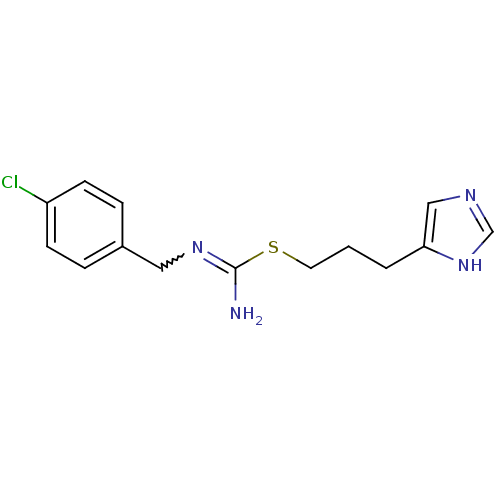
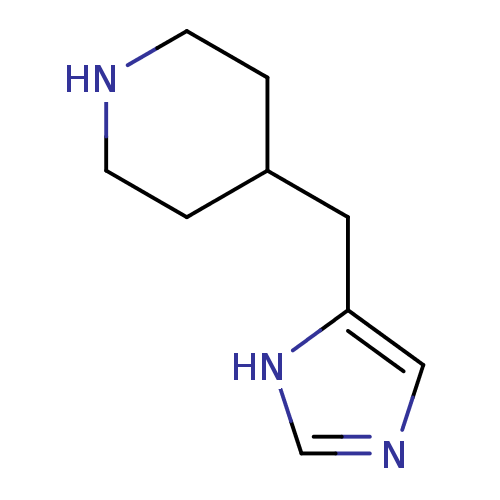
 3D Structure (crystal)
3D Structure (crystal)