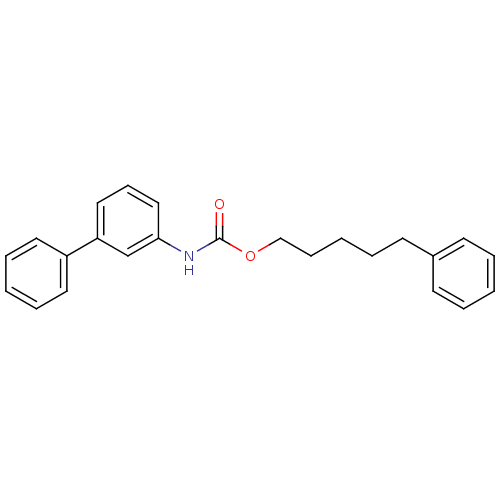
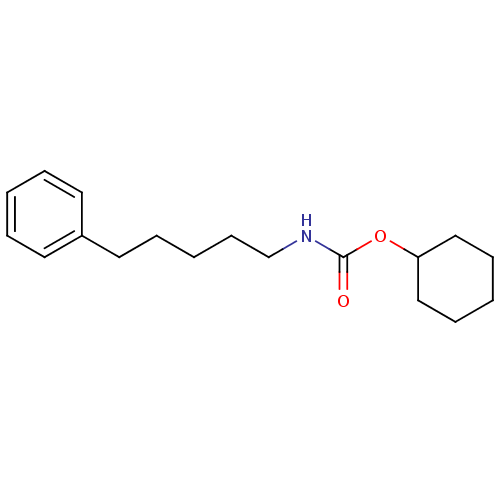
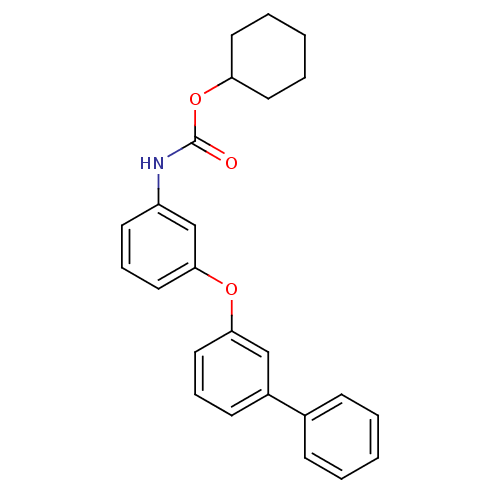
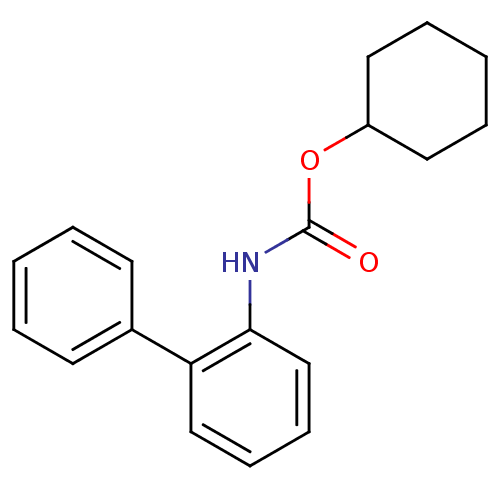
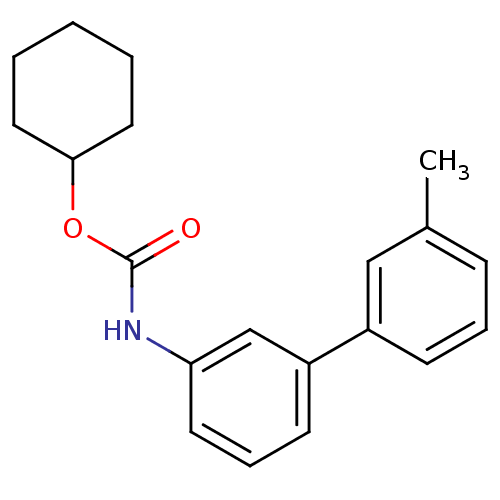
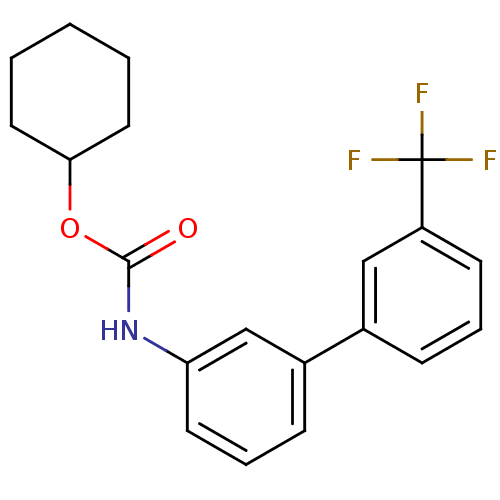
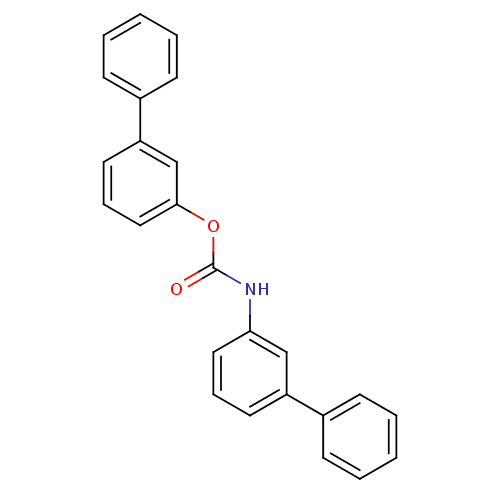
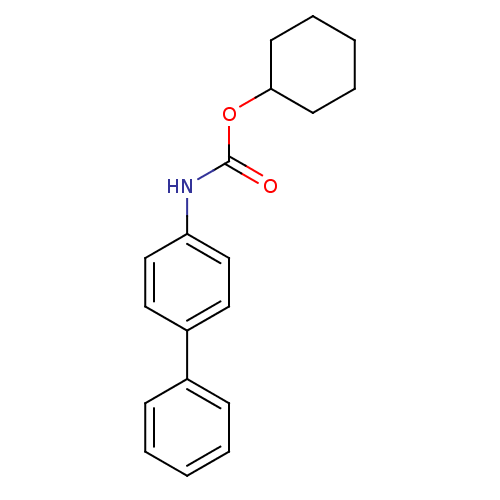
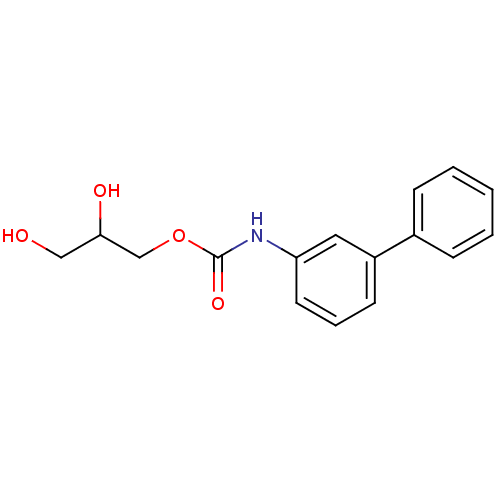
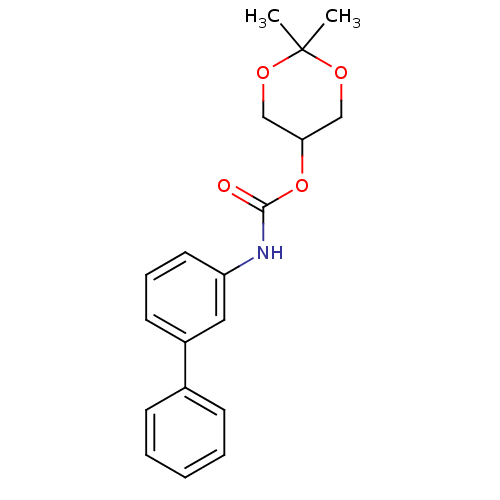
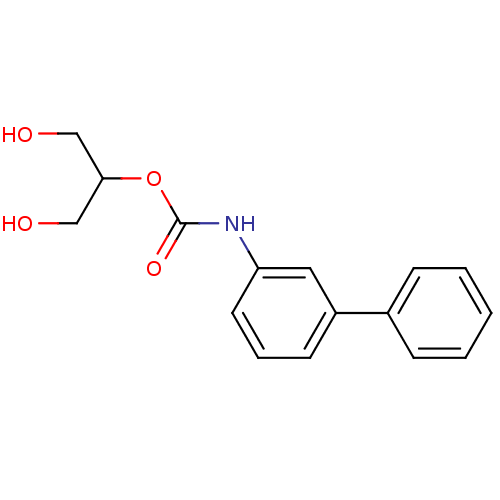
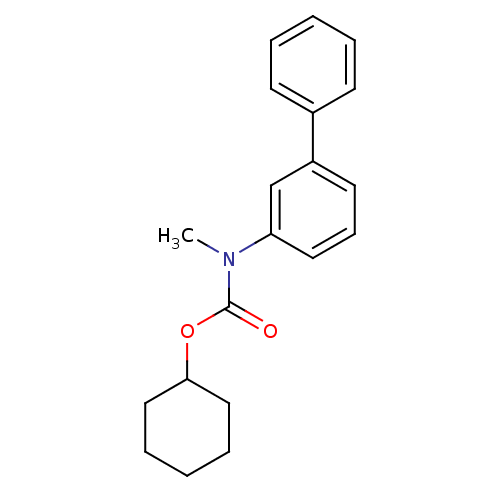
Report error Found 19 Enz. Inhib. hit(s) with all data for entry = 4617
Affinity DataIC50: 1.15E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 2.23E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 2.43E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMpH: 8.0Assay Description:The measurement of the ability of URB602 compound to inhibit MGL activity. More data for this Ligand-Target Pair