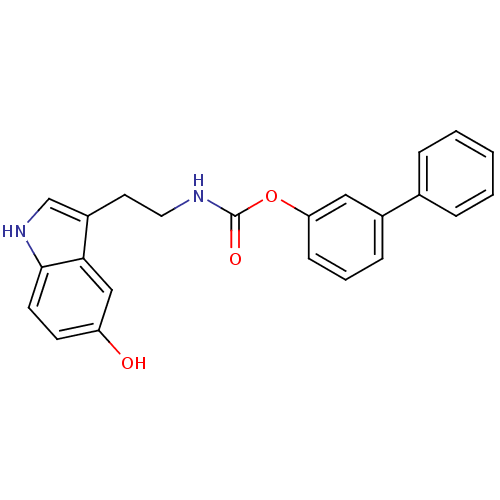
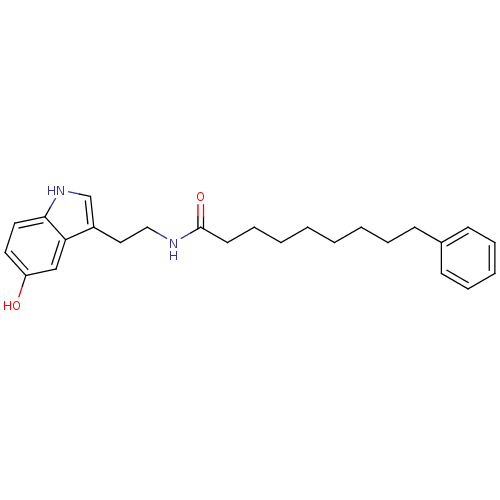
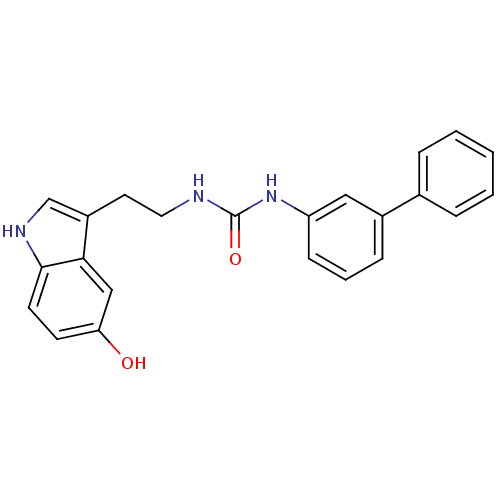
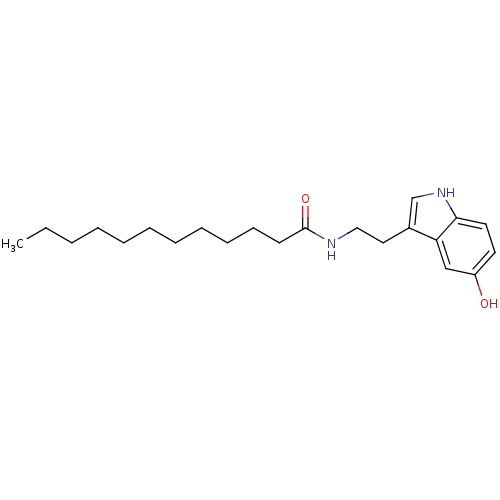
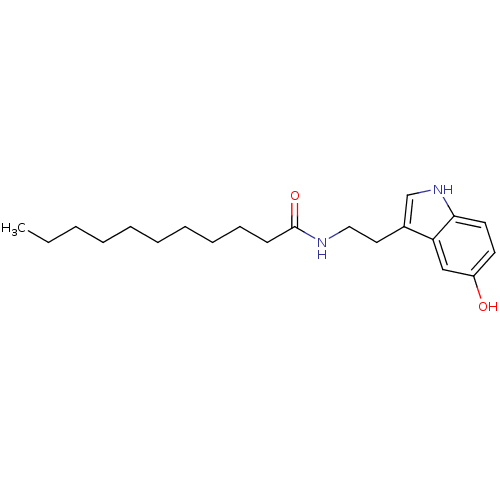
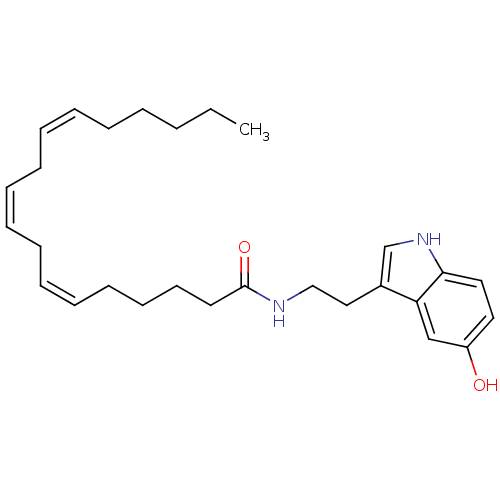
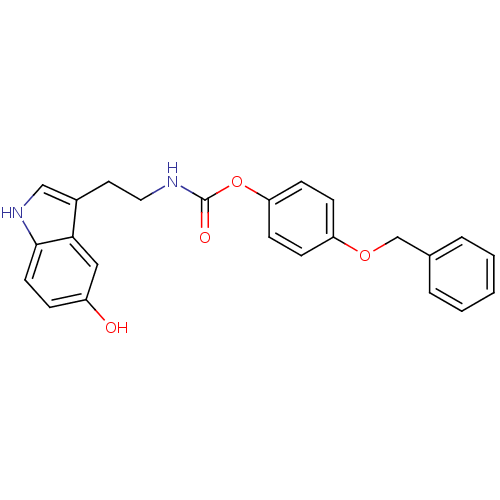
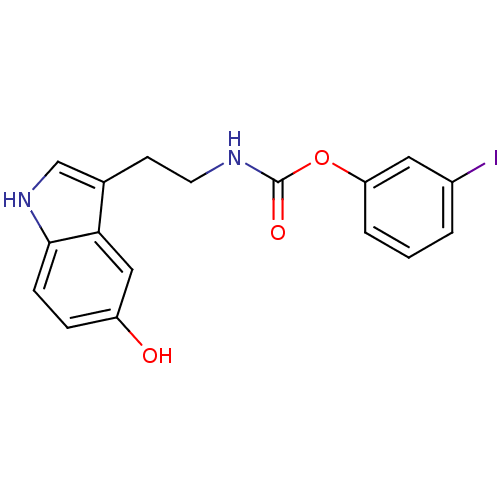
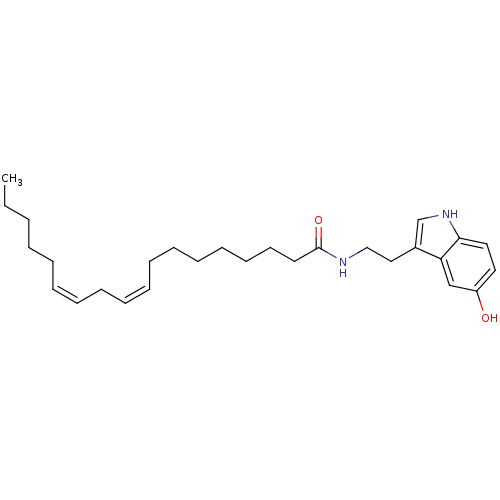
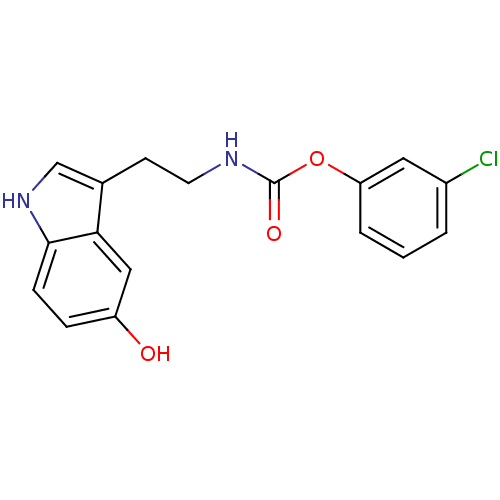
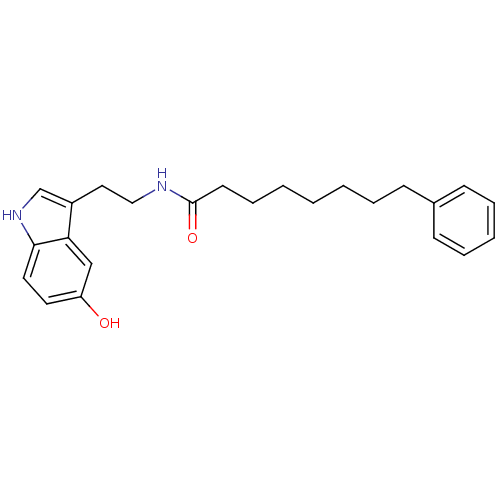
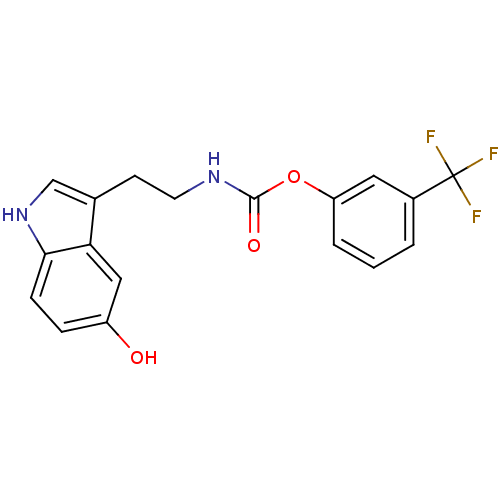
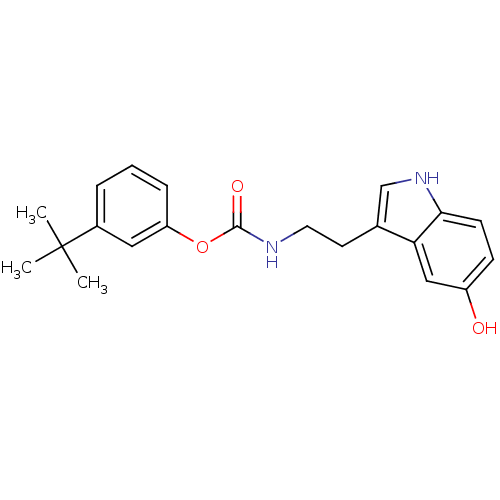
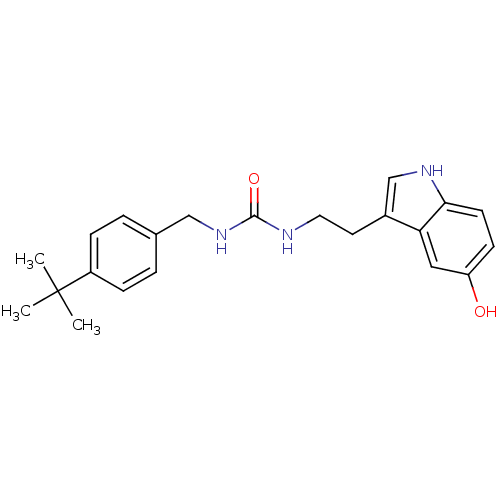
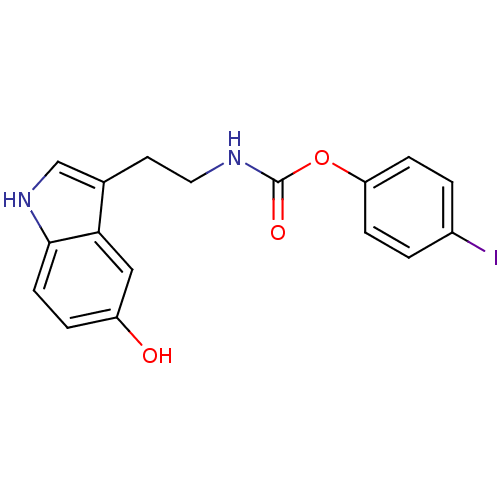
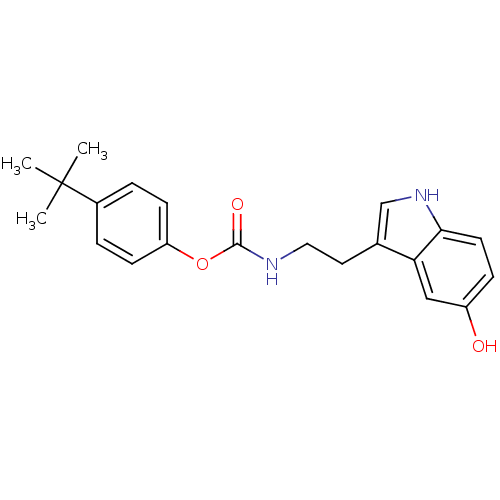
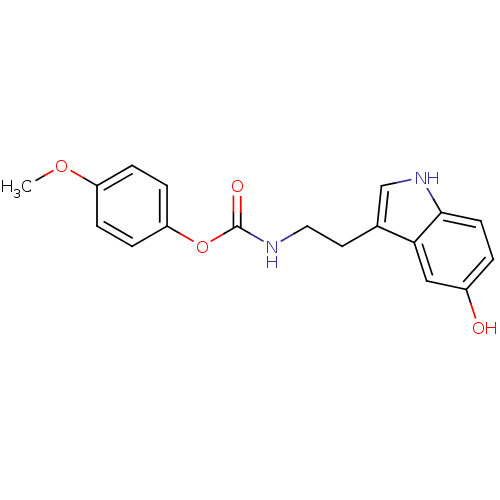
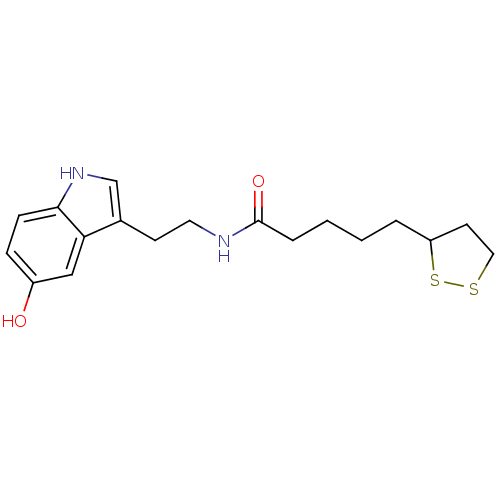
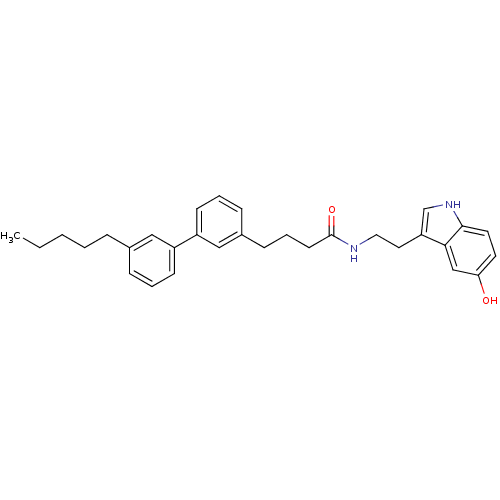
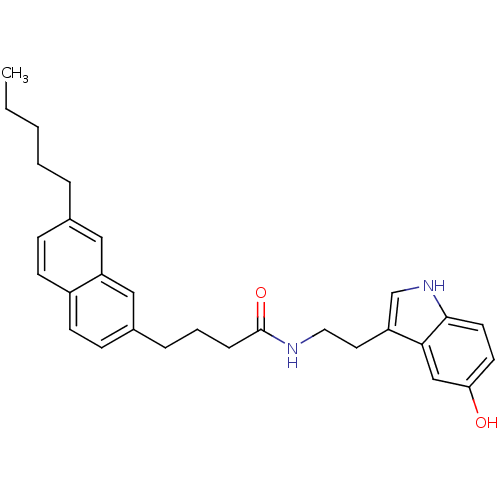
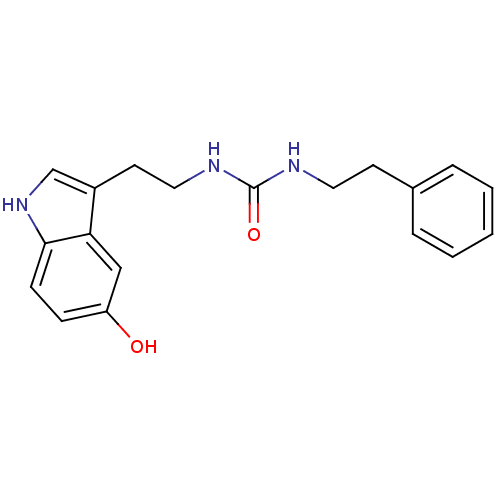
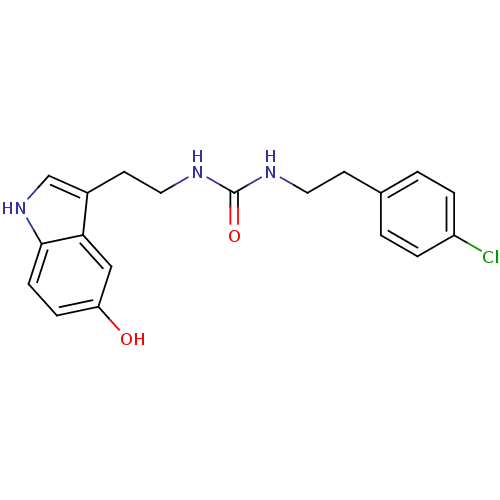
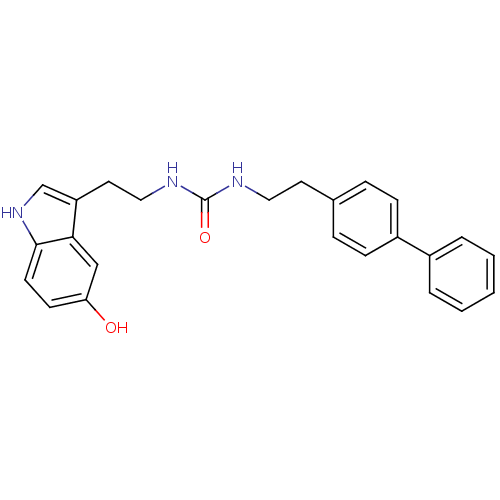
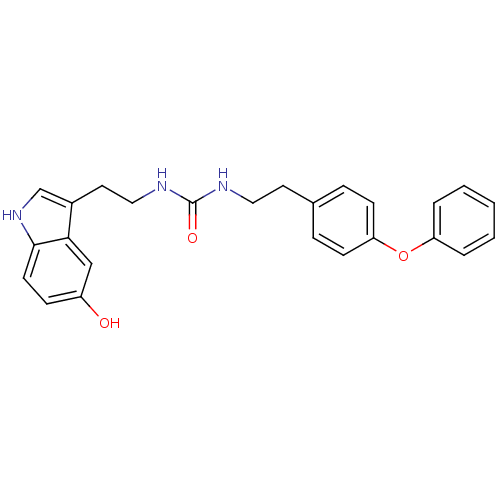
Report error Found 78 Enz. Inhib. hit(s) with all data for entry = 2657
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 240nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 270nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 430nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 470nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 9.0 T: 2°CAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 680nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 720nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 740nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 760nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 950nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.82E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 2.13E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 2.40E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 2.57E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 2.63E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 2.82E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMpH: 9.0 T: 2°CAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMpH: 9.0 T: 2°CAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 3.80E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 5.25E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 6.40E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 6.80E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMpH: 9.0 T: 2°CAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 9.12E+3nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 9.0 T: 2°CAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 9.0 T: 2°CAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMT: 2°CAssay Description:The antagonistic effects of test compounds were evaluated using HEK293 cells stably overexpressing human TRPV1. Experiments were carried out by measu...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 9.0 T: 2°CAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Human)
Sapienza University of Rome
Sapienza University of Rome
Affinity DataIC50: 1.00E+4nMAssay Description:The effect of the test compounds on the enzymatic hydrolysis of [14C]anandamide was evaluated by using membranes prepared from rat brain. [14C]Ethano...More data for this Ligand-Target Pair