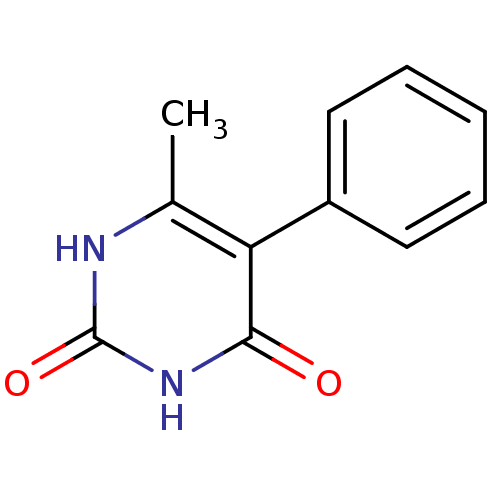
Report error Found 26 Enz. Inhib. hit(s) with all data for entry = 2401
Affinity DataKi: 1.30nM ΔG°: -52.8kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -39.8kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 280nM ΔG°: -38.9kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 400nM ΔG°: -38.0kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 410nM ΔG°: -37.9kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 420nM ΔG°: -37.9kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 490nM ΔG°: -37.5kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 500nM ΔG°: -37.4kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 630nM ΔG°: -36.8kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 710nM ΔG°: -36.5kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 1.03E+3nM ΔG°: -35.5kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 1.06E+3nM ΔG°: -35.5kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nM ΔG°: -35.2kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 1.21E+3nM ΔG°: -35.1kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 1.74E+3nM ΔG°: -34.2kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 1.83E+3nM ΔG°: -34.1kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 3.01E+3nM ΔG°: -32.8kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 3.34E+3nM ΔG°: -32.5kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 3.67E+3nM ΔG°: -32.3kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 4.59E+3nM ΔG°: -31.7kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 4.65E+3nM ΔG°: -31.7kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: 5.81E+3nM ΔG°: -31.1kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nM ΔG°: >-27.9kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nM ΔG°: >-27.9kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nM ΔG°: >-27.9kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nM ΔG°: >-27.9kJ/molepH: 6.4 T: 2°CAssay Description:The enzyme reaction was started by the addition of enzyme to the reaction mixture containing substrate and test compounds. The reaction was stopped b...More data for this Ligand-Target Pair
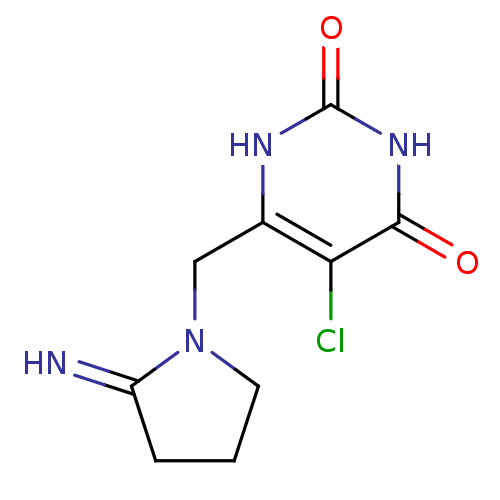
 3D Structure (crystal)
3D Structure (crystal)