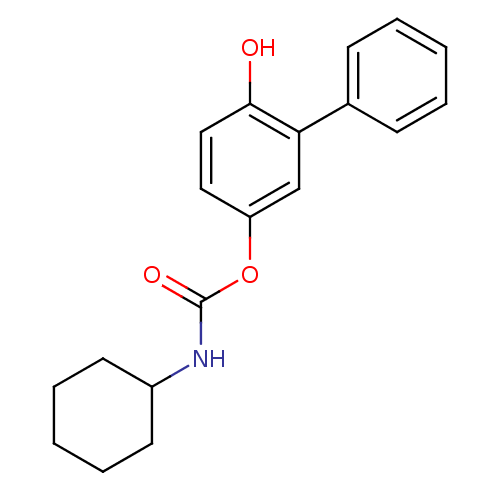
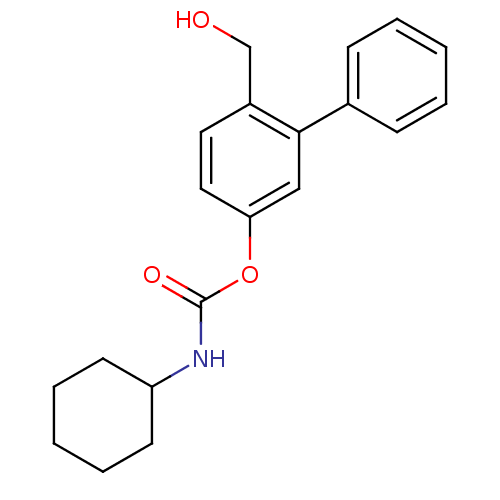
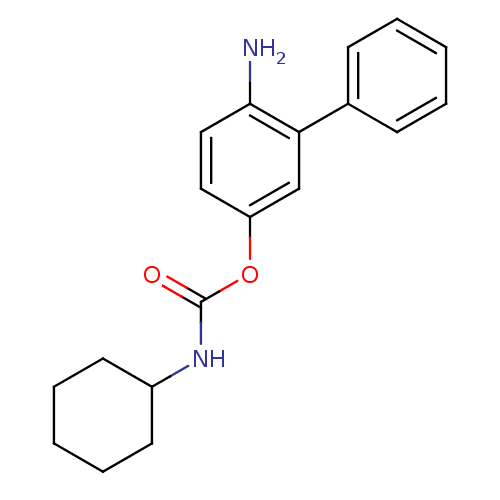
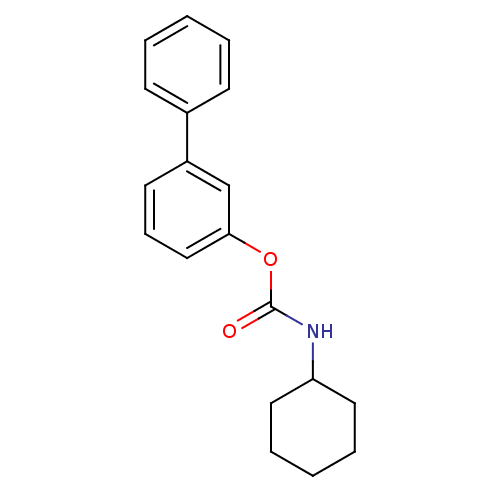
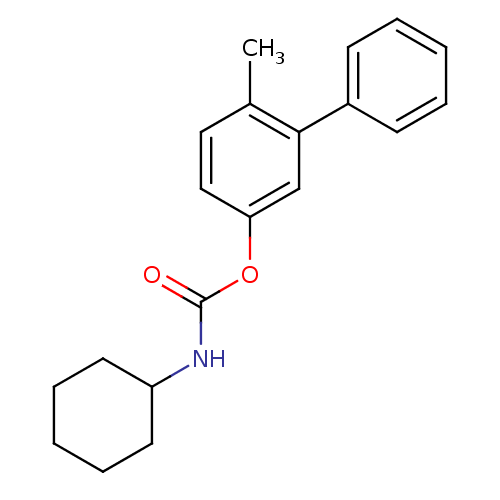
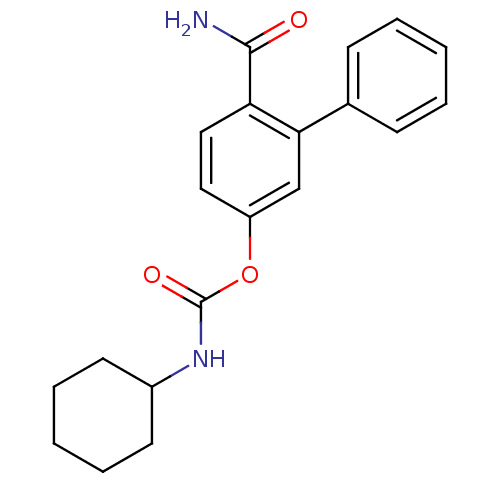
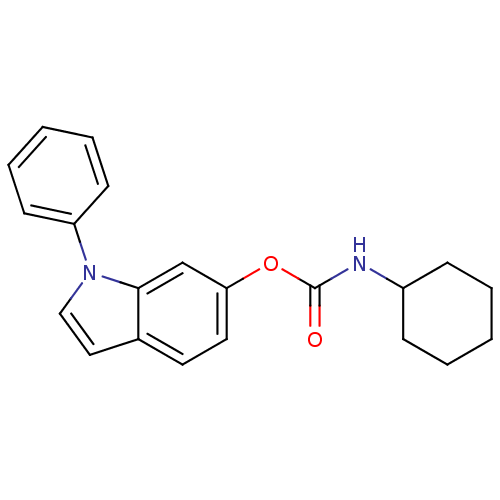
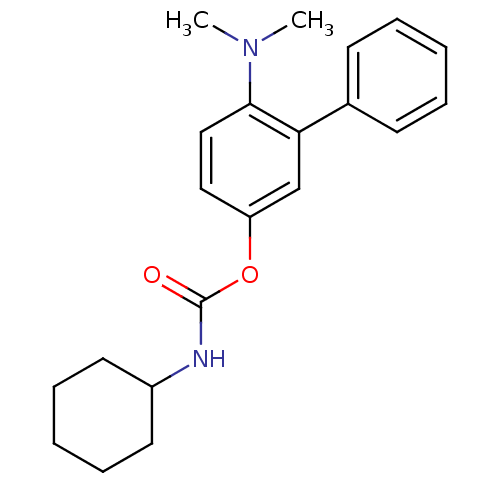
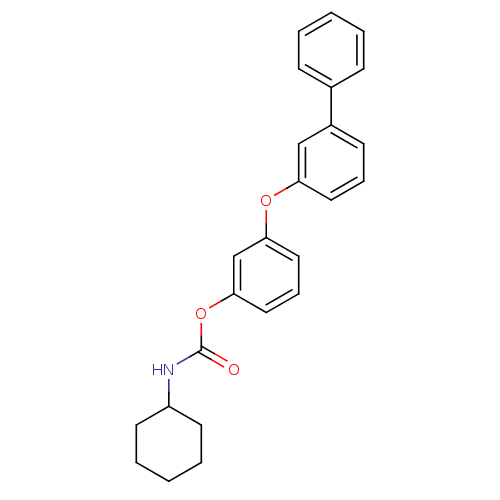
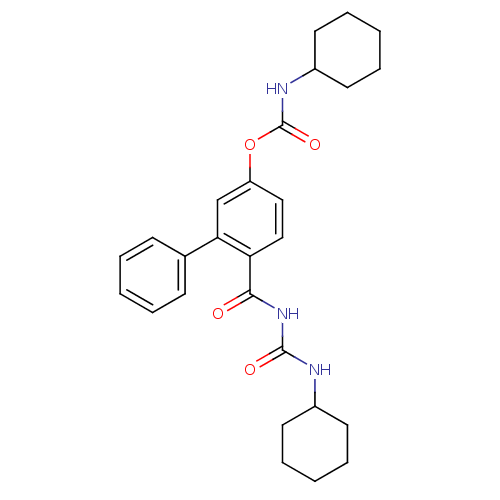
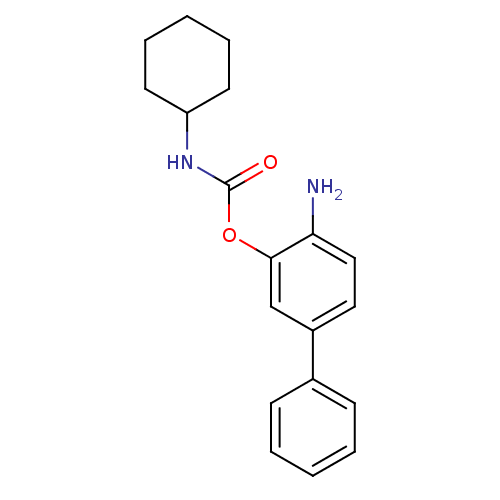
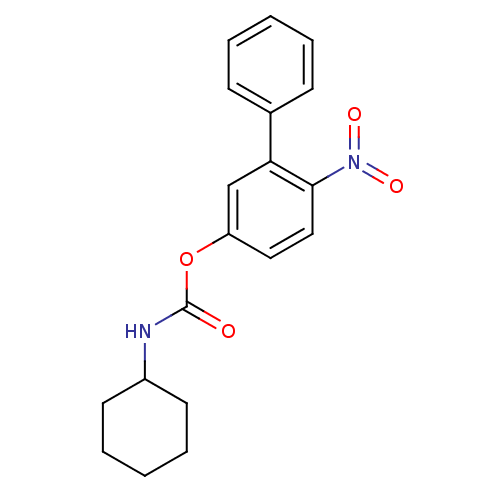
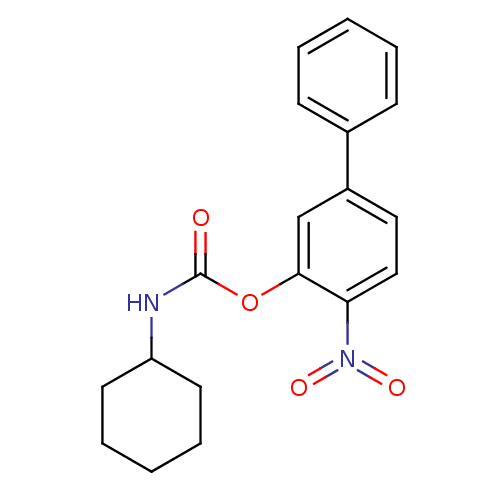
Report error Found 13 Enz. Inhib. hit(s) with all data for entry = 5849
Affinity DataIC50: 45nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 176nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 252nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+3nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 2.74E+3nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 2.74E+3nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 4.83E+3nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Membrane fractions were prepared from Wistar rat brain homogenates, and FAAH activity were assayed by using [3H]anandamide (anandamide[ethanolamine-3...More data for this Ligand-Target Pair