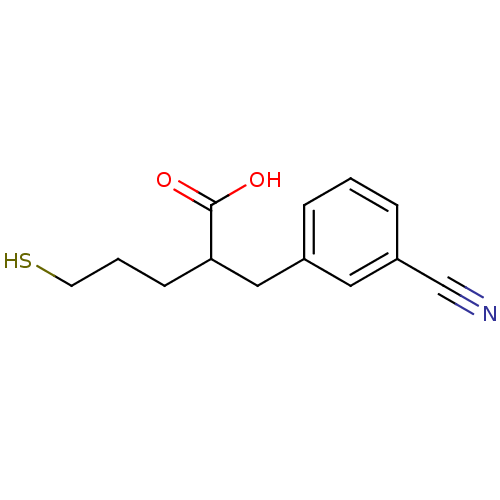
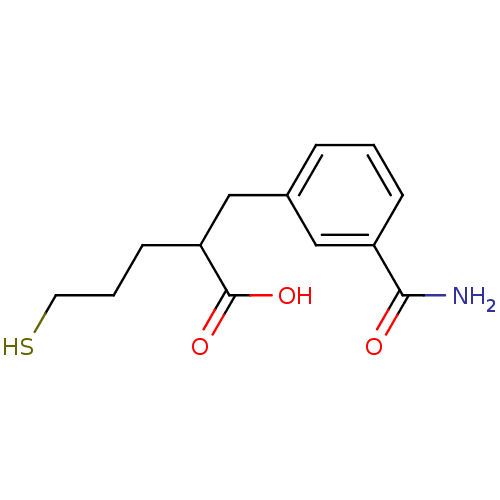
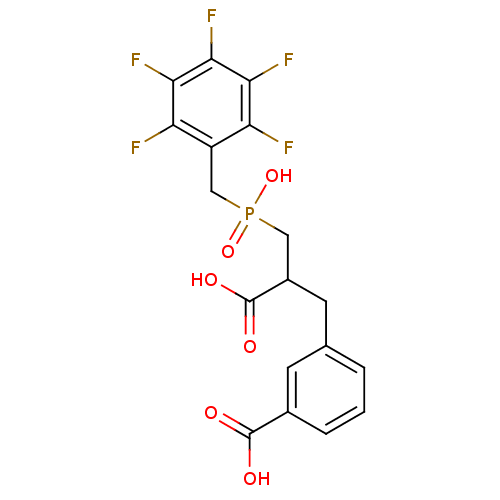
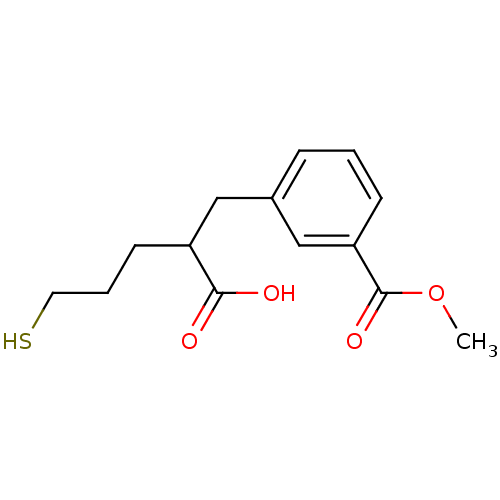
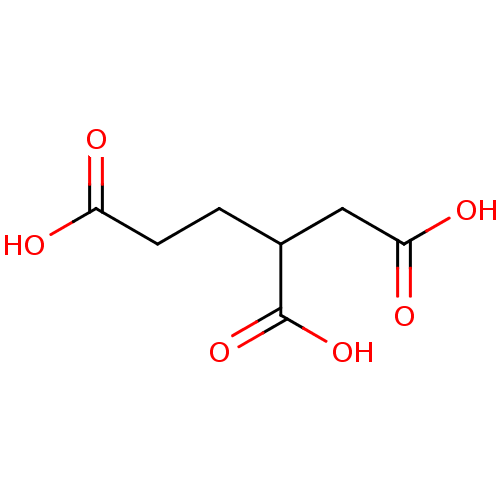
Report error Found 26 Enz. Inhib. hit(s) with all data for entry = 2176
Affinity DataIC50: 0.300nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 63nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 82nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 390nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 440nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 640nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 730nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 7.4 T: 2°CAssay Description:GCPII activity in vitro is monitored through the hydrolysis [3H]NAAG to NAA and [3H]Glu. The radioactivity-based assay was miniaturized to a 96-well ...More data for this Ligand-Target Pair
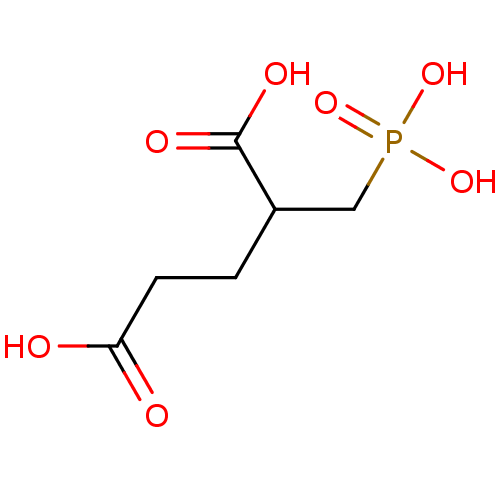
 3D Structure (crystal)
3D Structure (crystal)