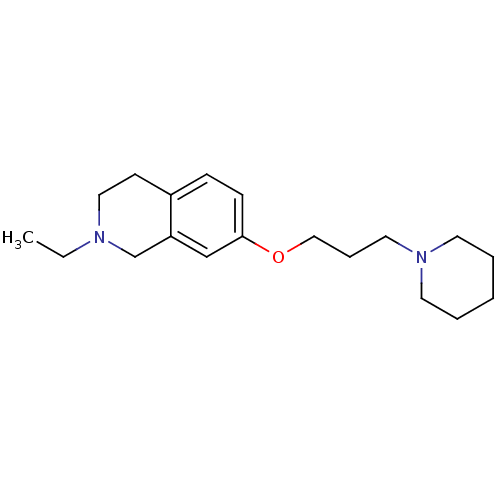
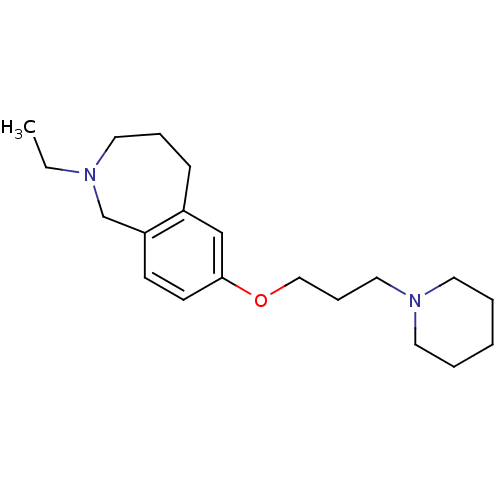
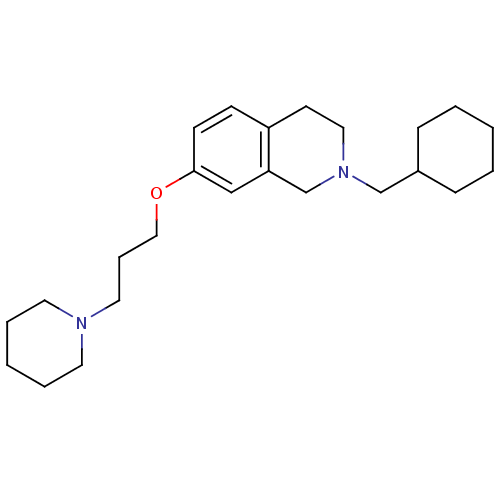
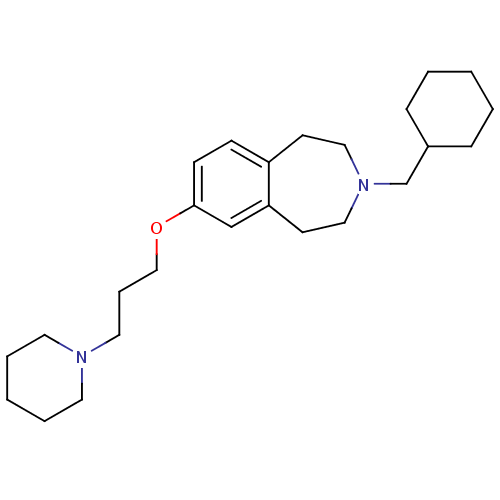
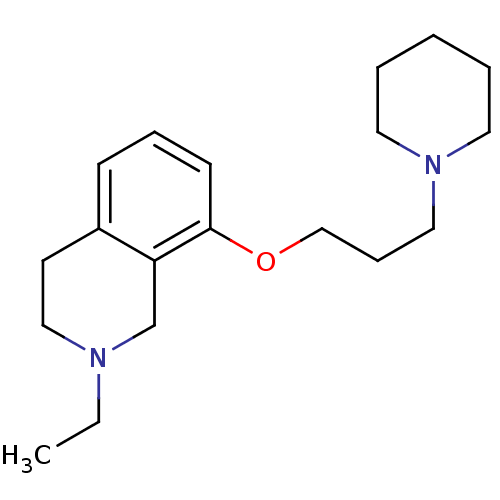
Report error Found 67 Enz. Inhib. hit(s) with all data for entry = 50017774
Affinity DataKi: 0.170nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.340nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.340nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.350nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.460nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.530nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.530nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.540nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.560nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.640nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.720nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.890nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.970nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.01nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.29nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4.90nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.30nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.40nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 8.10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 11.6nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 12.1nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 15.4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 17.1nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 19.4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 22.6nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 23.8nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 32.5nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 36.7nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 38.1nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 39.6nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 42.3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 48.8nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 54nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 59nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 70.1nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 74.5nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from rat cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 84.2nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 95.2nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from human cloned H3 receptorMore data for this Ligand-Target Pair