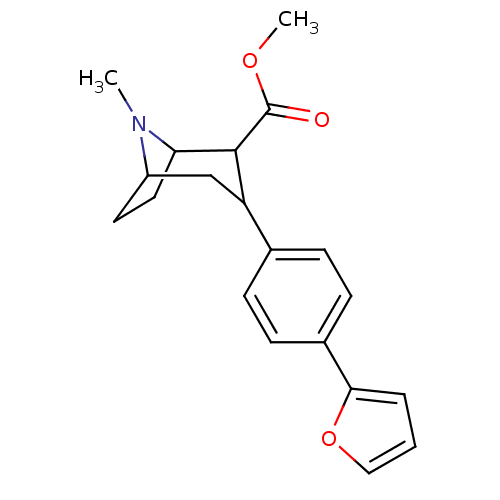
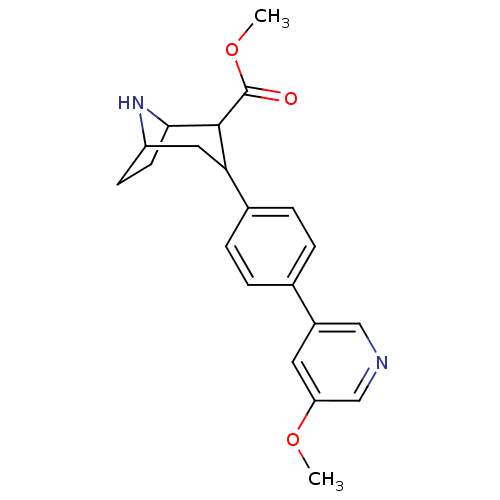
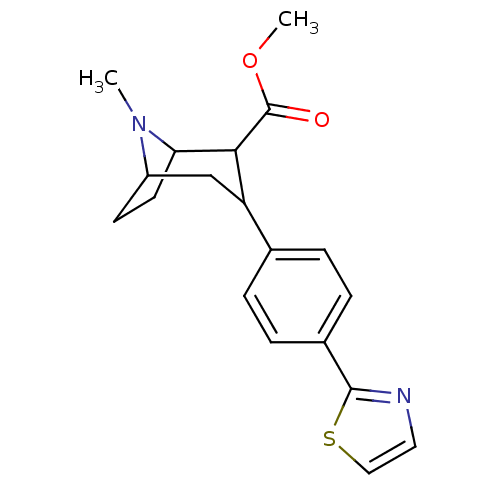
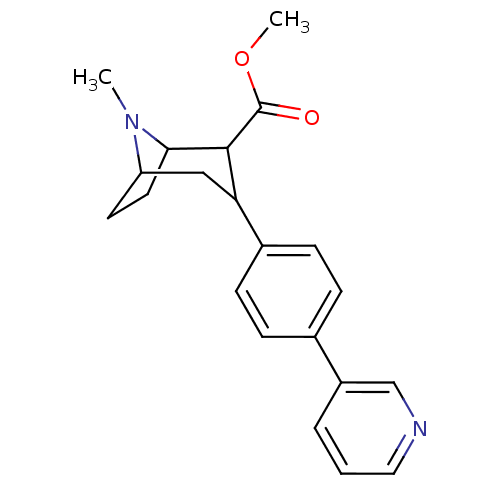
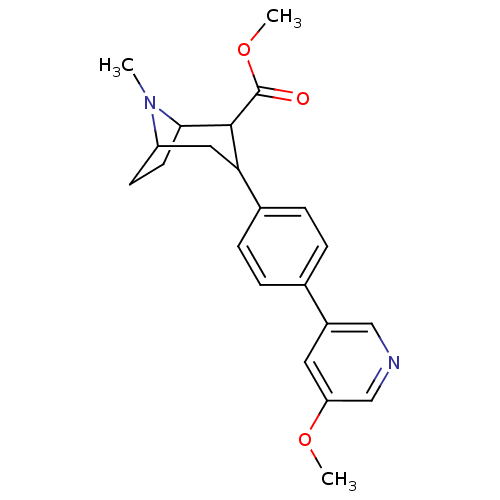
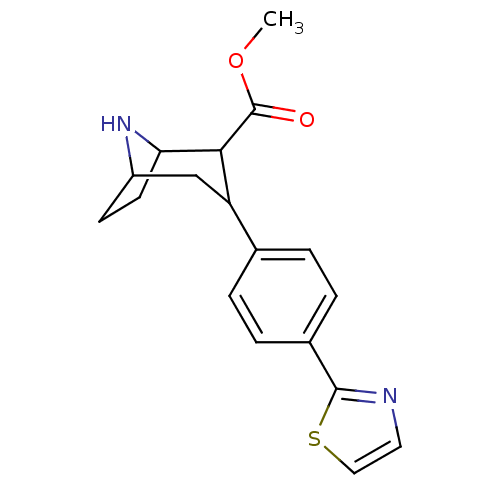
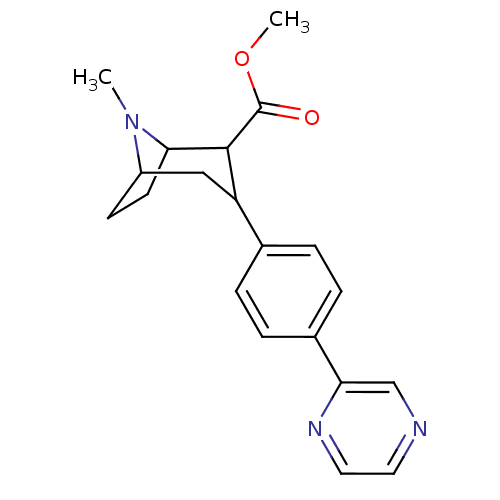
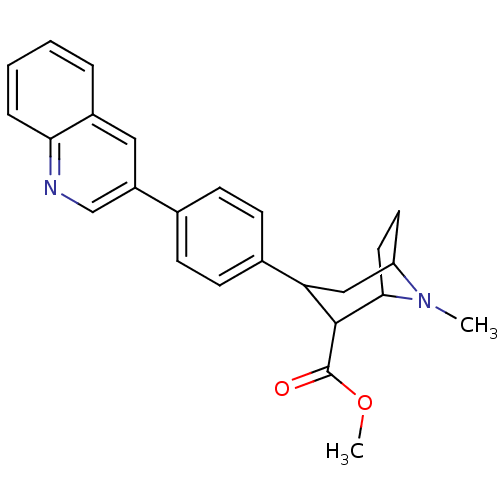
Report error Found 48 Enz. Inhib. hit(s) with all data for entry = 50016945
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.350nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.410nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.13nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.56nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.67nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 3.51nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 3.76nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 3.91nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 5.15nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 7.14nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 11.3nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 12.7nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 14.2nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 25.2nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 26.1nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 31.7nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 33.1nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 44nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 59.8nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 69.6nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 109nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 121nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 127nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 161nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 213nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 213nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 249nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 332nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 344nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 520nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 545nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 587nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 739nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [3H]paroxetine from SERT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 1.41E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: 1.57E+3nMAssay Description:Displacement of [3H]GBR-12935 from DAT in rat frontoparietal cerebral cortexMore data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nMAssay Description:Displacement of [3H]nisoxetine from NET in rat caudate-putamenMore data for this Ligand-Target Pair