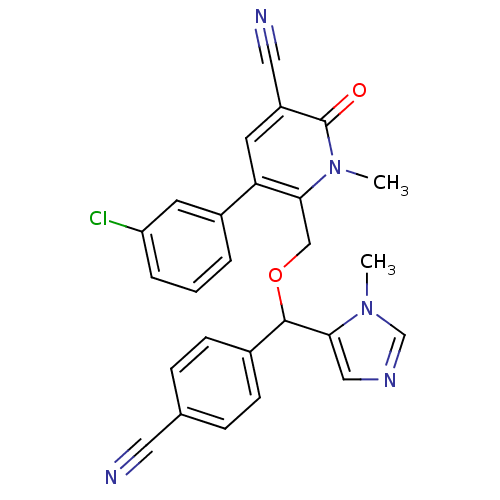
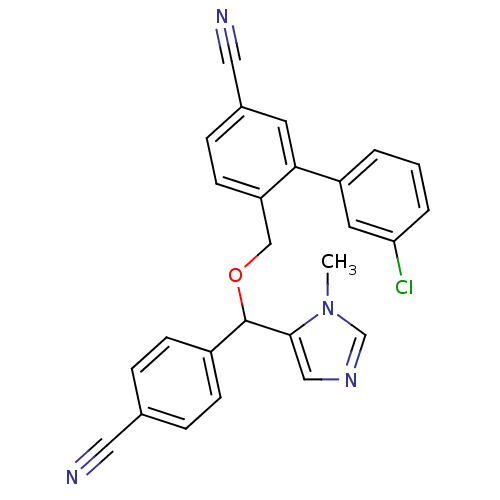
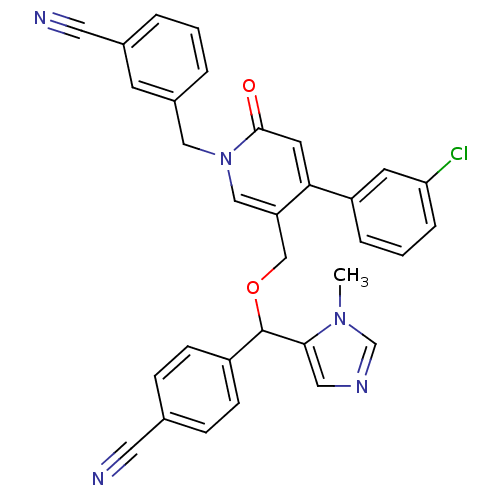
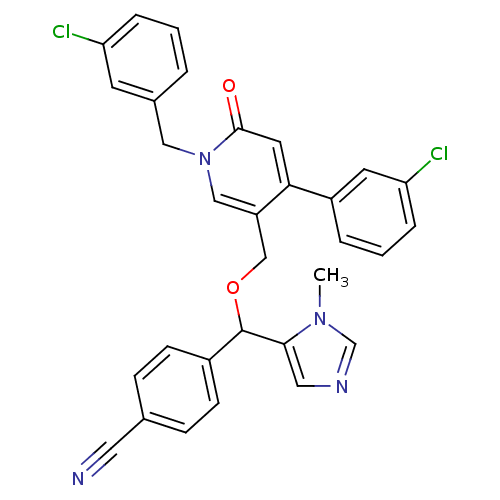
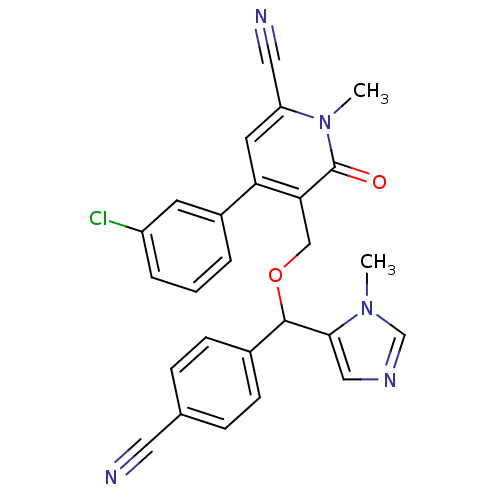
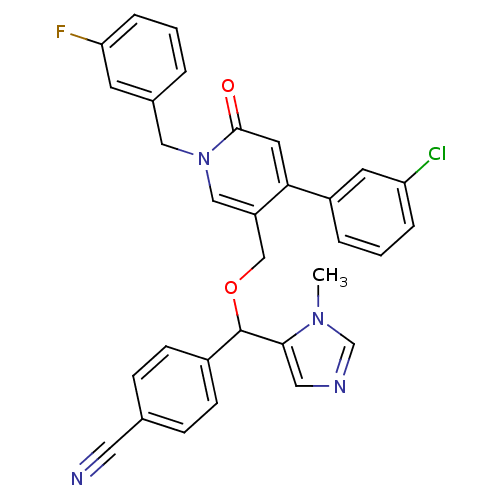
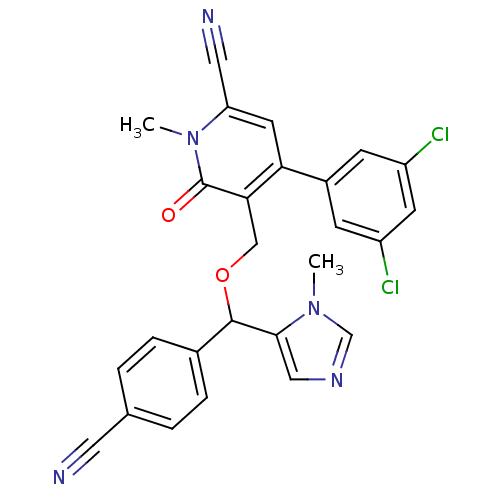
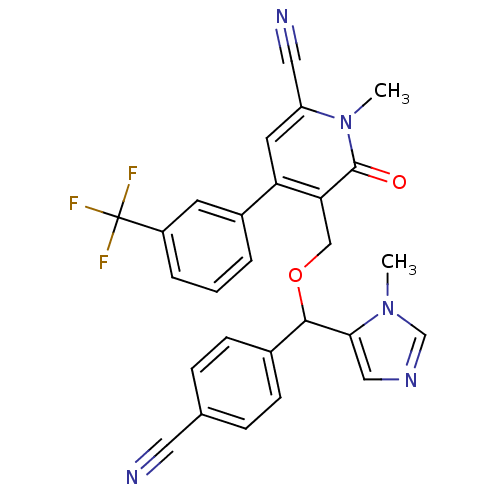
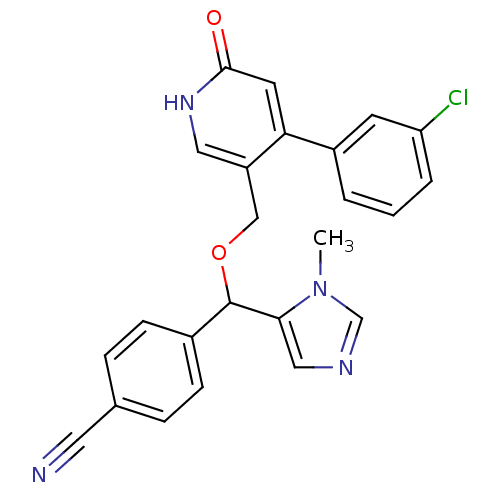
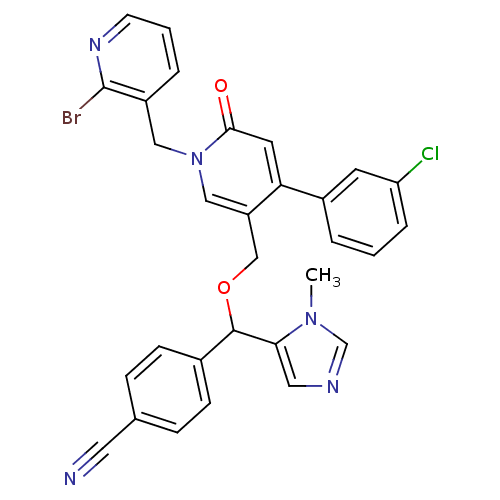
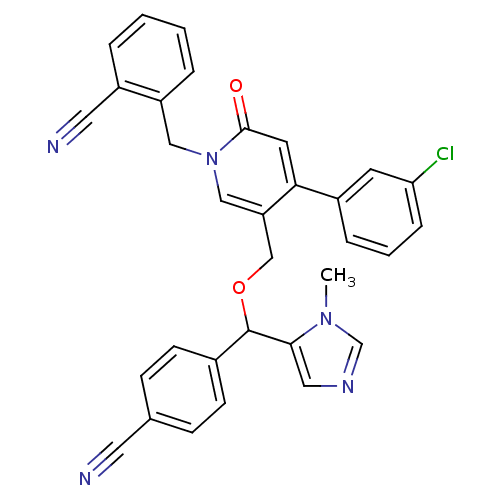
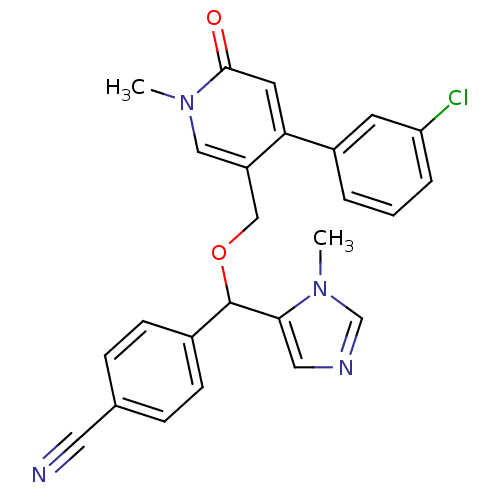
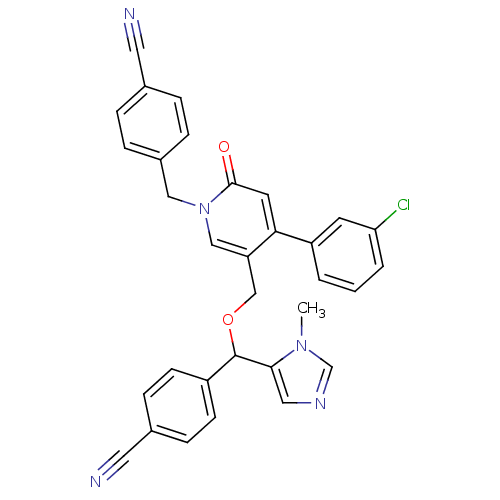
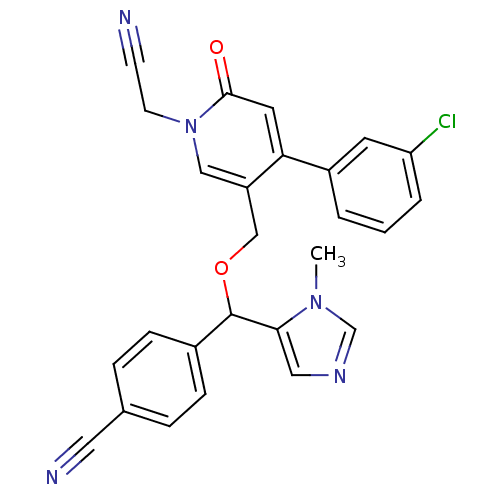
Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 50015107
TargetProtein farnesyltransferase subunit beta/geranylgeranyltransferase type-1 subunit alpha(Mouse)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 0.5nMAssay Description:Reduced farnesylation of H-ras transformed NIH3T3 cellsMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.30nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 2.70nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetProtein farnesyltransferase subunit beta/geranylgeranyltransferase type-1 subunit alpha(Mouse)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 5.20nMAssay Description:Reduced farnesylation of H-ras transformed NIH3T3 cellsMore data for this Ligand-Target Pair
TargetProtein farnesyltransferase subunit beta/geranylgeranyltransferase type-1 subunit alpha(Mouse)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 9nMAssay Description:Reduced farnesylation of H-ras transformed NIH3T3 cellsMore data for this Ligand-Target Pair
TargetProtein farnesyltransferase subunit beta/geranylgeranyltransferase type-1 subunit alpha(Mouse)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 10nMAssay Description:Reduced farnesylation of H-ras transformed NIH3T3 cellsMore data for this Ligand-Target Pair
TargetProtein farnesyltransferase subunit beta/geranylgeranyltransferase type-1 subunit alpha(Mouse)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 11nMAssay Description:Reduced farnesylation of H-ras transformed NIH3T3 cellsMore data for this Ligand-Target Pair
TargetProtein farnesyltransferase subunit beta/geranylgeranyltransferase type-1 subunit alpha(Mouse)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 15nMAssay Description:Reduced farnesylation of H-ras transformed NIH3T3 cellsMore data for this Ligand-Target Pair
TargetProtein farnesyltransferase subunit beta/geranylgeranyltransferase type-1 subunit alpha(Mouse)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataEC50: 15nMAssay Description:Reduced farnesylation of H-ras transformed NIH3T3 cellsMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.20E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.90E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 2.00E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 3.60E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 6.20E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 6.60E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 7.70E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 9.40E+3nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of [3H]FPP incorporation into biotin-linked K-ras decapeptide (CVLL) by bovine geranylgeranyltransferaseMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair
TargetGeranylgeranyl transferase type-1 subunit beta/Protein farnesyltransferase/geranylgeranyltransferase type-1 subunit alpha(Bovine)
Abbott Laboratories
Curated by ChEMBL
Abbott Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:inhibitory concentration needed to to reduce the bovine GGTase-catalyzed incorporation of [3H]-FPP into a biotin-linked K-ras (B) decapeptideMore data for this Ligand-Target Pair