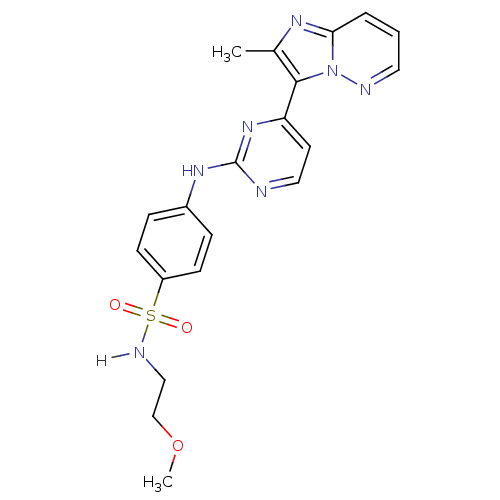
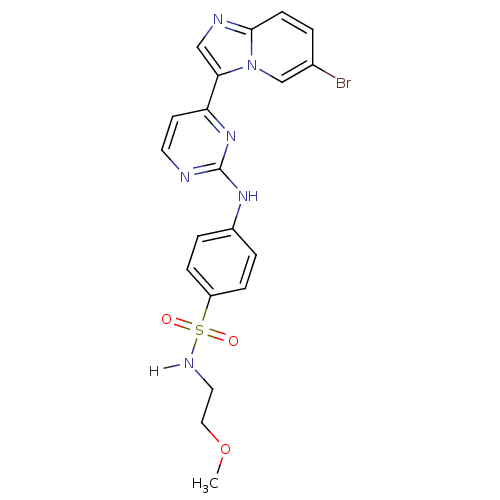
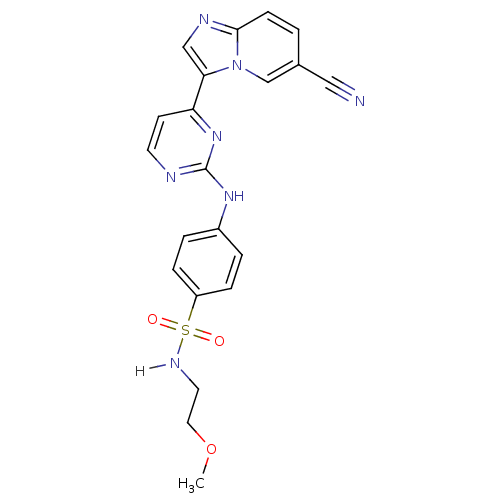
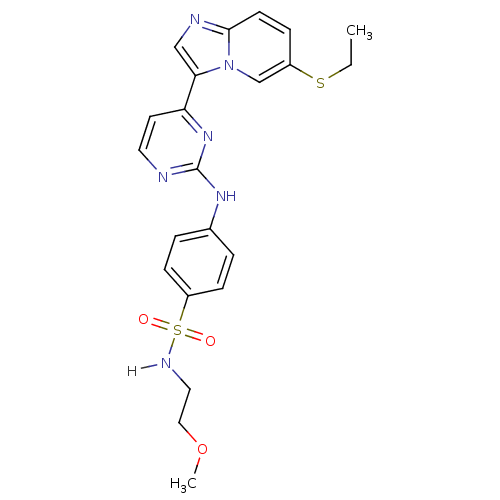
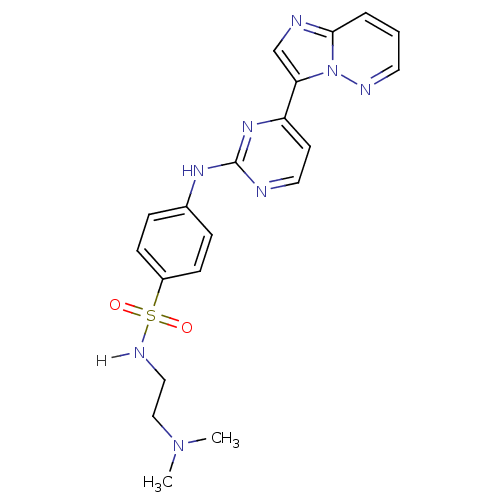
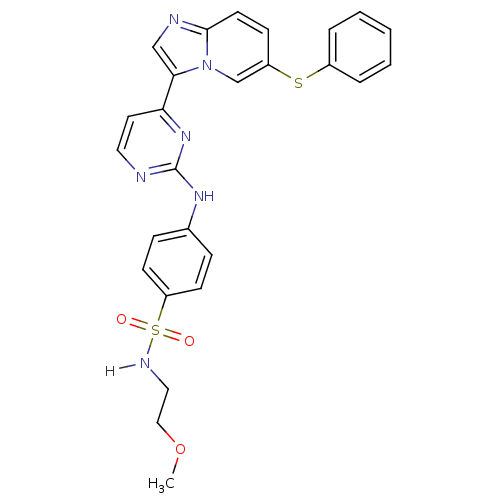
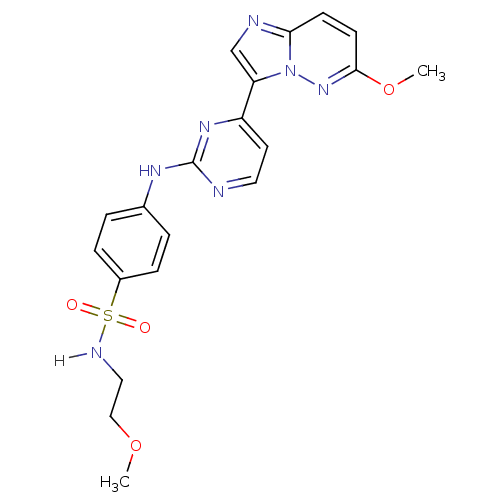
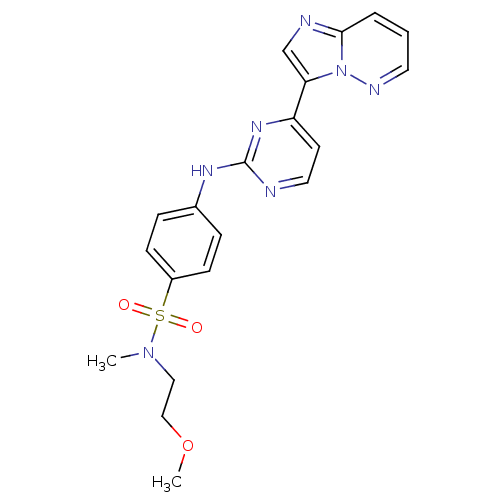
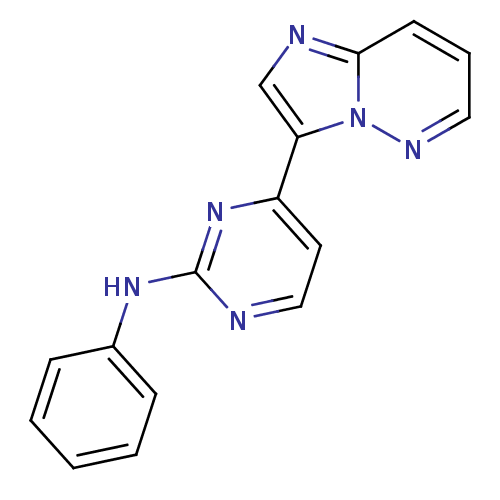
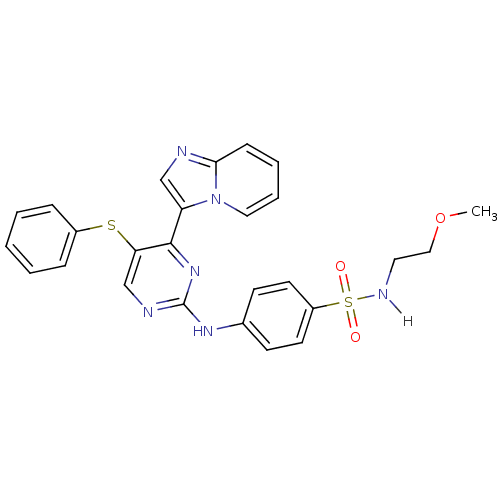
Report error Found 14 Enz. Inhib. hit(s) with all data for entry = 972
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 8nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 600nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 600nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.0 T: 2°CAssay Description:The kinase activity was assayed using an in vitro scintillation proximity assay (SPA) for measuring incorporation of [gamma-33P] ATP into GST-Rb.More data for this Ligand-Target Pair