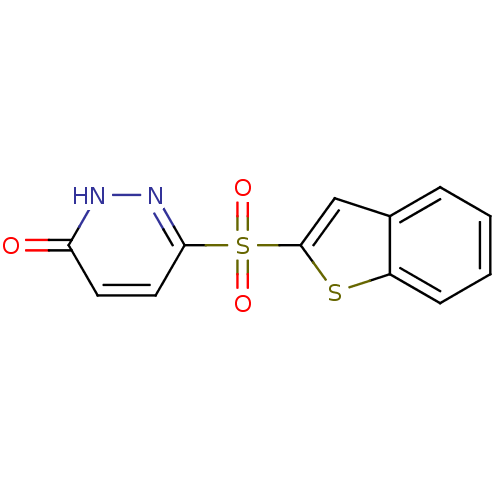
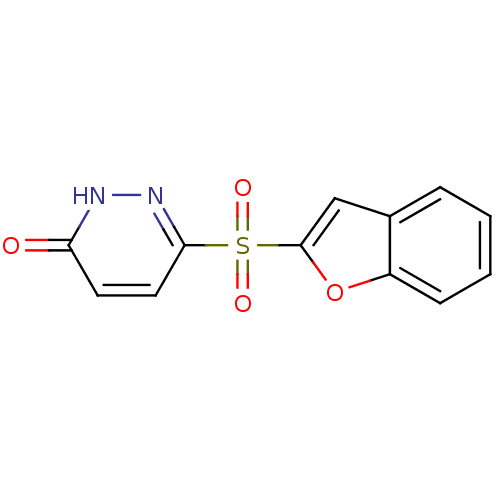
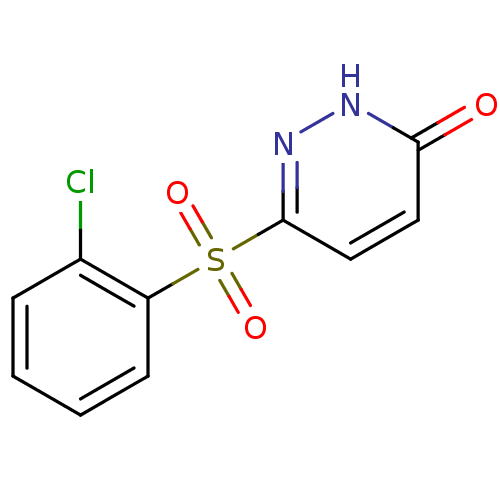
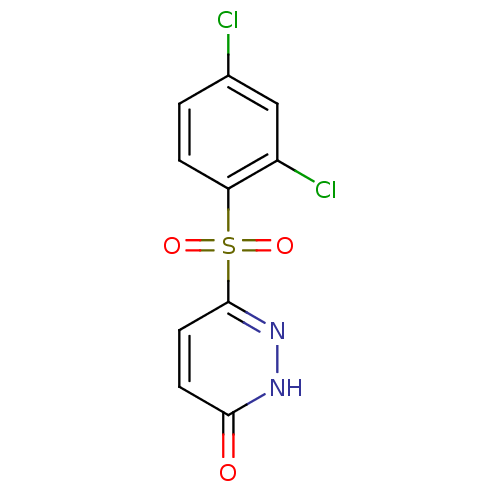
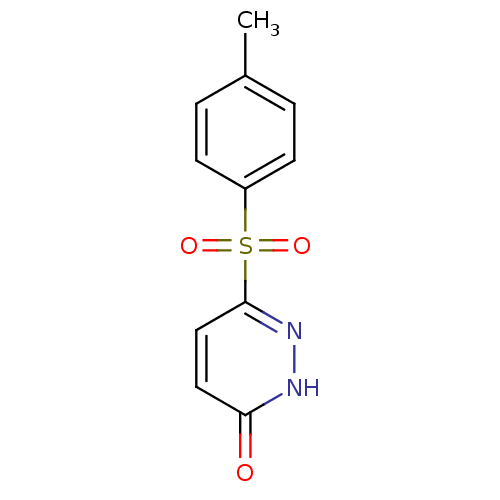
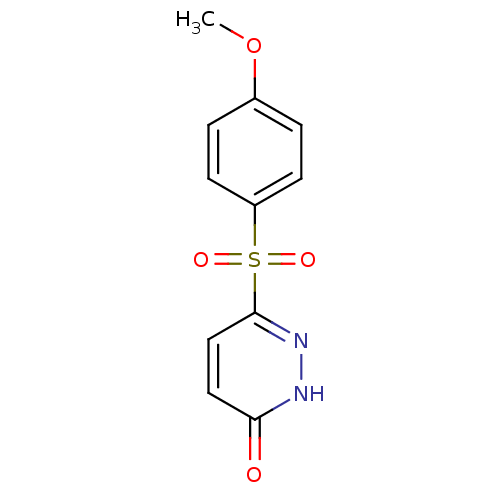
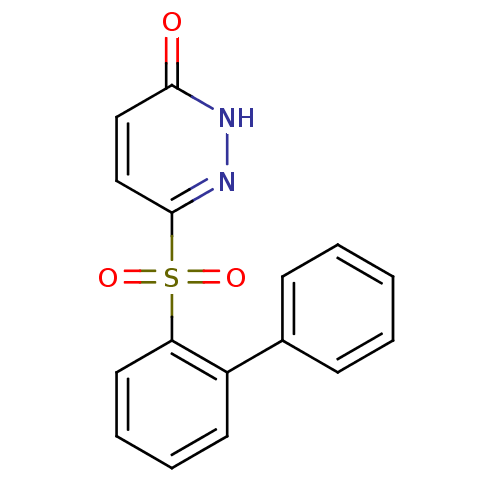
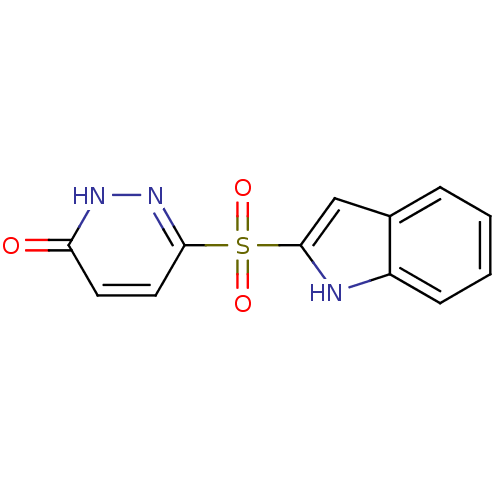
Report error Found 17 Enz. Inhib. hit(s) with all data for entry = 2052
Affinity DataIC50: 0.840nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 280nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 450nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 600nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMpH: 7.0 T: 2°CAssay Description:The activity of the enzyme was determined spectrophotometrically by monitoring the change in absorbance at 340 nm, which is due to the disappearance ...More data for this Ligand-Target Pair
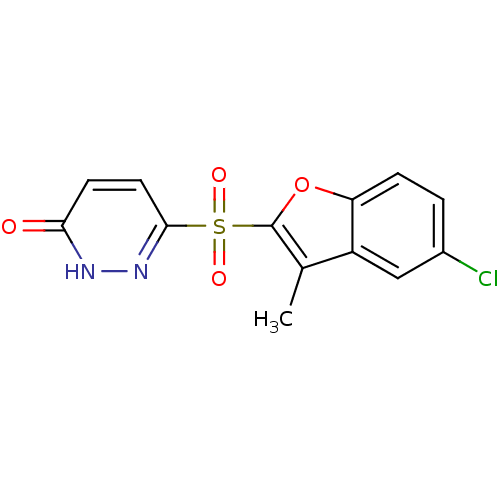
 3D Structure (crystal)
3D Structure (crystal)