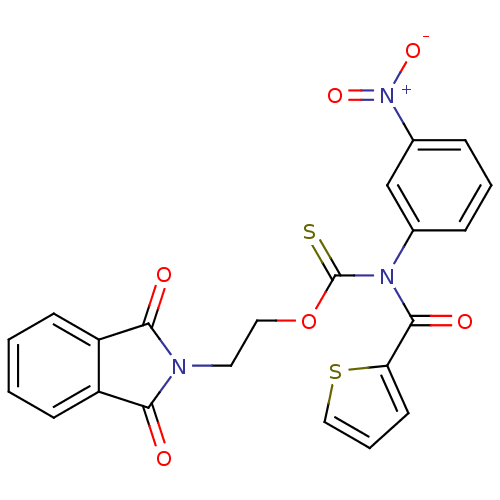
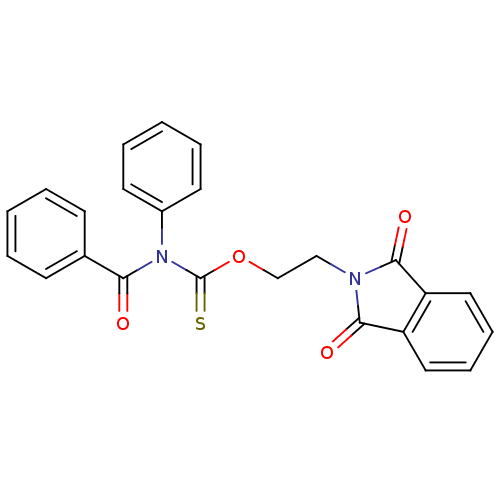
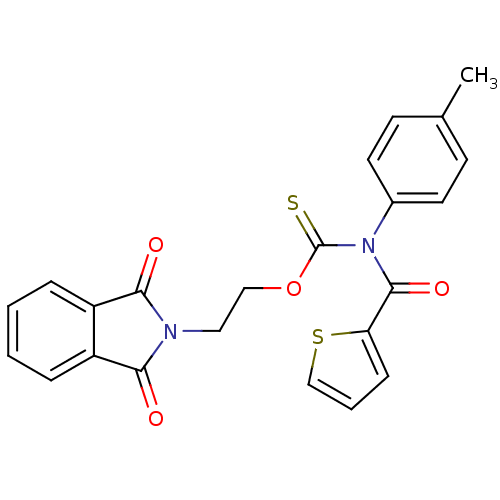
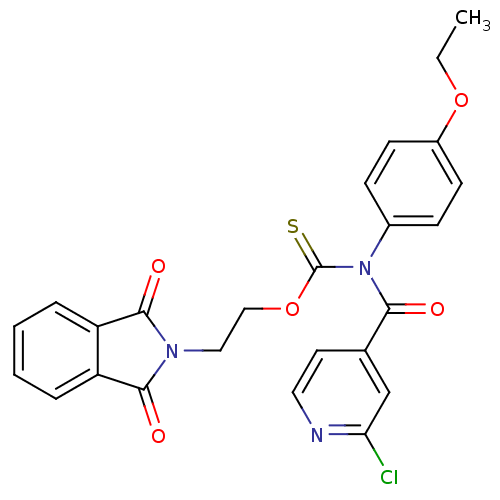
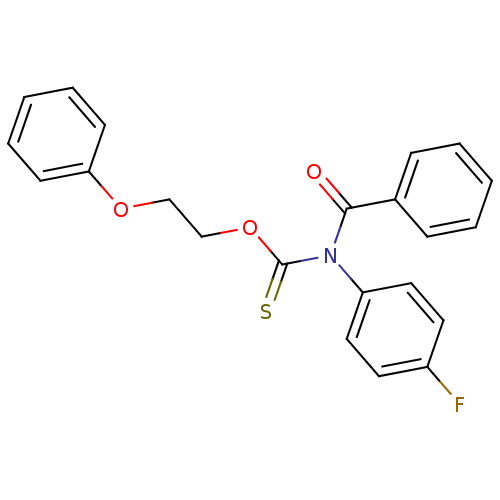
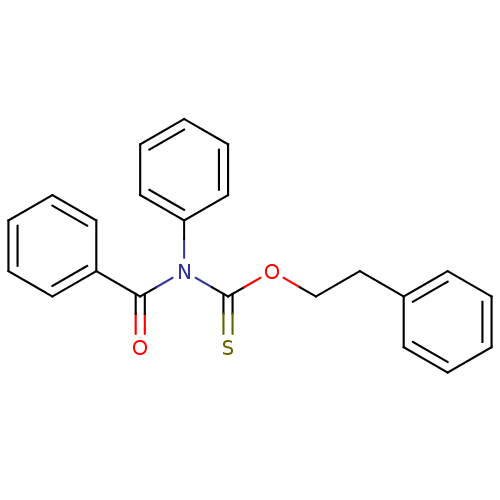
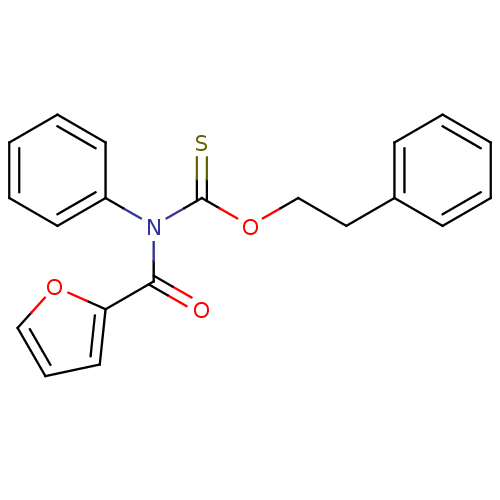
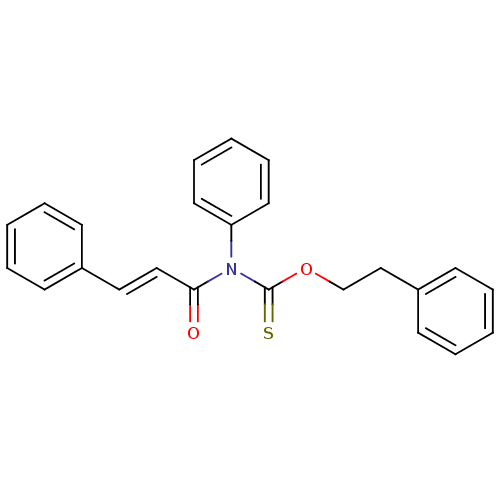
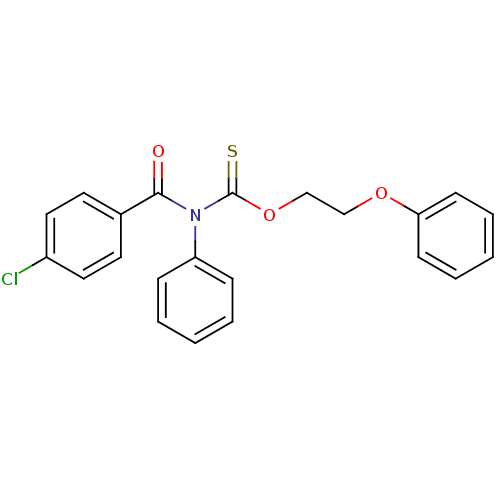
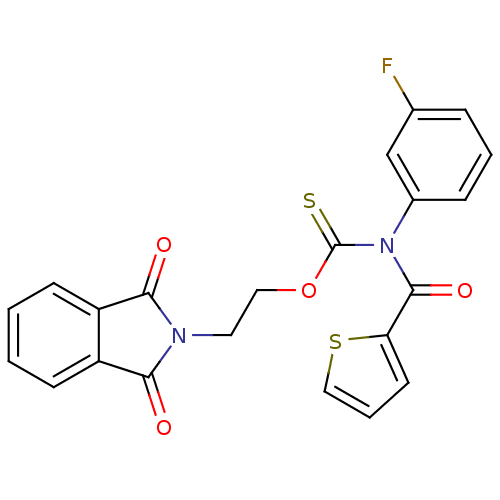
Report error Found 66 Enz. Inhib. hit(s) with all data for entry = 50012889
Affinity DataEC50: 5nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 7nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 8nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 8nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 25nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 35nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 380nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 400nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 400nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 500nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.20E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.30E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.30E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.40E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Activity of selected HIV-1 N-acylthiocarbamates in enzyme assay against Virion-Associated reverse transcriptaseMore data for this Ligand-Target Pair
Affinity DataEC50: 2.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:Activity of selected HIV-1 N-acylthiocarbamates in enzyme assay against Virion-Associated reverse transcriptaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Activity of selected HIV-1 N-acylthiocarbamates in enzyme assay against Virion-Associated reverse transcriptaseMore data for this Ligand-Target Pair
Affinity DataEC50: 3.50E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 3.60E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 4.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 4.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 4.20E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Activity of selected HIV-1 N-acylthiocarbamates in enzyme assay against Virion-Associated reverse transcriptaseMore data for this Ligand-Target Pair
Affinity DataEC50: 4.30E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 4.50E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 6.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 6.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 6.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 6.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:Activity of selected HIV-1 N-acylthiocarbamates in enzyme assay against Virion-Associated reverse transcriptaseMore data for this Ligand-Target Pair
Affinity DataEC50: 6.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 7.60E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 7.70E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 8.00E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 8.40E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 8.60E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 8.80E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 9.50E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 9.60E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 9.90E+3nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+4nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.03E+4nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair
Affinity DataEC50: 1.10E+4nMAssay Description:Concentration required to achieve 50% protection of MT-4 cell from the HIV-1 induced cytopathogenicity was determined by the MTT methodMore data for this Ligand-Target Pair