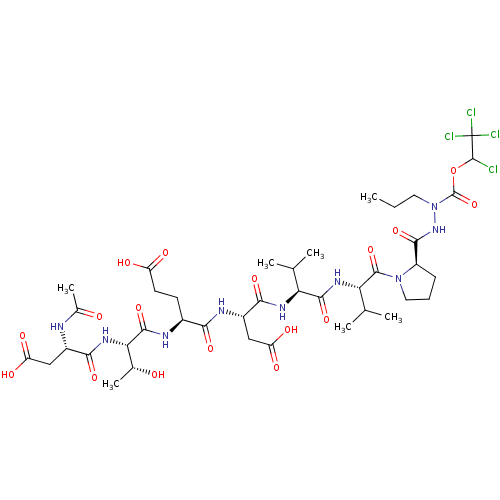
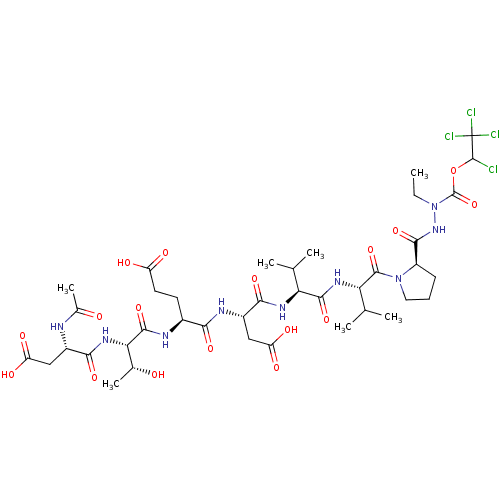
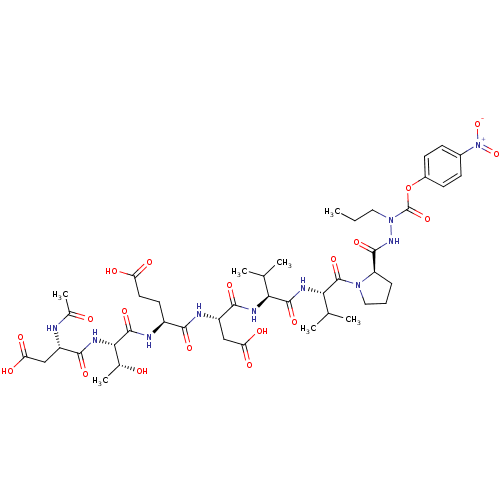
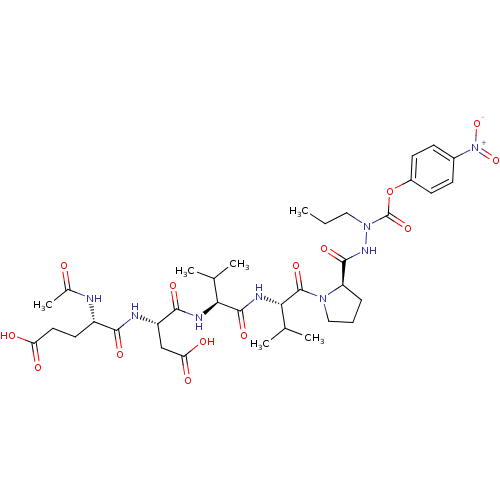
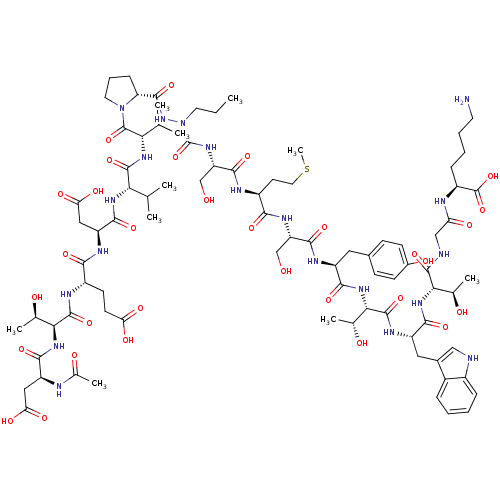
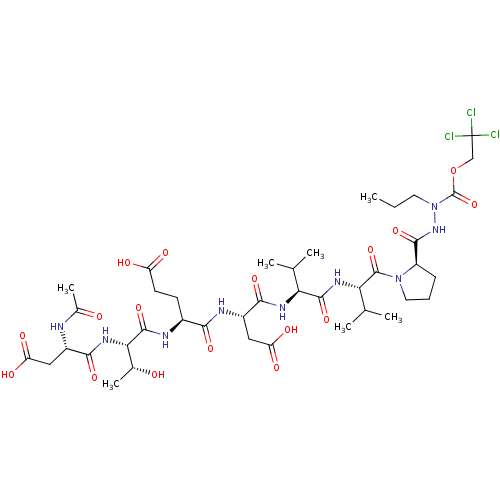
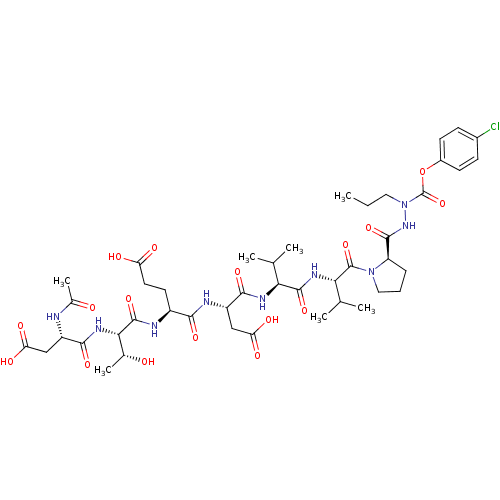
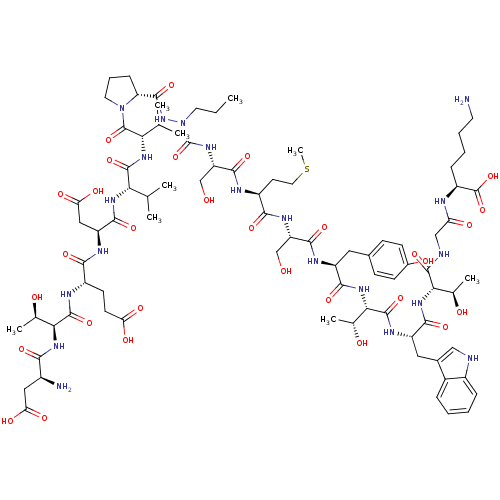
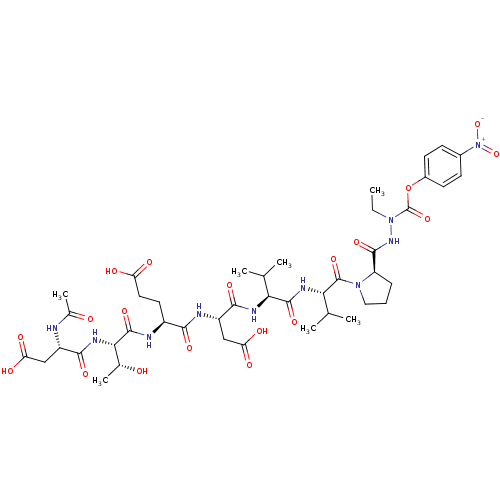
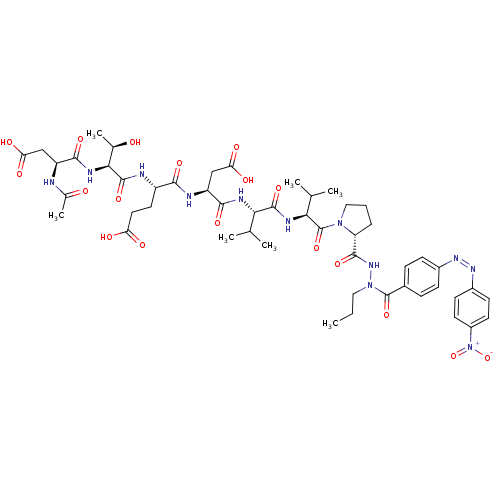
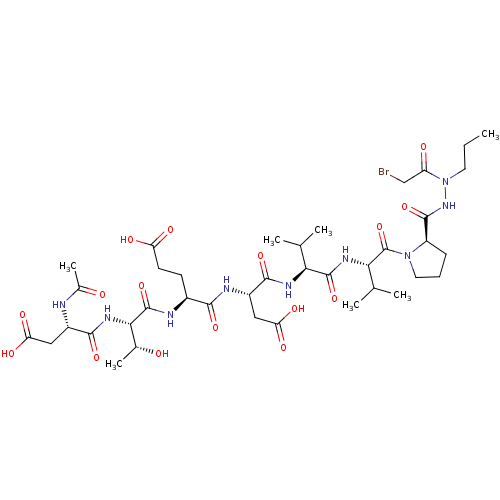
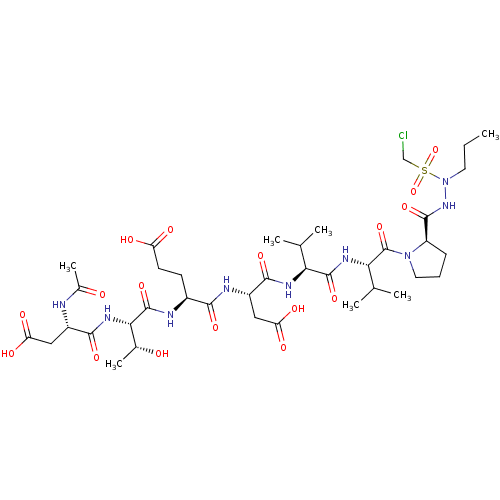
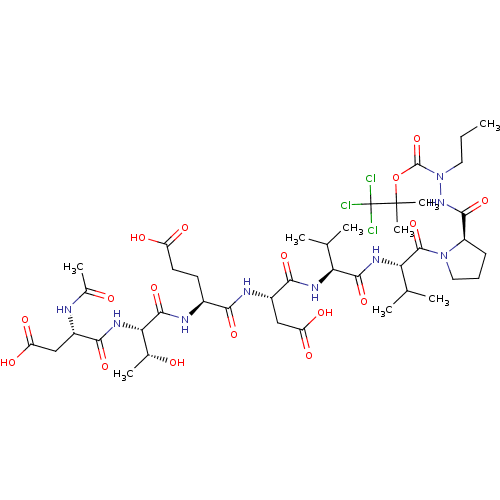
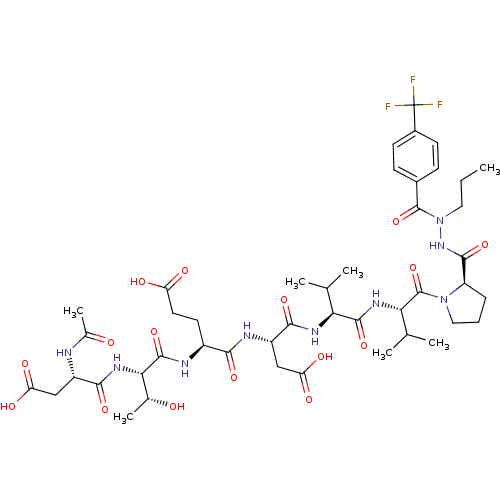
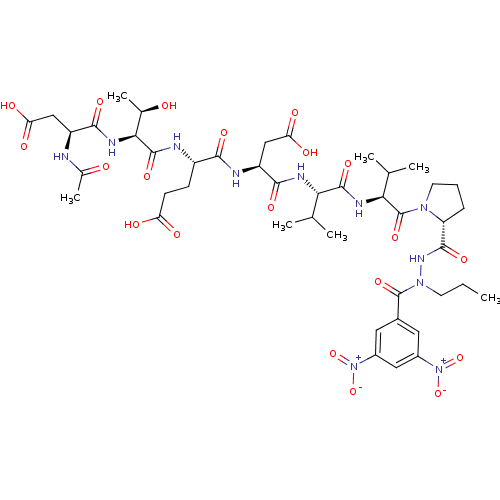
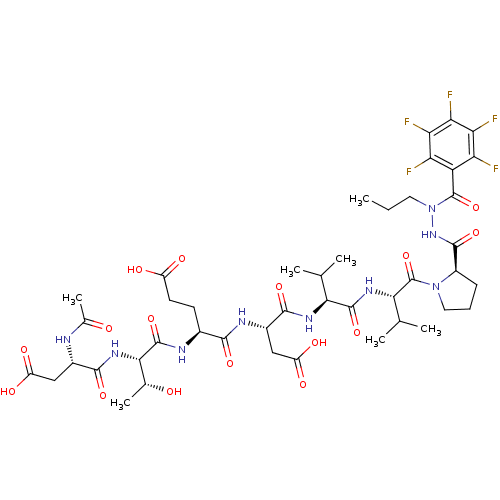
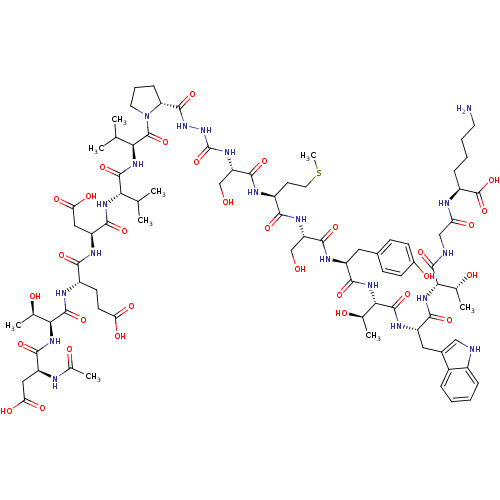
Report error Found 42 Enz. Inhib. hit(s) with all data for entry = 50011831
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
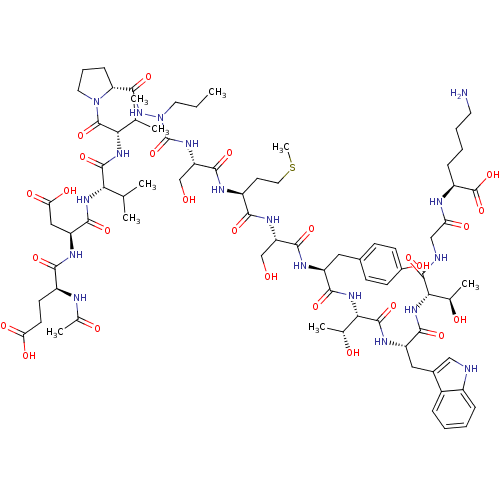
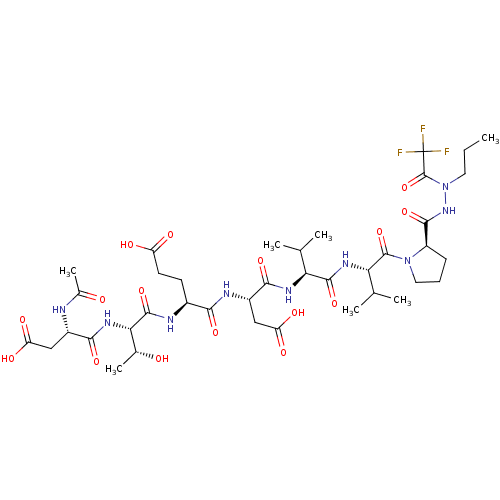
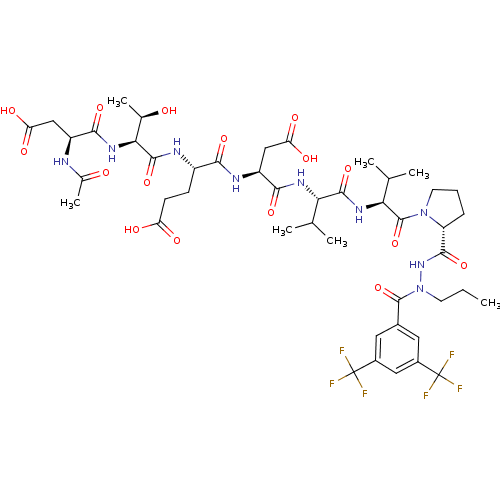
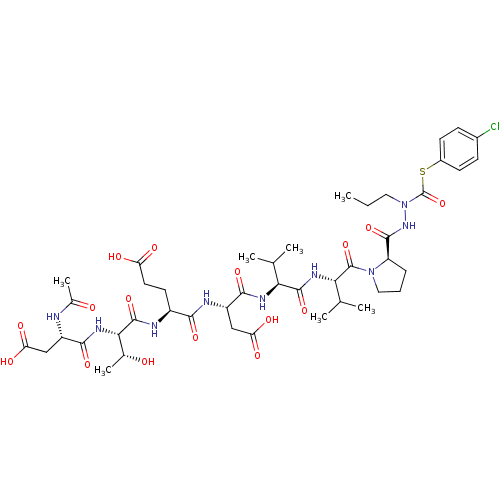
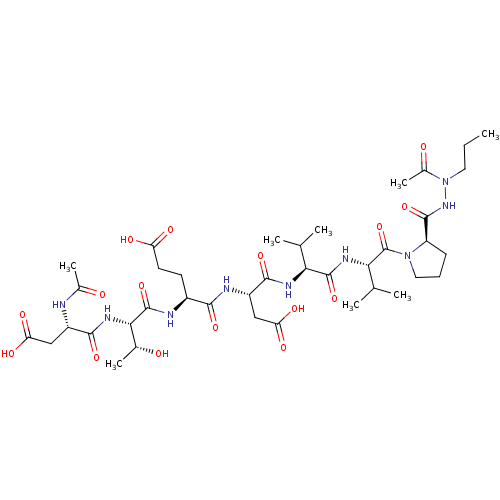
Affinity DataKi: 200nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
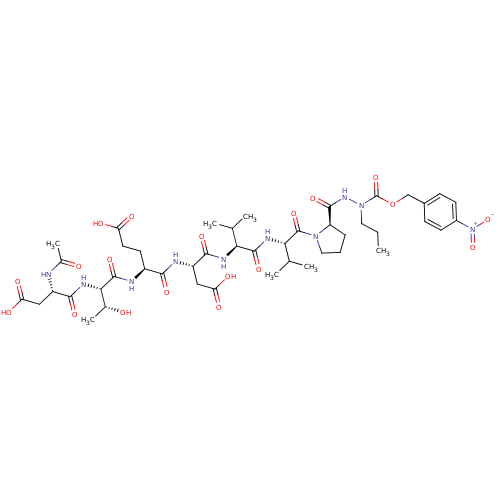
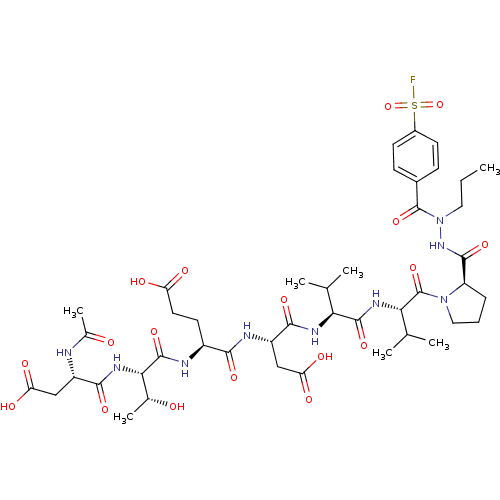
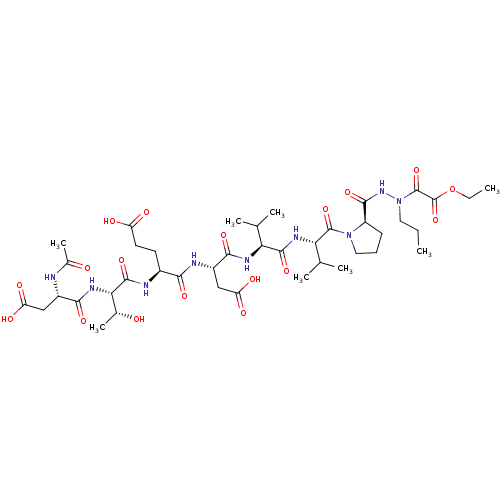
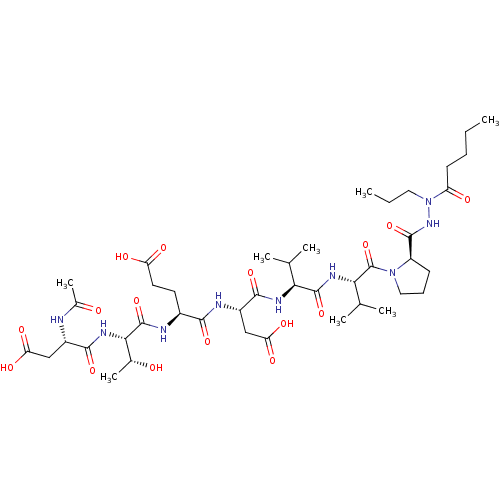
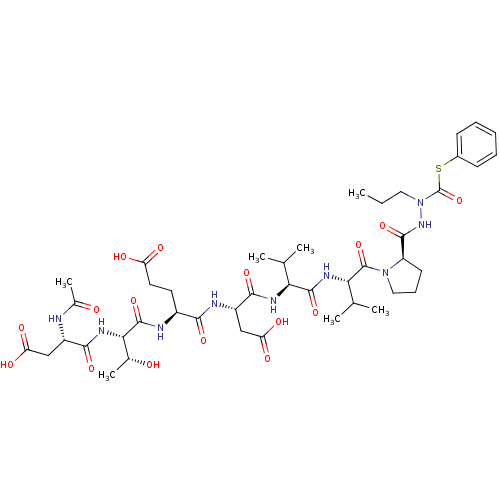
Affinity DataKi: 900nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
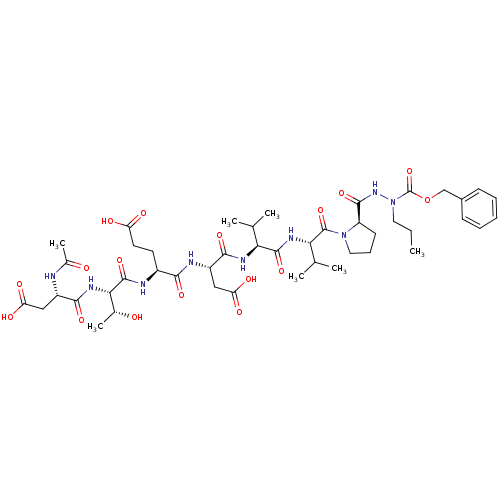
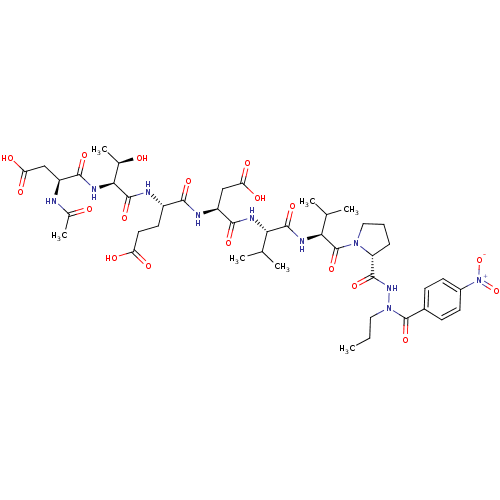
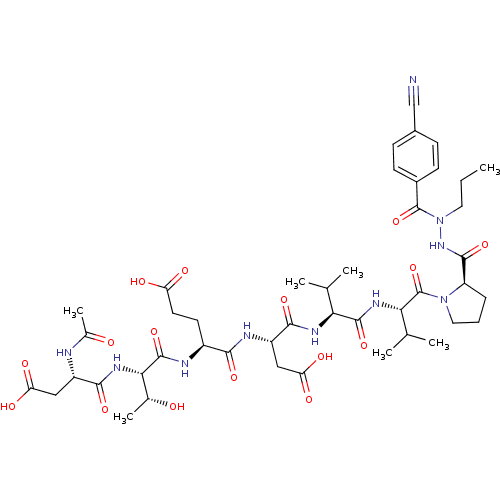
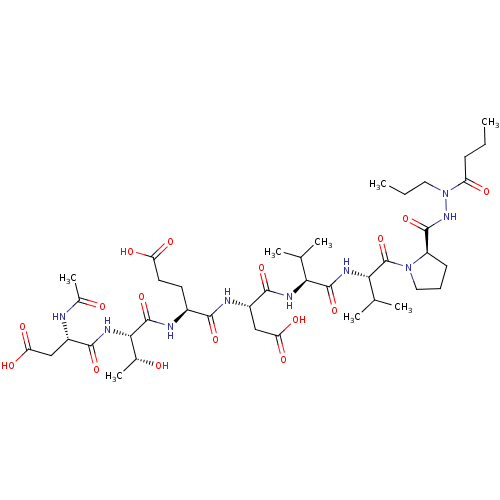
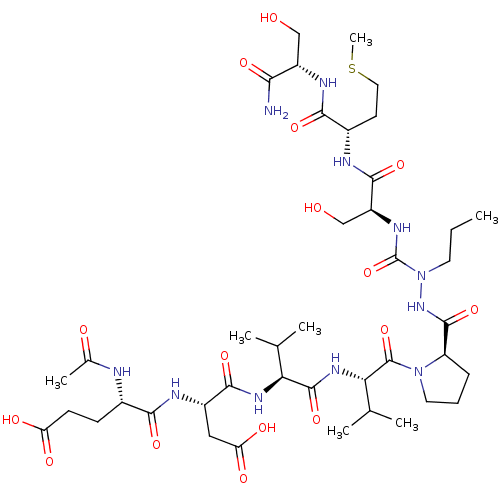
Affinity DataKi: 2.00E+3nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
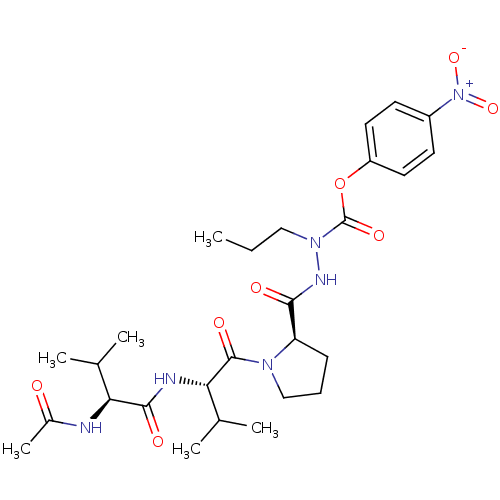
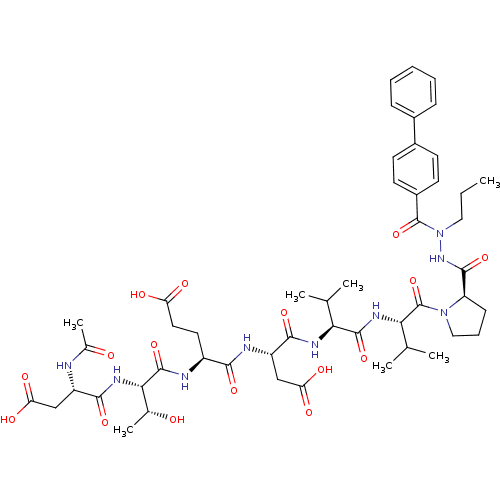
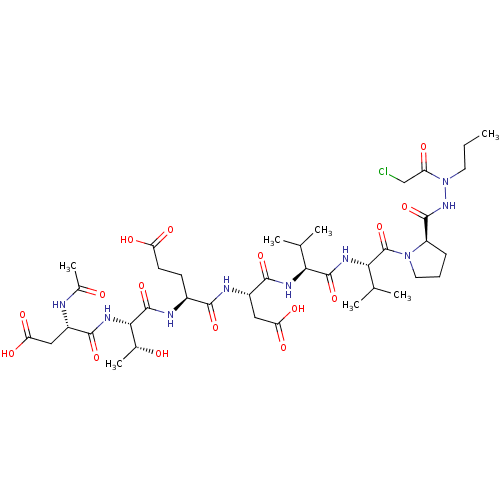
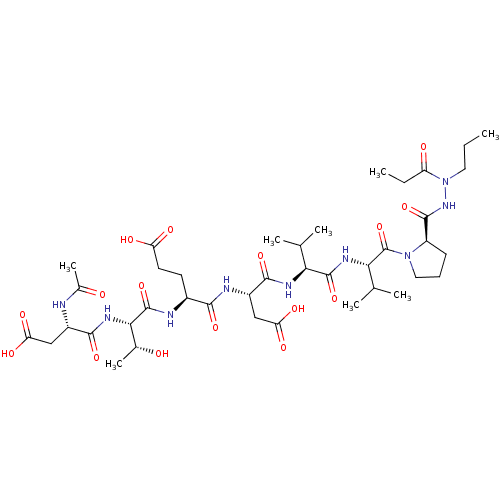
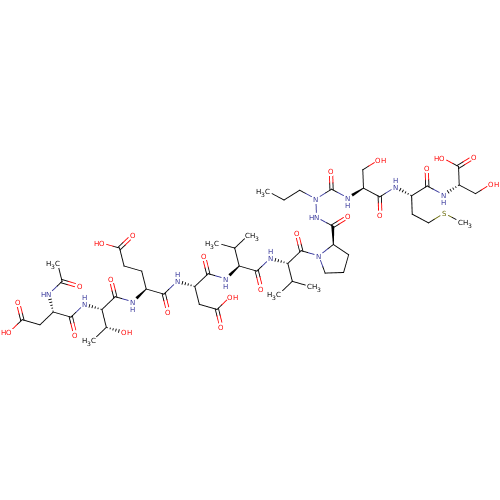
Affinity DataKi: 3.00E+3nMAssay Description:Compound was evaluated for its binding affinity against NS3 proteaseMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 6.00E+3nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 7.00E+3nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 8.00E+3nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 8.00E+3nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 9.00E+3nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 1.00E+4nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 1.60E+4nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 2.00E+4nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 2.00E+4nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 4.50E+4nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 8.00E+4nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >1.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 1.60E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 2.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: 2.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Schering-Plough Research Institute
Curated by ChEMBL
Schering-Plough Research Institute
Curated by ChEMBL
Affinity DataKi: >2.00E+5nMAssay Description:Michaelis-Menten constant was determined against Hepatitis C virus NS3 protease and represented as KmMore data for this Ligand-Target Pair