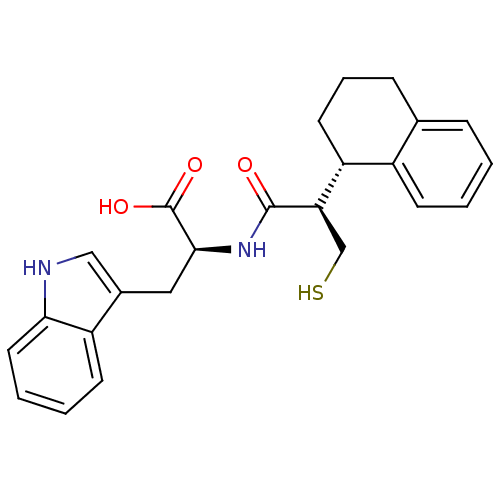
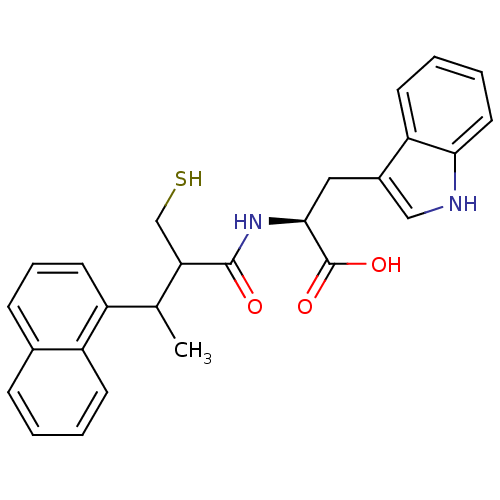
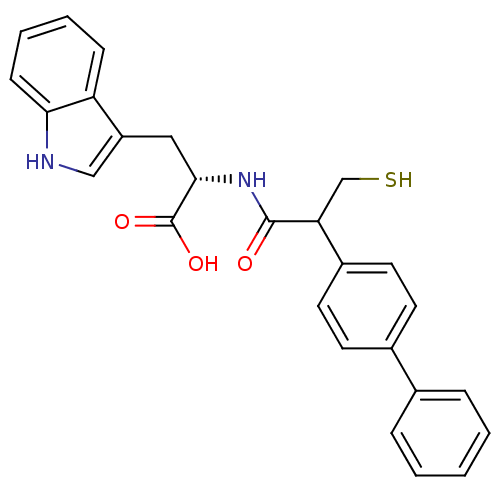
Report error Found 87 Enz. Inhib. hit(s) with all data for entry = 2526
Affinity DataKi: 0.700nM ΔG°: -54.4kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -52.4kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nM ΔG°: -51.9kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nM ΔG°: -51.5kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -51.4kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nM ΔG°: -51.4kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 2.40nM ΔG°: -51.2kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nM ΔG°: -51.1kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -50.6kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -50.6kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -50.6kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -49.9kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nM ΔG°: -49.6kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 4.80nM ΔG°: -49.4kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 5nM ΔG°: -49.3kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 5nM ΔG°: -49.3kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 5.40nM ΔG°: -49.1kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -48.1kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -47.5kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 10.8nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 22nM ΔG°: -45.5kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 27nM ΔG°: -44.9kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 30nM ΔG°: -44.7kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 35nM ΔG°: -44.3kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 35nM ΔG°: -44.3kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 43nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 43nM ΔG°: -43.7kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 51nM ΔG°: -43.3kJ/molepH: 7.4 T: 2°CAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 62nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 65nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 79nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: >100nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:NEP was preincubated in black 96-well microplates with or without increasing concentrations of inhibitors. DGPA was added, and the reaction was stopp...More data for this Ligand-Target Pair
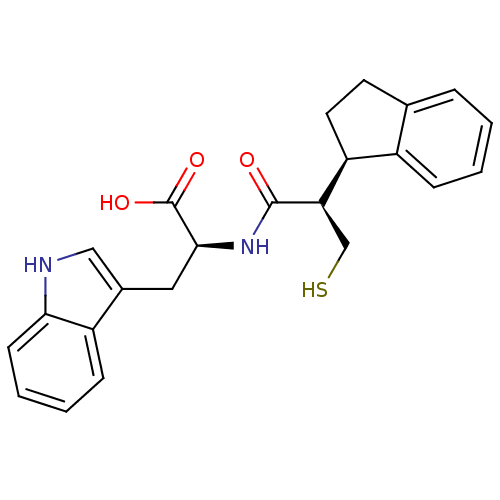
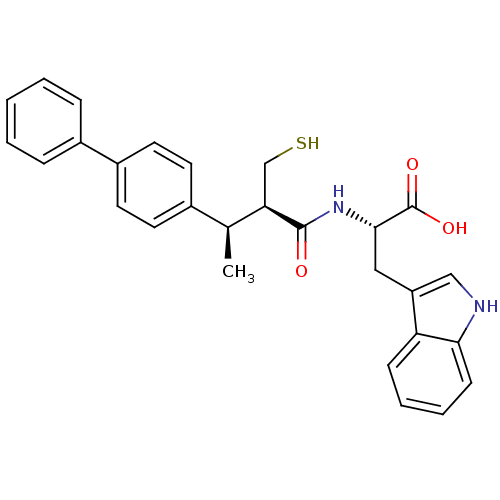
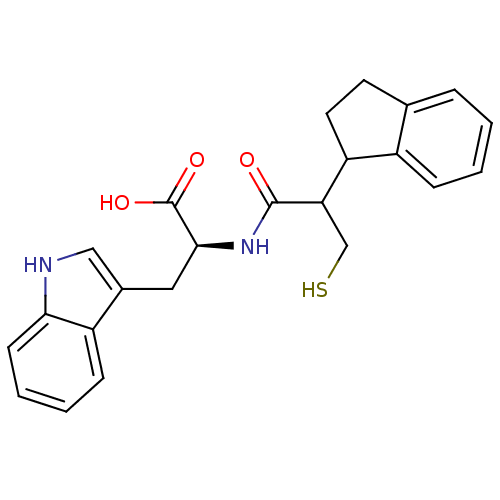
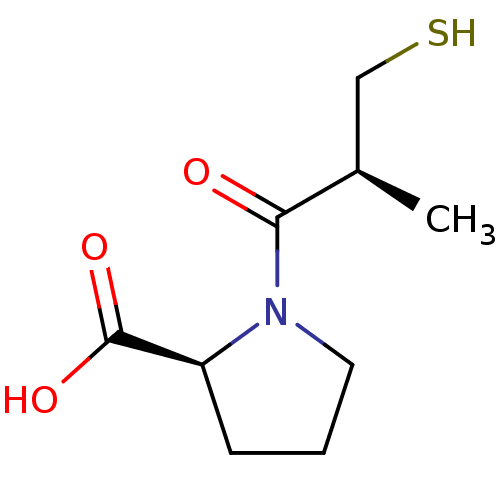
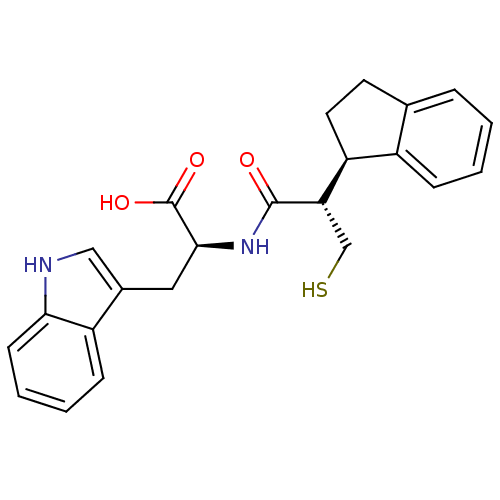
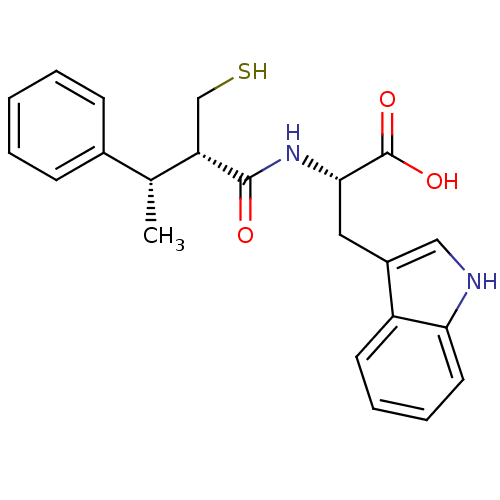
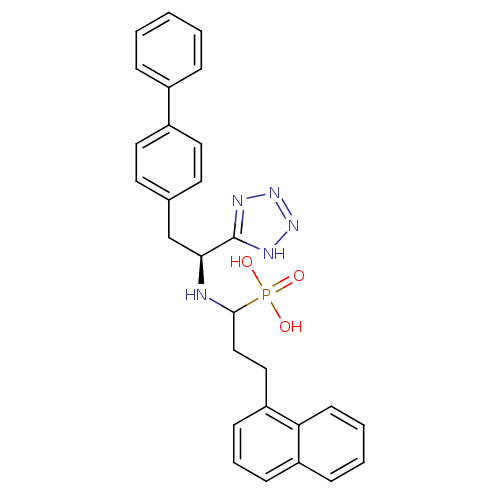
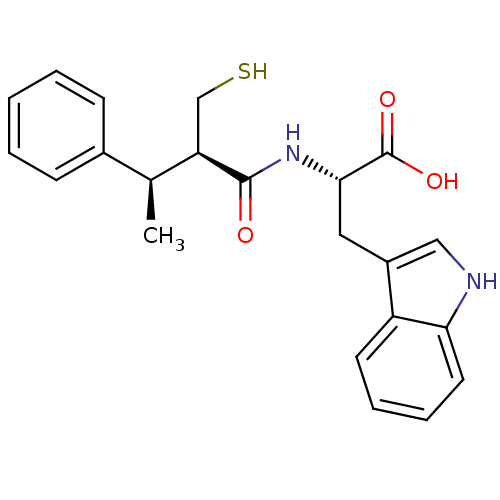
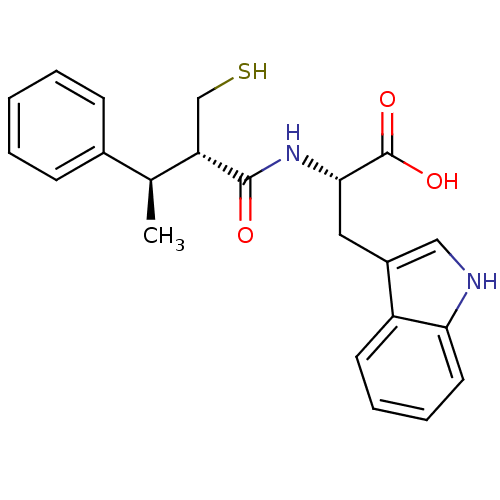
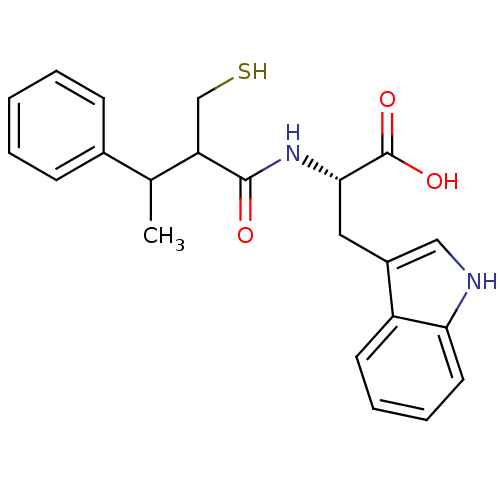
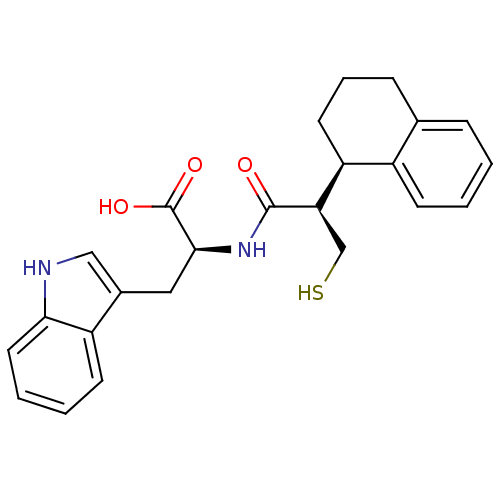
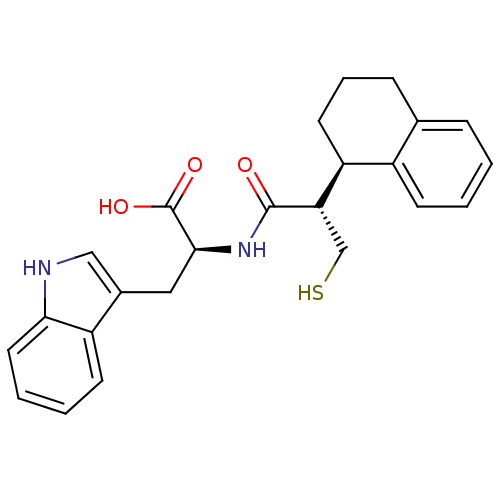
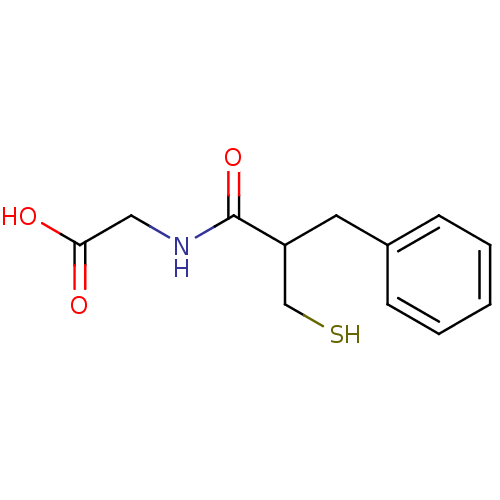
 3D Structure (crystal)
3D Structure (crystal)