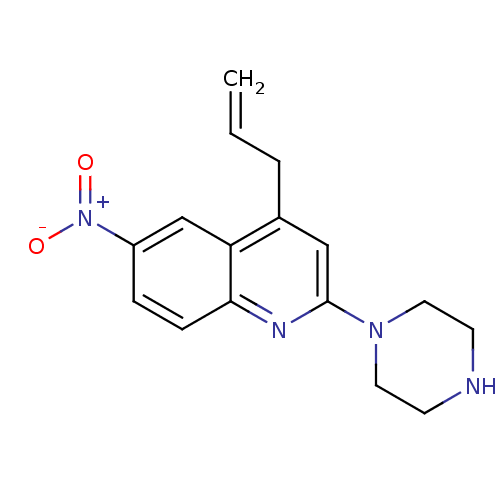
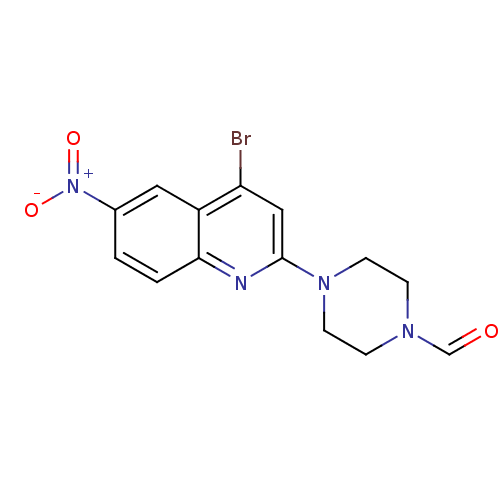
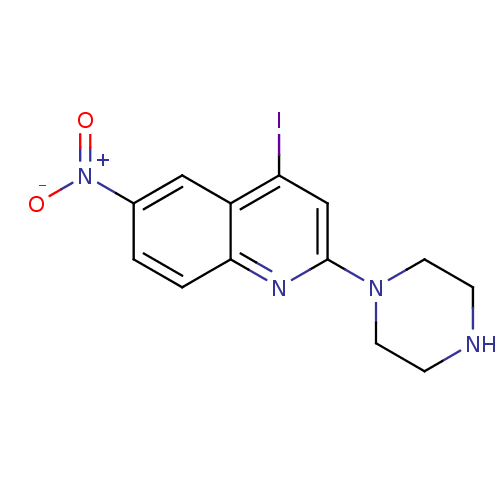
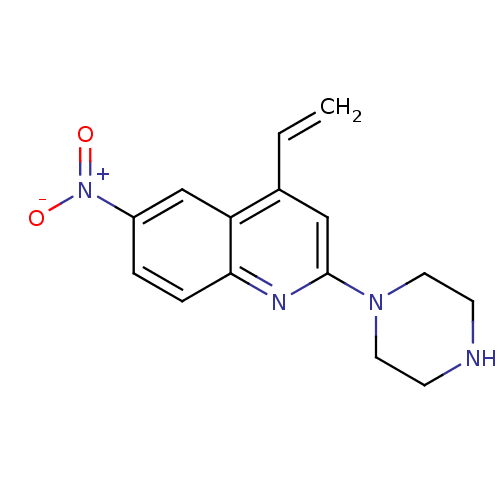
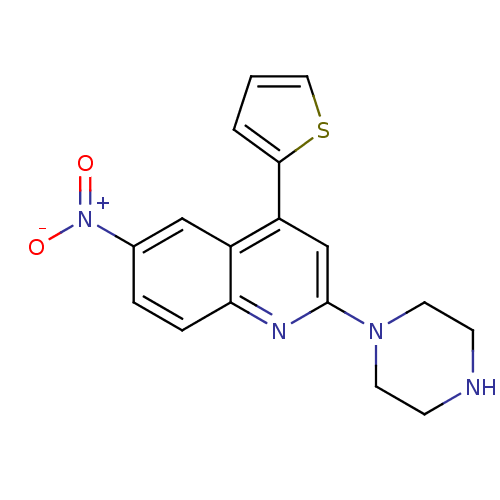
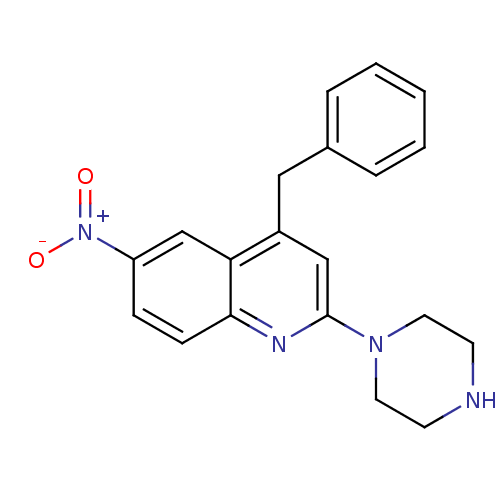
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 50035277
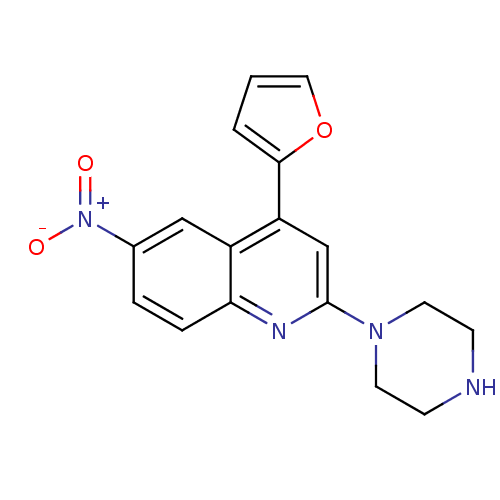
Affinity DataKi: 0.0130nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
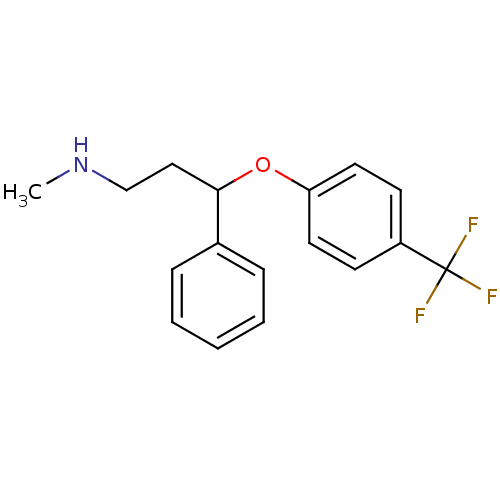
Affinity DataKi: 0.0300nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
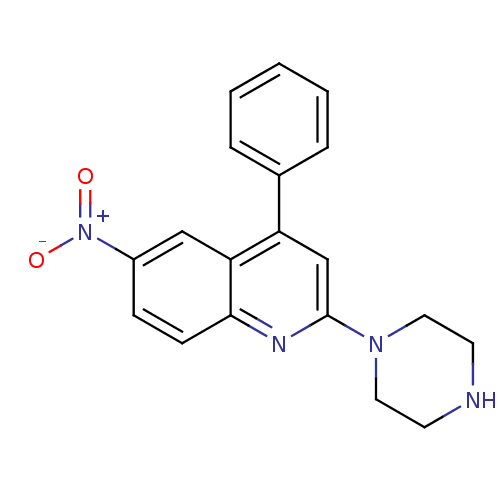
Affinity DataKi: 0.170nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
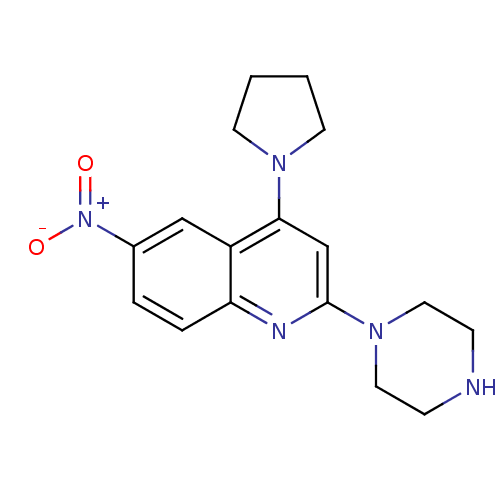
Affinity DataKi: 0.320nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 0.370nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 0.530nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 22nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 67nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
Affinity DataKi: 127nMAssay Description:Compound was evaluated for its ability to displace [3H]citalopram binding to the rat cortical Serotonin transporterMore data for this Ligand-Target Pair
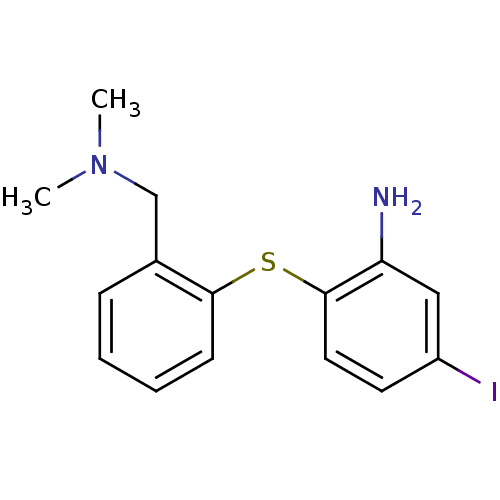
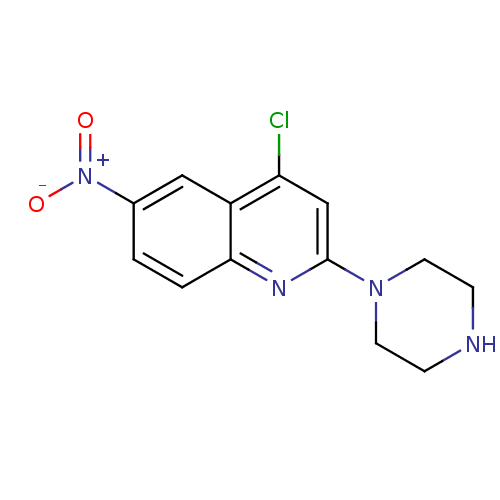
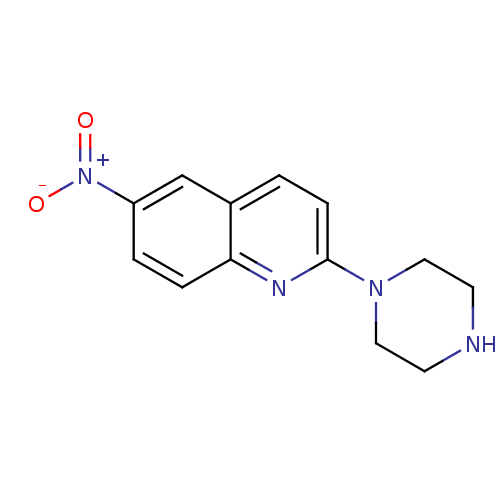
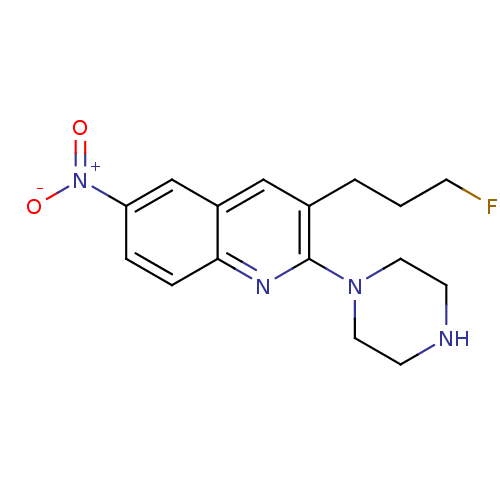
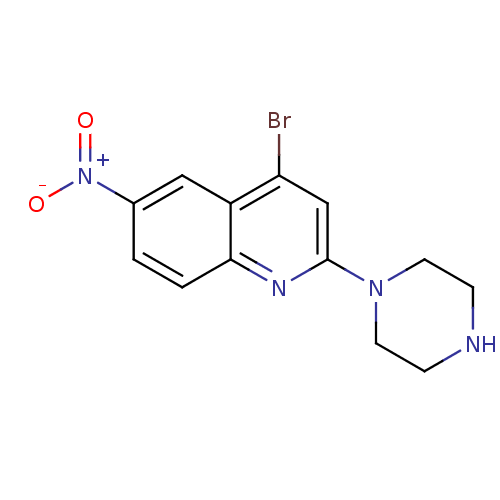
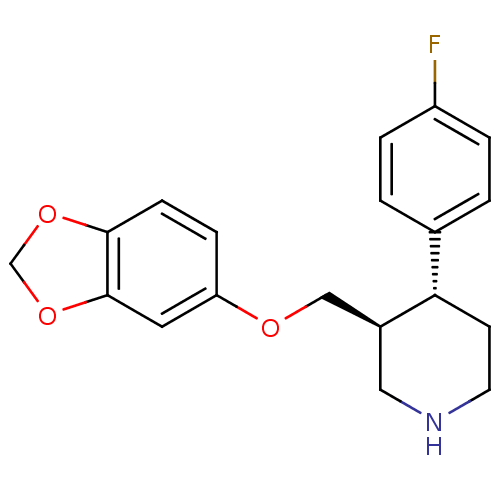
 3D Structure (crystal)
3D Structure (crystal)