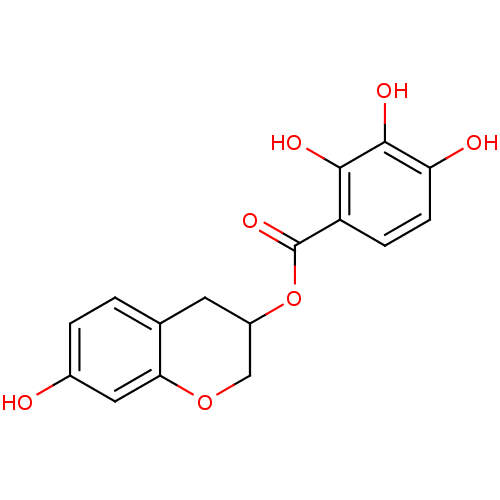
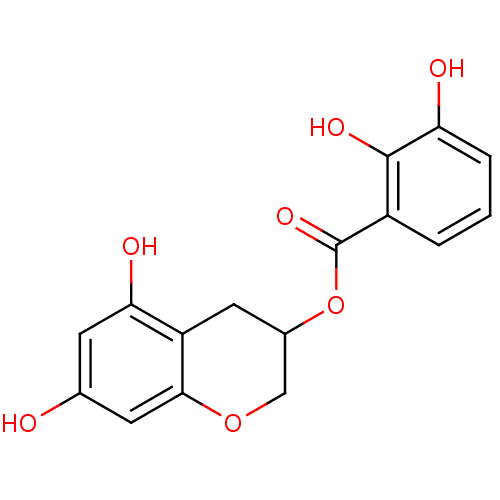
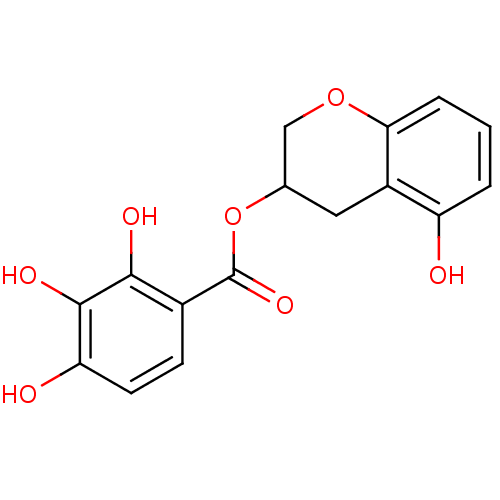
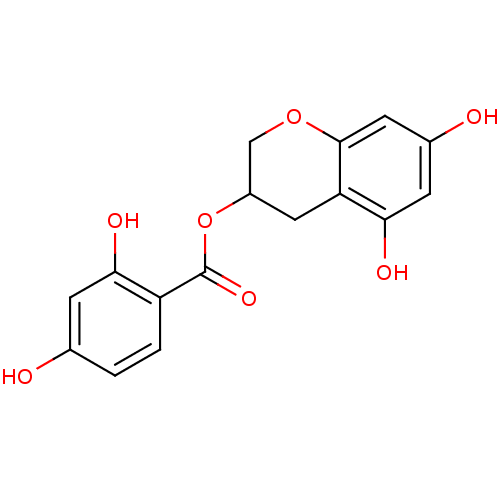
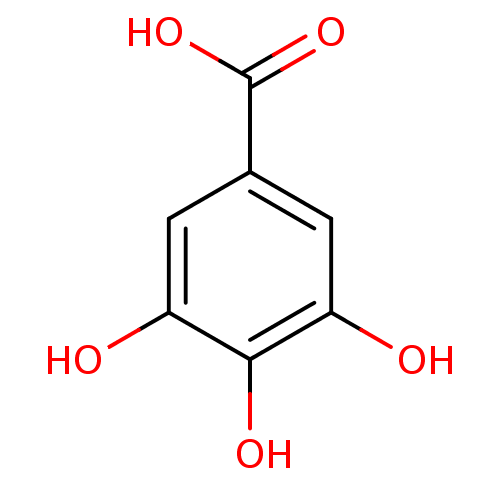
Report error Found 20 Enz. Inhib. hit(s) with all data for entry = 50035174
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 730nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 760nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 2.36E+3nMAssay Description:Inhibitory concentration against polymerization in A17 double mutant HIV-1 RT using a template primer of poly rC/dG12-18 and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 3.83E+3nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 6.33E+3nMAssay Description:Inhibitory concentration against polymerization in A17 double mutant HIV-1 RT using a template primer of poly rC/dG12-18 and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.07E+4nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 3.08E+4nMAssay Description:Inhibitory concentration against polymerization in A17 double mutant HIV-1 RT using a template primer of poly rC/dG12-18 and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 3.63E+4nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 3.72E+4nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 4.55E+4nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 6.47E+4nMAssay Description:Inhibitory concentration against polymerization in A17 double mutant HIV-1 RT using a template primer of poly rC/dG12-18 and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 9.63E+4nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory concentration against polymerization in A17 double mutant HIV-1 RT using a template primer of poly rC/dG12-18 and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory concentration against polymerization in A17 double mutant HIV-1 RT using a template primer of poly rC/dG12-18 and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibitory concentration against polymerization in A17 double mutant HIV-1 RT using a template primer of poly rC/dG12-18 and [3H]dGTPMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
University of Toledo
Curated by ChEMBL
University of Toledo
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of polymerization in wild type HIV-1 RT with poly rC/dG12-18 template primer and [3H]dGTPMore data for this Ligand-Target Pair