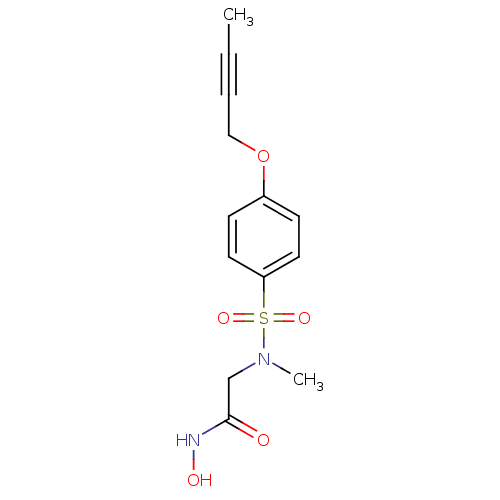
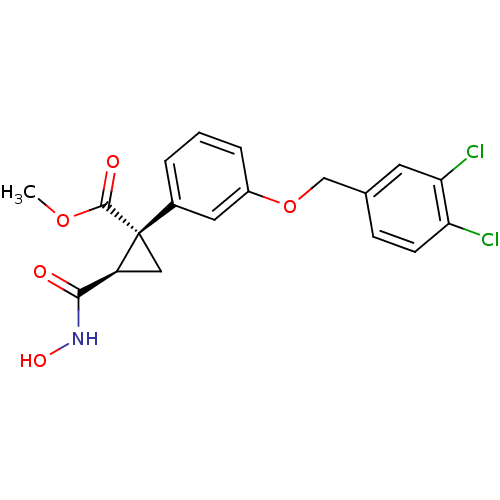
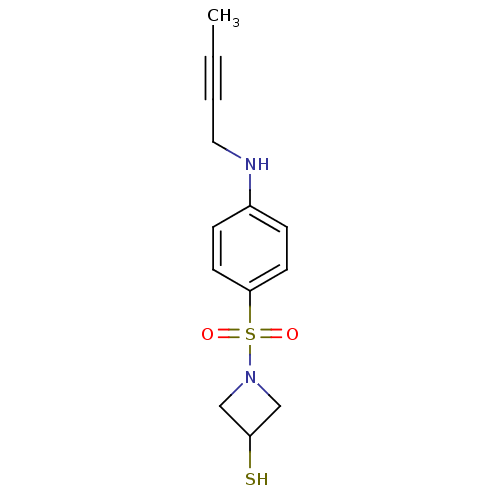
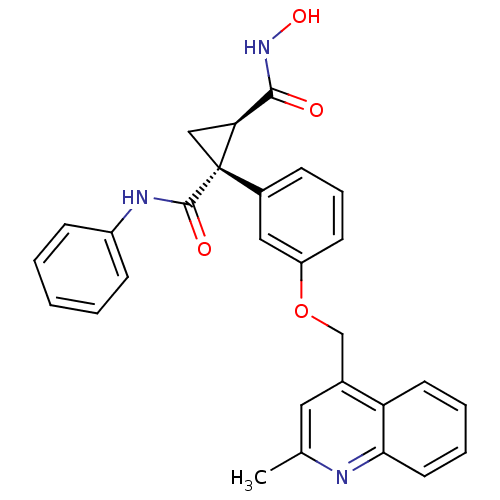
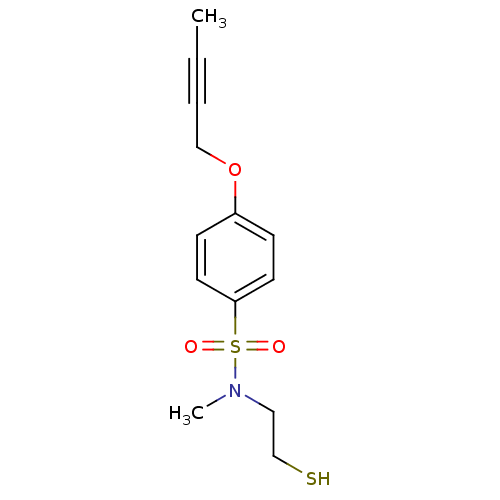
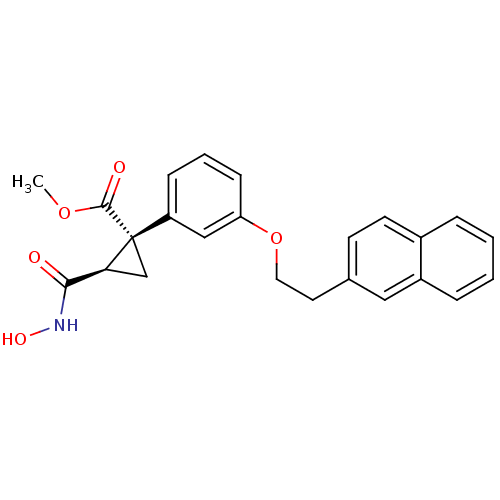
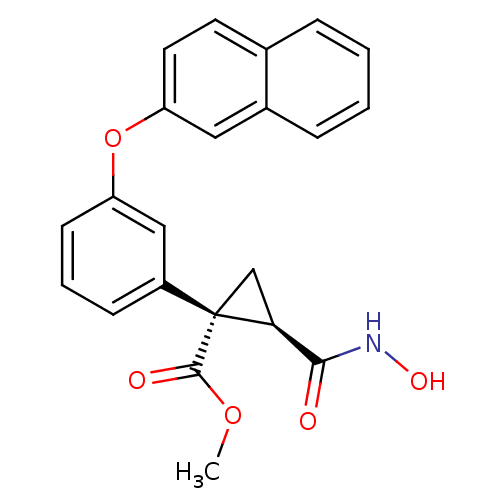
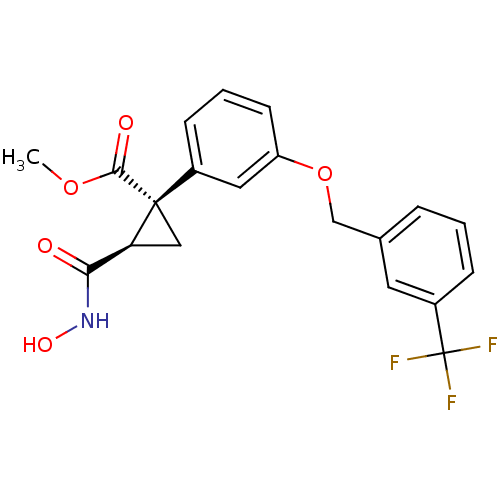
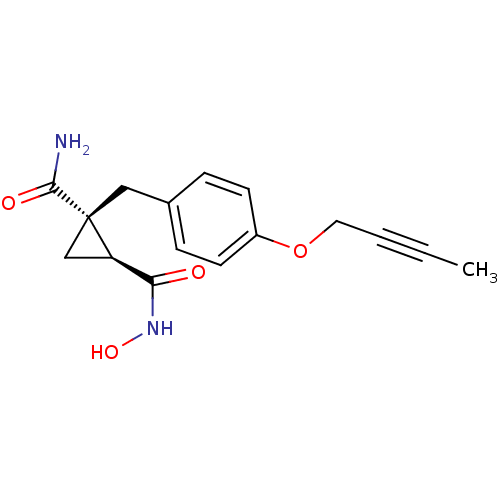
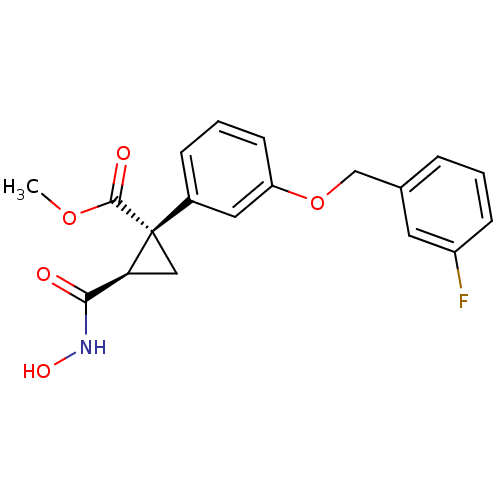
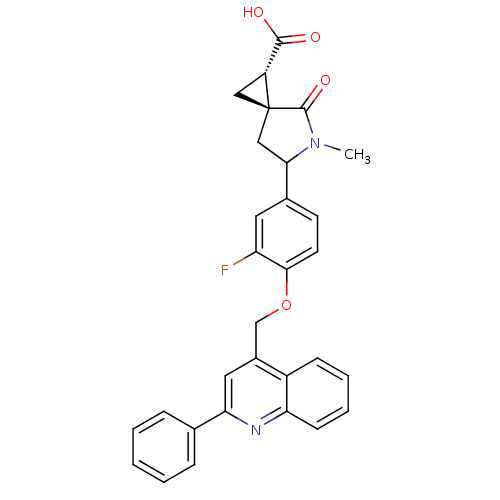
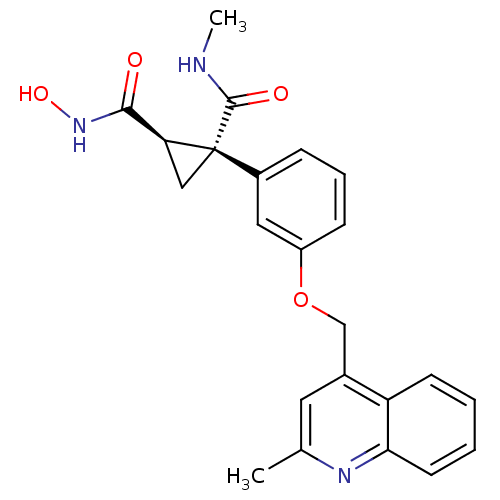
Report error Found 86 Enz. Inhib. hit(s) with Target = 'Disintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q]'
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 0.0600nM ΔG°: -57.8kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 0.140nM ΔG°: -55.7kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 0.180nM ΔG°: -55.1kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 1nMAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2nM ΔG°: -49.2kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nMAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nM ΔG°: -48.2kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 4.5nM ΔG°: -47.2kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5nM ΔG°: -46.9kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5nM ΔG°: -46.9kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 6.70nM ΔG°: -46.2kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 8nMAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 8nM ΔG°: -45.7kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 8nM ΔG°: -45.7kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 9nM ΔG°: -45.5kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 10nM ΔG°: -45.2kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 10nM ΔG°: -45.2kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 10nM ΔG°: -45.2kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 11nM ΔG°: -45.0kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 11nM ΔG°: -45.0kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 11nM ΔG°: -45.0kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 12nM ΔG°: -44.8kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 13nM ΔG°: -44.6kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 14nMAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 16nM ΔG°: -44.0kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 17nM ΔG°: -43.9kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 18nM ΔG°: -43.8kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 27nM ΔG°: -42.8kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 28nM ΔG°: -42.7kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 28nM ΔG°: -42.7kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 30nM ΔG°: -42.5kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 30nM ΔG°: -42.5kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 32nM ΔG°: -42.3kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 32nM ΔG°: -42.3kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 33nM ΔG°: -42.3kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 33nM ΔG°: -42.3kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 40nM ΔG°: -41.8kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 50nM ΔG°: -41.3kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 50nM ΔG°: -41.3kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 50nM ΔG°: -41.3kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 53nM ΔG°: -41.1kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 55nM ΔG°: -41.0kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 56nM ΔG°: -41.0kJ/molepH: 7.5 T: 2°CAssay Description:The compounds were tested for TACE inhibition using fluorescence resonance energy transfer (FRET) assay. TACE catalyzed cleavage of the substrate pep...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 60nM ΔG°: -40.8kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 70nM ΔG°: -40.4kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 70nM ΔG°: -40.4kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 80nMAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 80nM ΔG°: -40.1kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 94nM ΔG°: -39.7kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
TargetDisintegrin and metalloproteinase domain-containing protein 17 [215-477,S266A,N452Q](Human)
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 100nM ΔG°: -39.6kJ/molepH: 7.3 T: 2°CAssay Description:Enzyme activity was determined by a kinetic assay measuring the rate of increase in fluorescent intensity generated by the cleavage of an internally ...More data for this Ligand-Target Pair
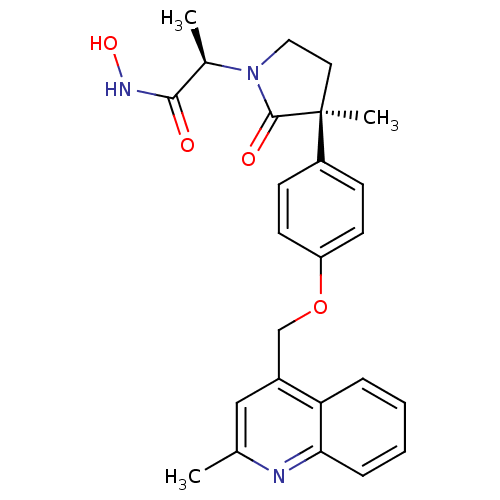
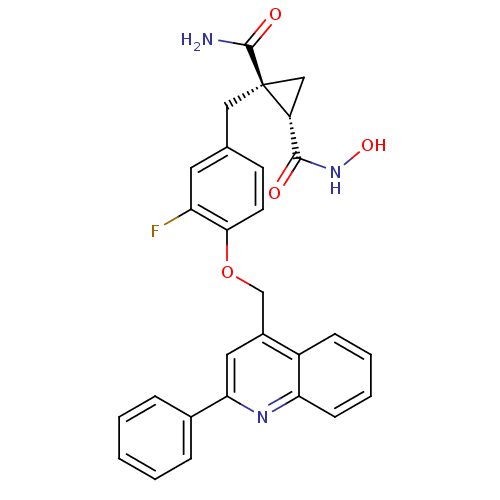
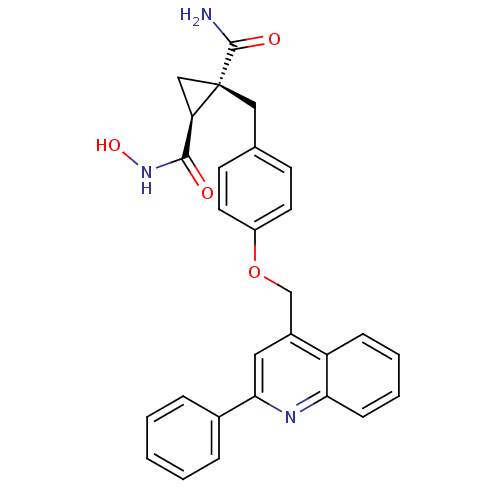
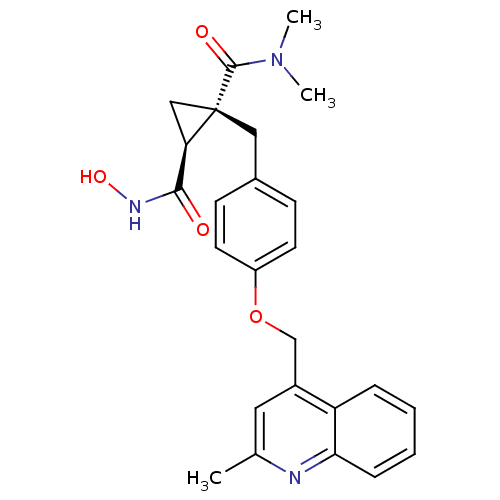
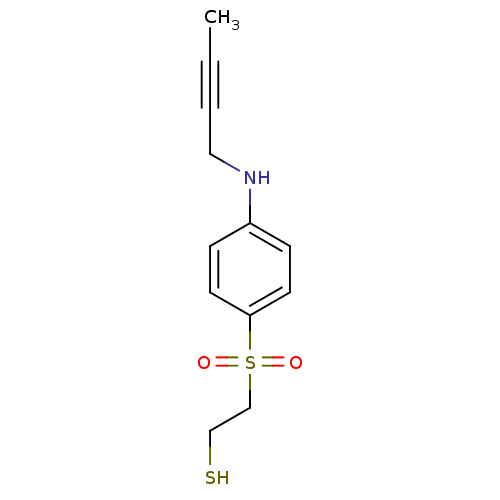
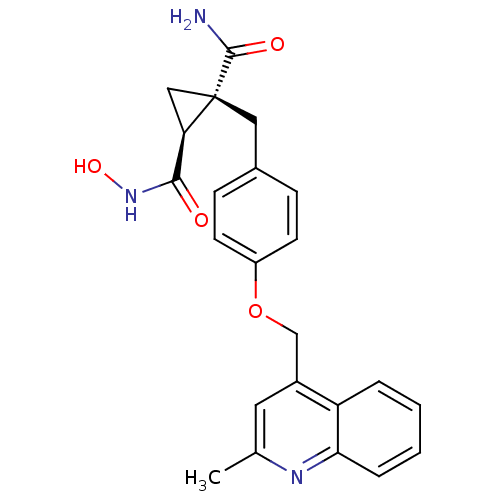
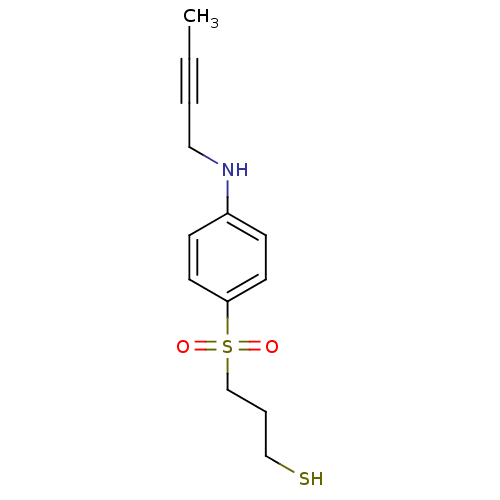
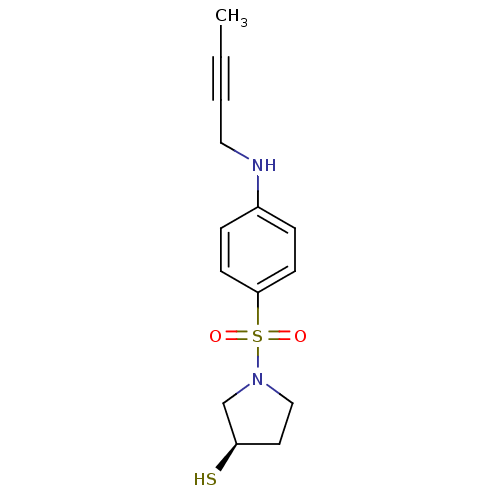
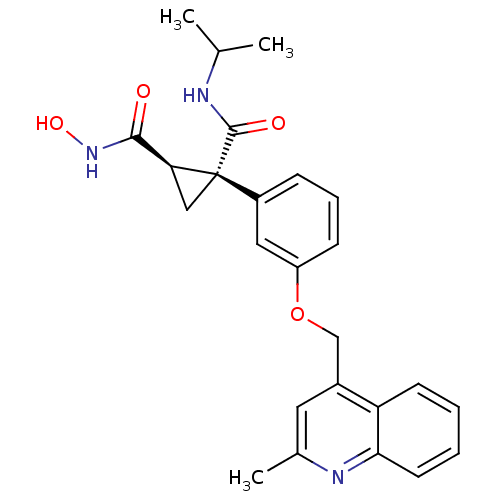
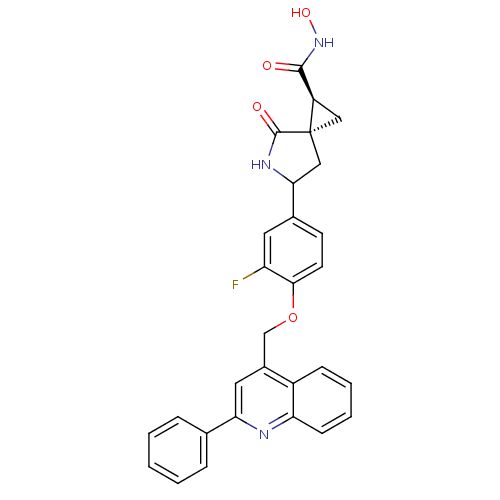
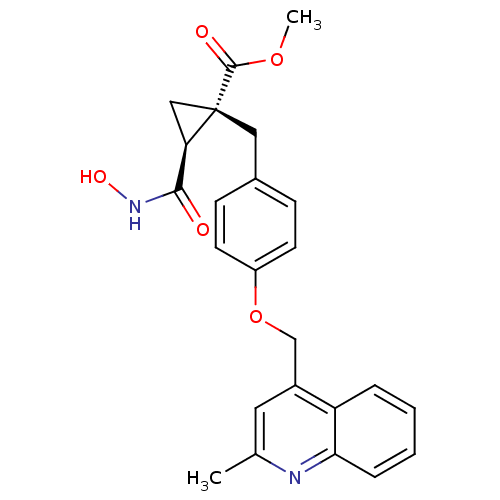
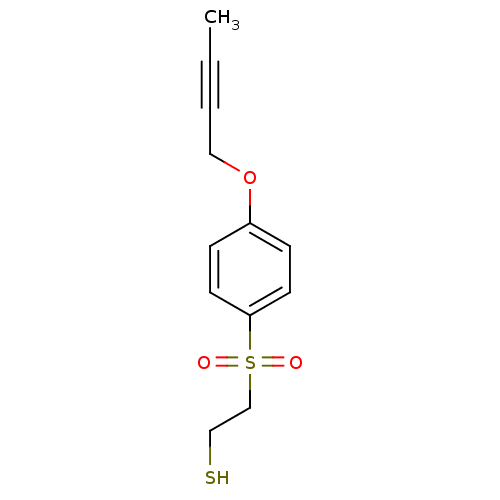
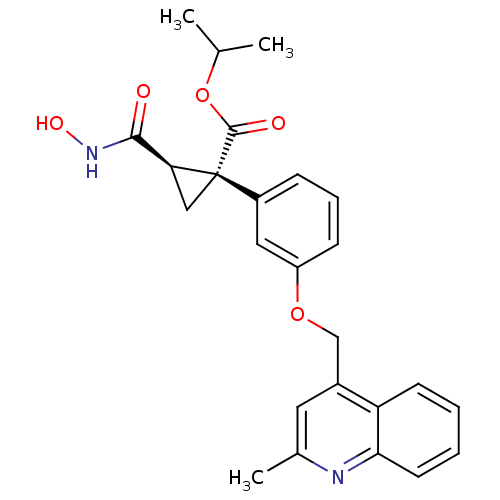
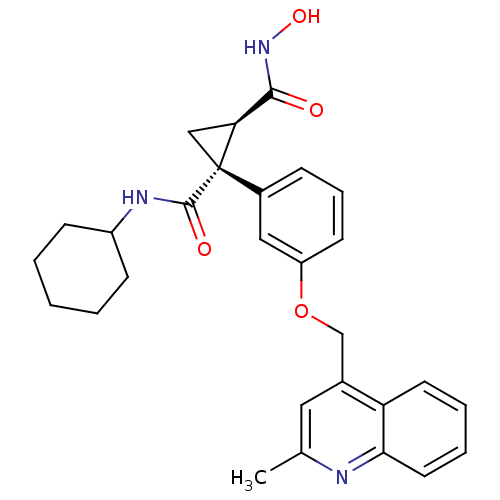
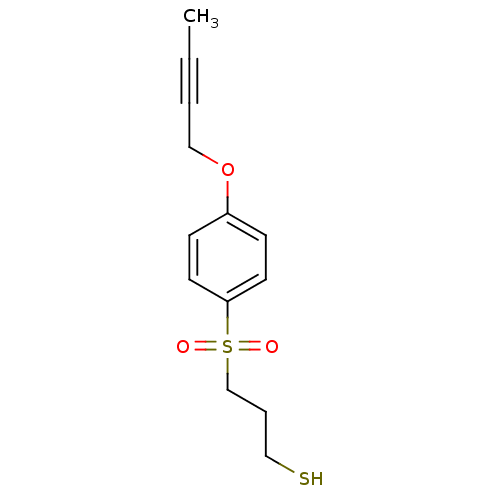
 3D Structure (crystal)
3D Structure (crystal)