BDBM195611 GSKJ1
SMILES OC(=O)CCNc1cc(nc(n1)-c1ccccn1)N1CCc2ccccc2CC1
InChI Key InChIKey=AVZCPICCWKMZDT-UHFFFAOYSA-N
Data 28 IC50
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
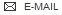 Found 28 hits for monomerid = 195611
Found 28 hits for monomerid = 195611
Affinity DataIC50: 1.70nMAssay Description:Inhibition of KDM5B (1 to 809 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.70nMAssay Description:Inhibition of epitope-tagged KDM6B (1026 to 1682 residues)(unknown origin) transfected in human U2OS cells using H3K27me2 peptide as substrateMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 2.10nMAssay Description:Inhibition of KDM2B (1 to 650 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 2.80nMAssay Description:Inhibition of KDM6B (1043 to 1643 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 2.80nMAssay Description:Inhibition of PHF8 (1 to 1024 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 3.40nMAssay Description:Inhibition of KDM4C (1 to 349 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 5nMAssay Description:Inhibition of KDM4A (1 to 350 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 5.30nMAssay Description:Inhibition of KDM6A (919 to 1401 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 5.5nMAssay Description:Inhibition of KDM5C (2 to 1560 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 6.80nMAssay Description:Inhibition of KDM5A (1 to 1090 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 7.30nMAssay Description:Inhibition of KDM4B (2 to 500 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 7.5nMAssay Description:Inhibition of KDM3B (842 to 1761 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 7.5nMAssay Description:Inhibition of KDM3A (2 to 1322 residues)(unknown origin) by Alphalisa assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 60nMAssay Description:Inhibition of EZH2 Y641F mutant (unknown origin)More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 60nMAssay Description:Competitive inhibition of KDM6B (unknown origin) using biotinylated H3K27me3 peptide as substrate preincubated for 15 mins followed by substrate addi...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 60nMAssay Description:Inhibition of recombinant human N-terminal His6/Flag-tagged TEV-protease cleavage site fused JMJD3 1637-1675 deletion mutant (1141 to 1682 residues) ...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 170nMAssay Description:Inhibition of KDM5B (1 to 809 residues) (unknown origin) AlphaLISA assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of recombinant human N-terminal His6/Flag-tagged TEV-protease cleavage site fused JMJD3 1637-1675 deletion mutant (1141 to 1682 residues) ...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of recombinant human N-terminal His6/Flag-tagged TEV-protease cleavage site fused JMJD3 1637-1675 deletion mutant (1141 to 1682 residues) ...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.96E+4nMAssay Description:Inhibition of recombinant human JmjD2e using biotinylated H3K9(1-15)me3 as substrate preincubated for 15 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of recombinant human JmjD1a using biotinylated H3K9(1-21)me2 as substrate preincubated for 15 mins followed by substrate addition and meas...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 4.43E+4nMAssay Description:Inhibition of human JmjD2a by Alphascreen assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 5.60E+4nMAssay Description:Inhibition of human UTX using Biotin-KAPRKQLATKAARK(me3)SAPATGG as substrate by MALDI-TOF analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 6.30E+4nMAssay Description:Inhibition of human JmjD2e using H3K27(me3) as substrate preincubated for 10 mins followed by substrate addition and measured after 7 mins in presenc...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 7.90E+4nMAssay Description:Inhibition of human JmjD2d using H3K27(me3) as substrate preincubated for 10 mins followed by substrate addition and measured after 7 mins in presenc...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of human JmjD2c using H3K27(me3) as substrate preincubated for 10 mins followed by substrate addition and measured after 7 mins in presenc...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of human JmjD2a using H3K27(me3) as substrate preincubated for 10 mins followed by substrate addition and measured after 7 mins in presenc...More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of human Jarid1c by MALDI-TOF analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
