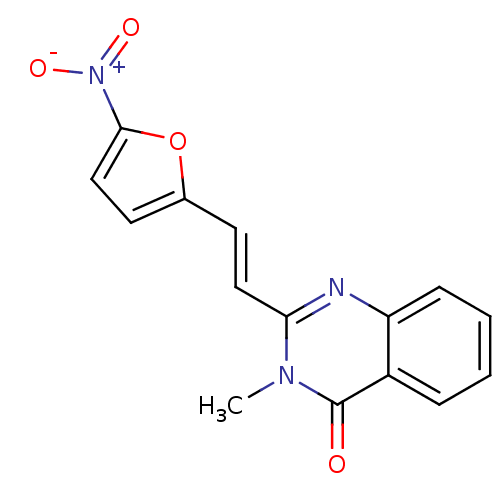
BDBM71904 3-Methyl-2-[2-(5-nitro-furan-2-yl)-vinyl]-3H-quinazolin-4-one::3-methyl-2-[(E)-2-(5-nitro-2-furanyl)ethenyl]-4-quinazolinone::3-methyl-2-[(E)-2-(5-nitro-2-furyl)vinyl]quinazolin-4-one::3-methyl-2-[(E)-2-(5-nitrofuran-2-yl)ethenyl]quinazolin-4-one::MLS000529350::SMR000121825::cid_5731073
SMILES Cn1c(\C=C\c2ccc(o2)[N+]([O-])=O)nc2ccccc2c1=O
InChI Key InChIKey=SBNIQRGKIQSRPT-UHFFFAOYSA-N
Data 2 IC50
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 71904
Found 2 hits for monomerid = 71904
TargetMitochondrial import inner membrane translocase subunit TIM23(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 6.28E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair
TargetMitochondrial import inner membrane translocase subunit TIM10(Baker's yeast)
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Burnham Center For Chemical Genomics
Curated by PubChem BioAssay
Affinity DataIC50: 7.30E+3nMAssay Description:Data Source: Sanford-Burnham Center for Chemical Genomics (SBCCG) Source Affiliation: Sanford-Burnham Medical Research Institute(SBMRI, San Diego, CA...More data for this Ligand-Target Pair