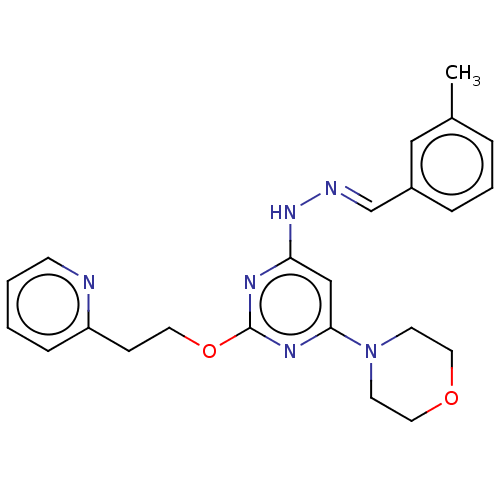
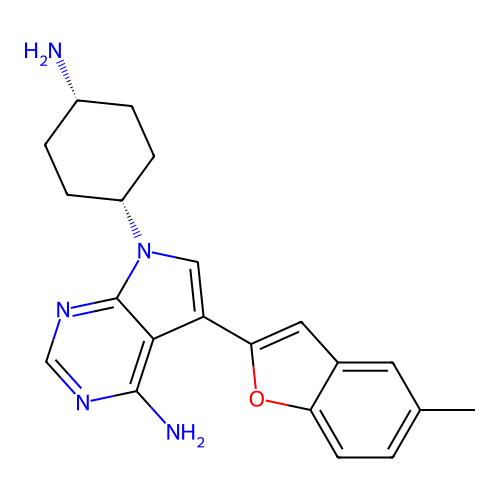
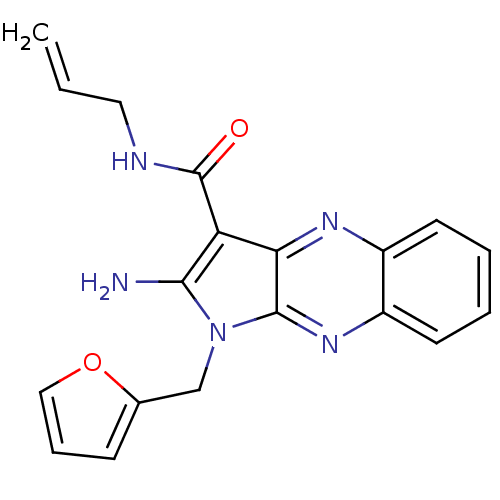
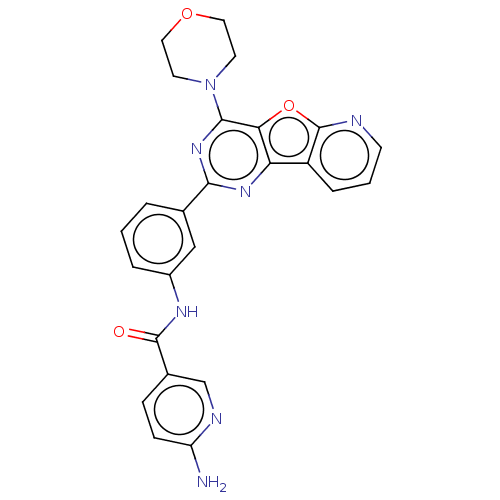
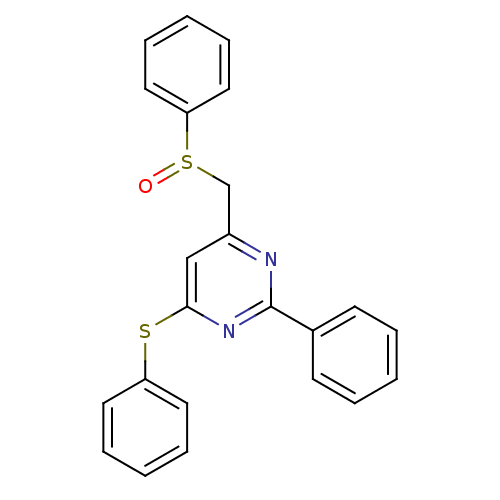
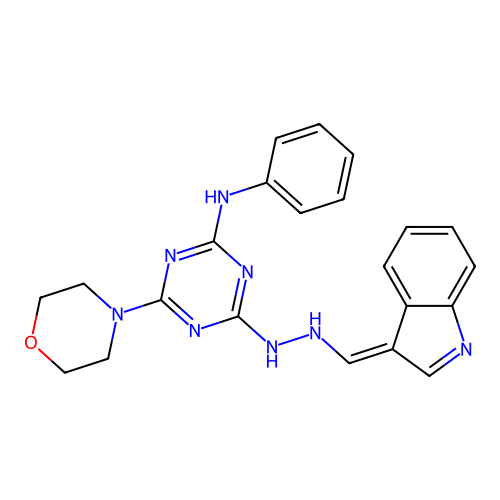
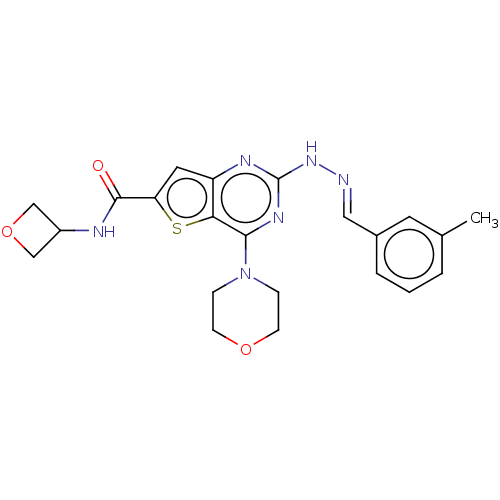
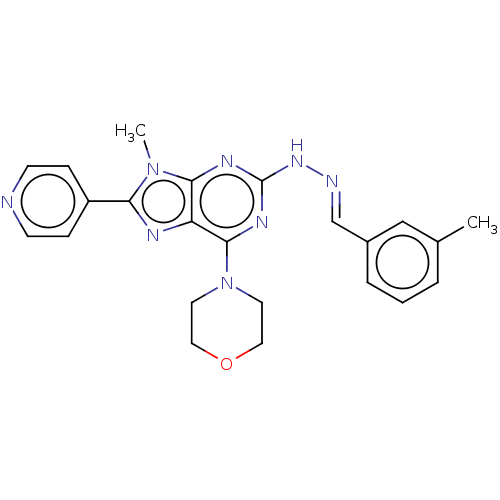
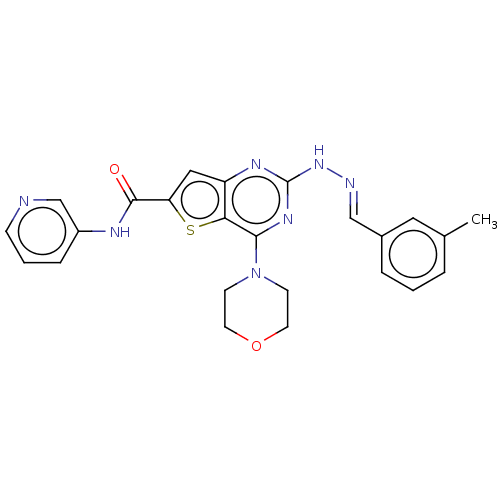
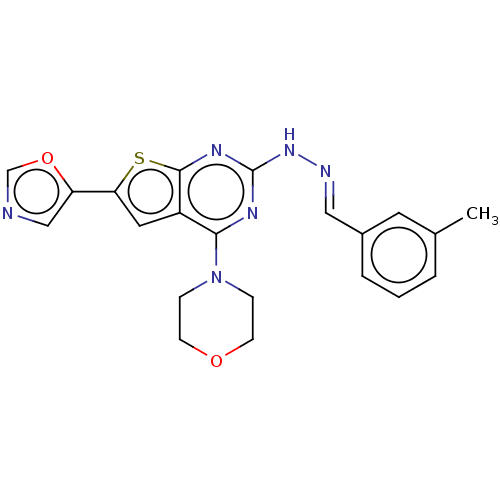
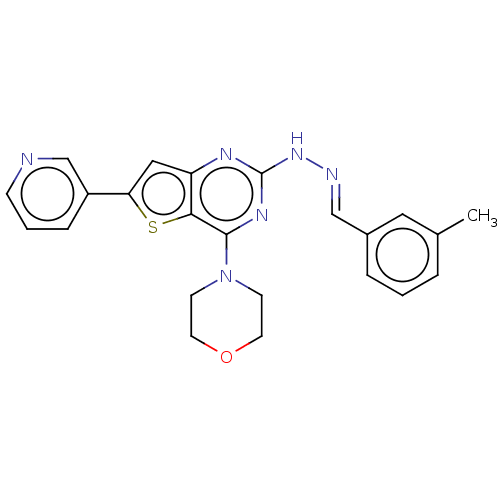
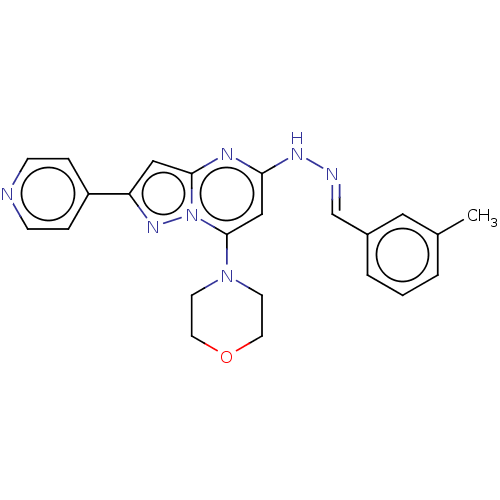
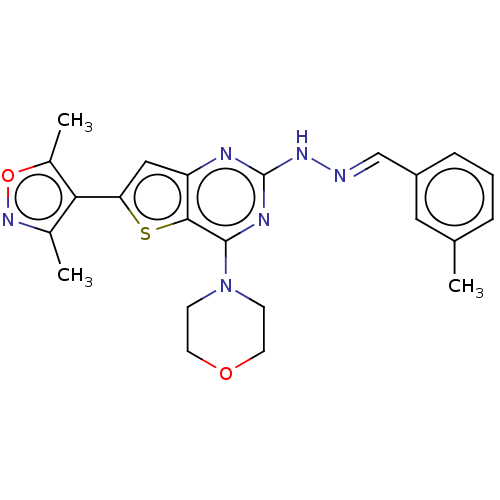
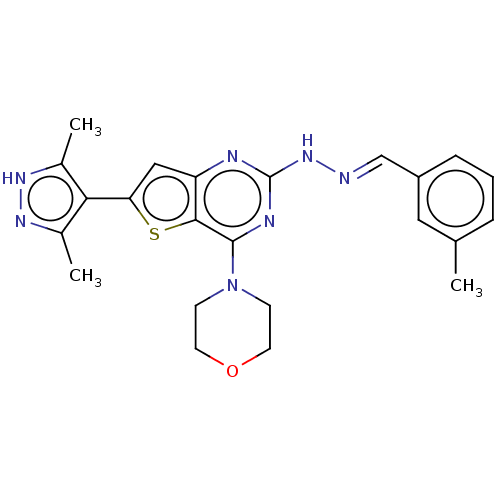
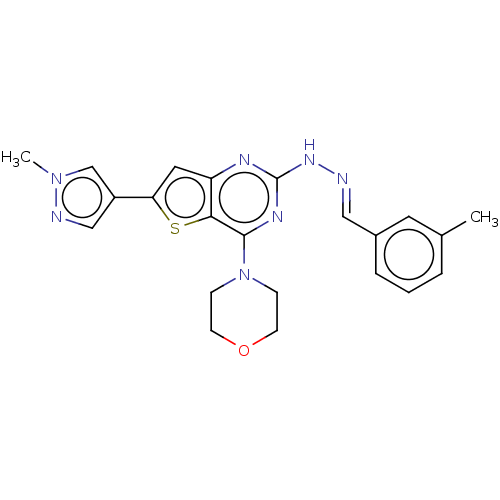
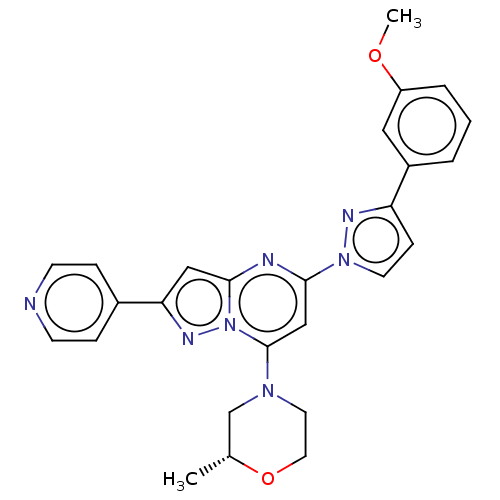
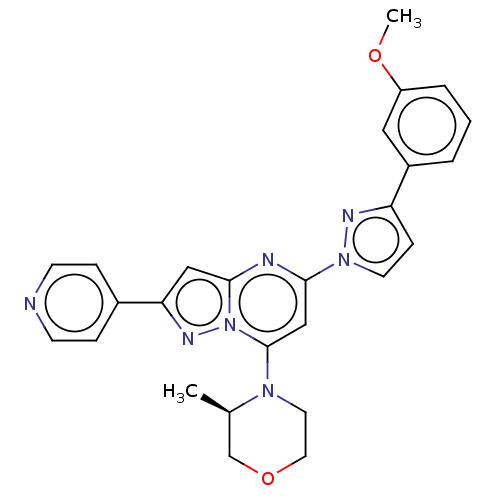
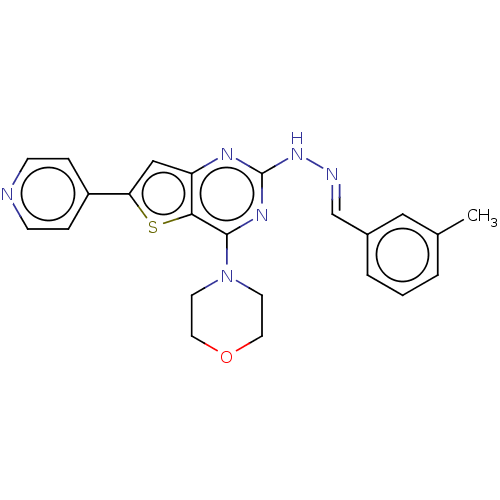
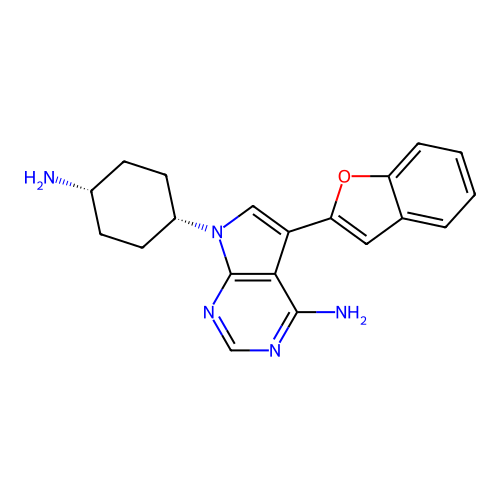
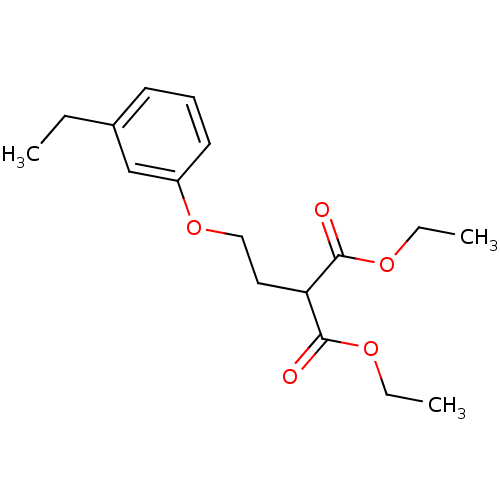
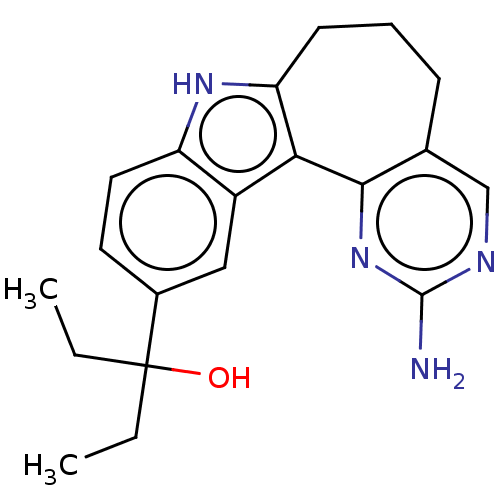
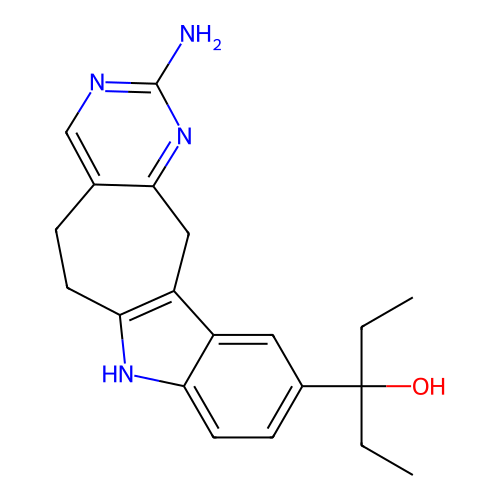
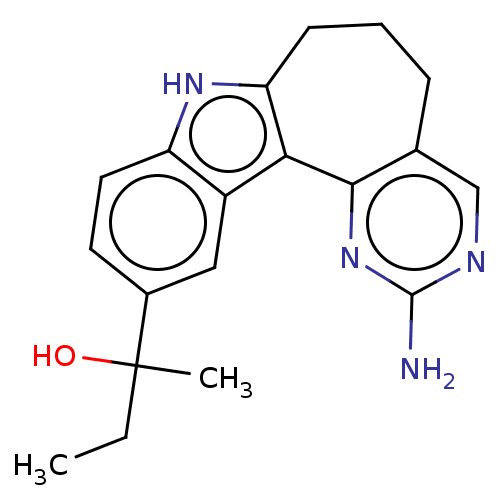
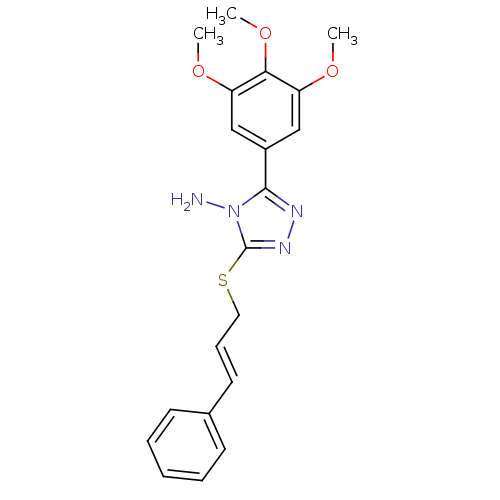
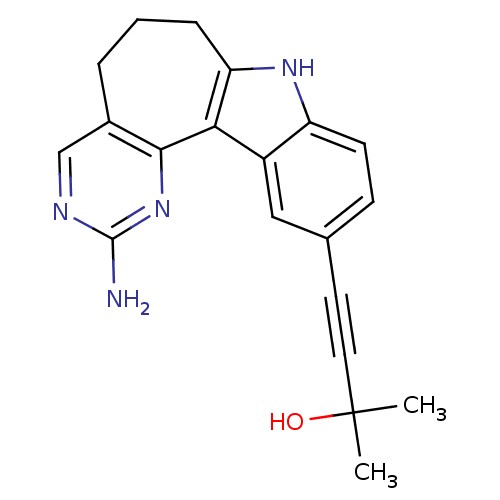
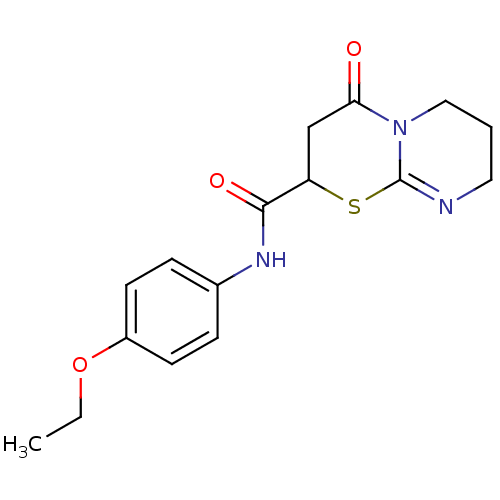
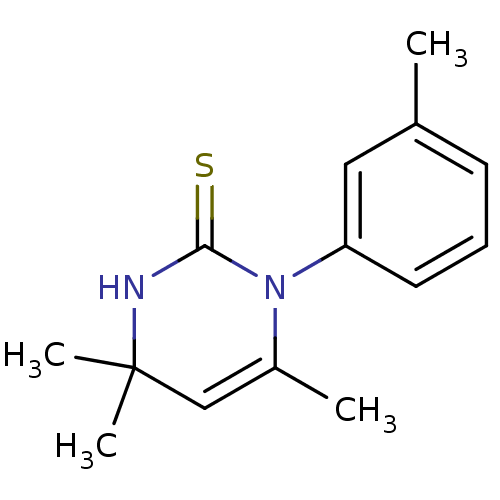
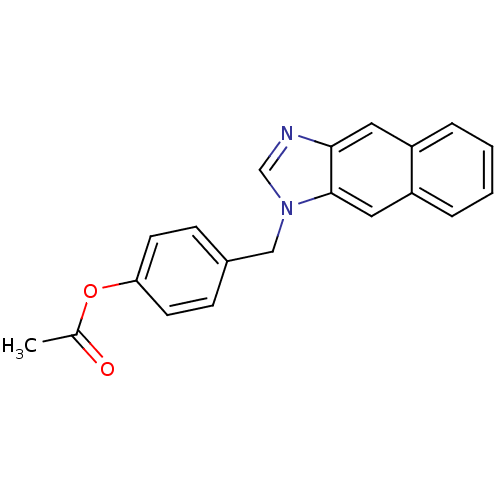
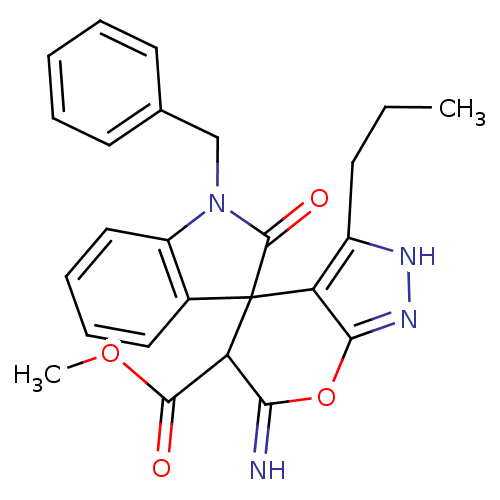
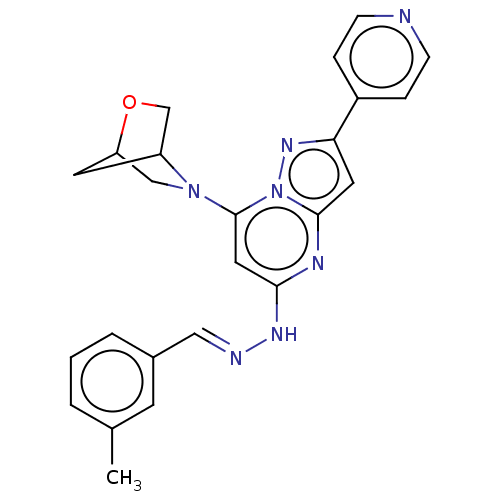
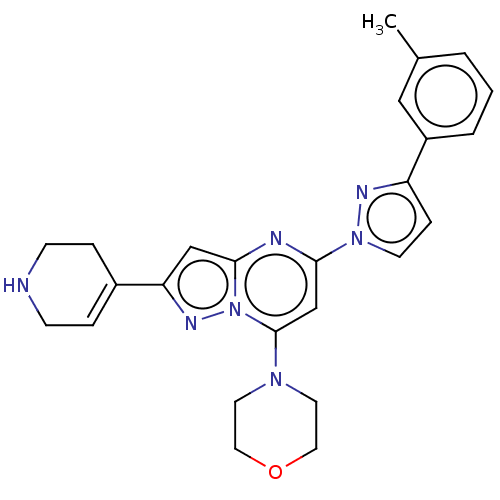
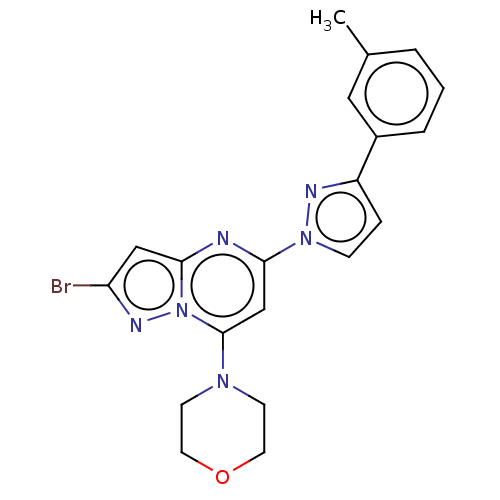
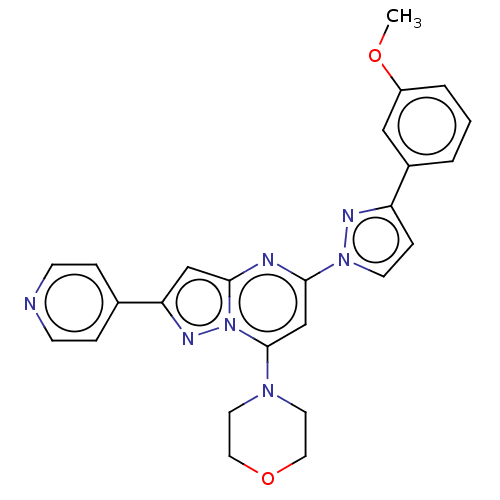
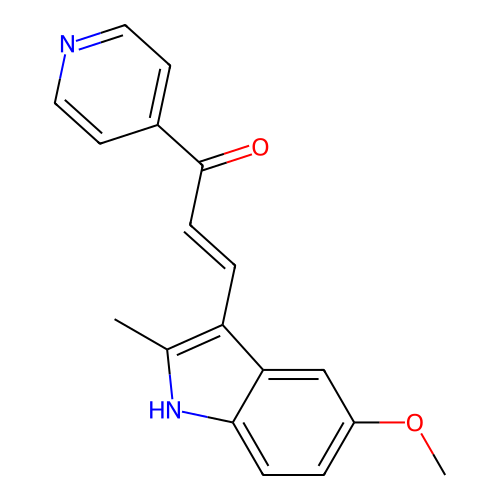
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 0.0750nMAssay Description:Binding affinity to human wild type partial length PIKFYVE (F1512 to C2098 residues) mammalian expression assessed as dissociation constant by Kinome...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 0.309nMAssay Description:Inhibition of PIKFYVE in HEK293 cells by NanoBRET assayMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 0.312nMAssay Description:Inhibition of PIKFYVE in HEK293 cells by NanoBRET assayMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 0.470nMAssay Description:Binding affinity to PIKfyve (unknown origin) assessed as dissociation constant incubated for 1 hr by qPCR analysisMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 0.700nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 0.820nMAssay Description:Inhibition of human GST tagged PIKFYVE (1493 to end residues) expressed in baculovirus-infected Sf9 cells using PI(3)P:PS as substrate incubated for ...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 0.900nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 0.910nMAssay Description:Binding affinity to N-terminal His-tagged human PIKfyve expressed in Sf9 cells assessed as dissociation constantMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:The biochemical PIKFyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo Kinase assa...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:The biochemical PIKFyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo Kinase assa...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:Kinase-tagged T7 phage strains are prepared in an E. coli host derived from the BL21 strain. E. coli are grown to log-phase and infected with T7 phag...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase [1-2098,S696N,L932L,Q995L,T998S,S1033A,Q1183K](Human)
Kineta
US Patent
Kineta
US Patent
Affinity DataIC50: 1nMAssay Description:The biochemical PIKFyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo Kinase assa...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:The biochemical PIKfyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo™ Kinase ass...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase [1-2098,S696N,L932L,Q995L,T998S,S1033A,Q1183K](Human)
Kineta
US Patent
Kineta
US Patent
Affinity DataIC50: 1nMAssay Description:PIKfyve Biochemical Assay. The biochemical PIKFyve inhibition assays were run by Cama Biosciences according to proprietary methodology based on the P...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:Kinase-tagged T7 phage strains are prepared in an E. coli host derived from the BL21 strain. E. coli are grown to log-phase and infected with T7 phag...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: <1nMAssay Description:For most assays, kinase-tagged T7 phage strains were prepared in an E. colihost derived from the BL21 strain. E. coli were grown to log-phase and inf...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1.20nMAssay Description:Inhibition of PIKFYVE in HEK293 cells by NanoBRET assayMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1.20nMAssay Description:Inhibition of PIKFYVE in HEK293 cells by NanoBRET assayMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 1.70nMAssay Description:Binding affinity to PIKfyve (unknown origin) assessed as dissociation constant incubated for 1 hr by qPCR analysisMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1.70nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1.80nMAssay Description:Inhibition of human GST tagged PIKFYVE (1493 to end residues) expressed in baculovirus-infected Sf9 cells using PI(3)P:PS as substrate incubated for ...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1.80nMAssay Description:Inhibition of PIKfyve (unknown origin)More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 1.90nMAssay Description:Inhibition of human GST tagged PIKFYVE (1493 to end residues) expressed in baculovirus-infected Sf9 cells using PI(3)P:PS as substrate incubated for ...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.20nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.20nMAssay Description:Inhibition of human GST tagged PIKFYVE (1493 to end residues) expressed in baculovirus-infected Sf9 cells using PI(3)P:PS as substrate incubated for ...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.20nMAssay Description:Inhibition of human wild type partial length PIKfyve (F1512 to C2098 residues) expressed in mammalian expression system by Kinomescan assayMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.20nMAssay Description:Inhibition of PIKfyve (unknown origin)More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.40nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.5nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.60nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2.70nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:The biochemical PIKFyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo Kinase assa...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 3nMAssay Description:Kinase-tagged T7 phage strains are prepared in an E. coli host derived from the BL21 strain. E. coli are grown to log-phase and infected with T7 phag...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase [1-2098,S696N,L932L,Q995L,T998S,S1033A,Q1183K](Human)
Kineta
US Patent
Kineta
US Patent
Affinity DataIC50: 3nMAssay Description:The biochemical PIKFyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo Kinase assa...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 3nMAssay Description:Kinase-tagged T7 phage strains are prepared in an E. coli host derived from the BL21 strain. E. coli are grown to log-phase and infected with T7 phag...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:The biochemical PIKfyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo™ Kinase ass...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 3nMAssay Description:Kinase-tagged T7 phage strains are prepared in an E. coli host derived from the BL21 strain. E. coli are grown to log-phase and infected with T7 phag...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase [1-2098,S696N,L932L,Q995L,T998S,S1033A,Q1183K](Human)
Kineta
US Patent
Kineta
US Patent
Affinity DataIC50: 3nMAssay Description:PIKfyve Biochemical Assay. The biochemical PIKFyve inhibition assays were run by Cama Biosciences according to proprietary methodology based on the P...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:The biochemical PIKFyve inhibition assays were run by Carna Biosciences according to proprietary methodology based on the Promega ADP-Glo Kinase assa...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 3.40nMAssay Description:Inhibition of human GST tagged PIKFYVE (1493 to end residues) expressed in baculovirus-infected Sf9 cells using PI(3)P:PS as substrate incubated for ...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 3.70nMAssay Description:Inhibition of human GST tagged PIKFYVE (1493 to end residues) expressed in baculovirus-infected Sf9 cells using PI(3)P:PS as substrate incubated for ...More data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataKd: 3.80nMAssay Description:Binding affinity to PIKfyve (unknown origin) assessed as dissociation constant incubated for 1 hr by qPCR analysisMore data for this Ligand-Target Pair
Target1-phosphatidylinositol 3-phosphate 5-kinase(Human)
University of North Carolina at Chapel Hill
Curated by ChEMBL
University of North Carolina at Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of Nano-Luc fused PIKFYVE (unknown origin) expressed in HEK293T cells pretreated for 3 hrs followed by K8-tarcer addition and measured aft...More data for this Ligand-Target Pair