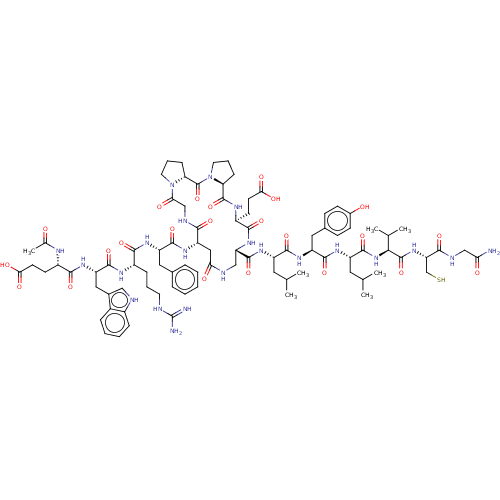
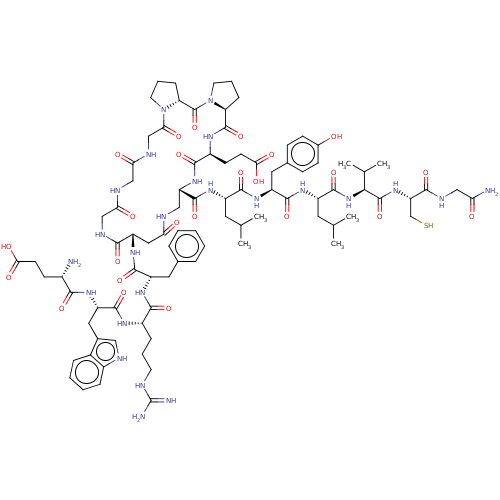
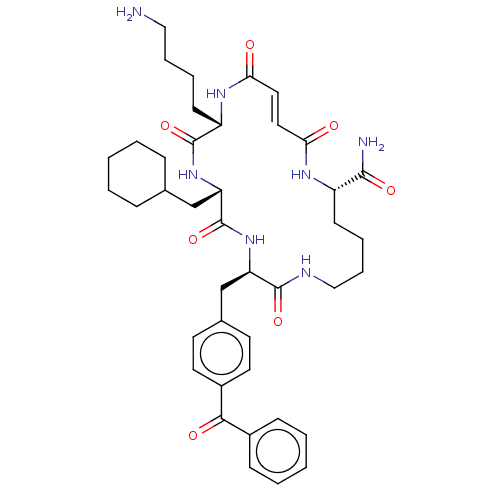
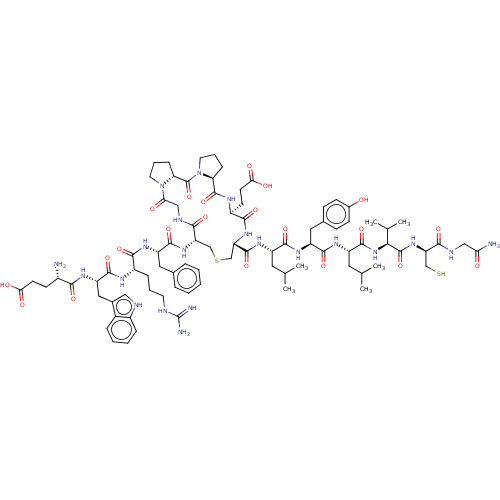
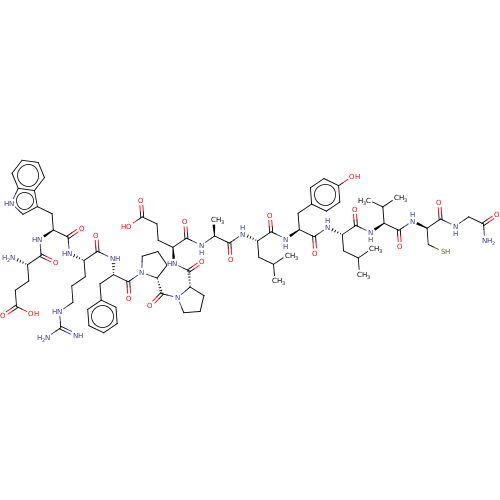
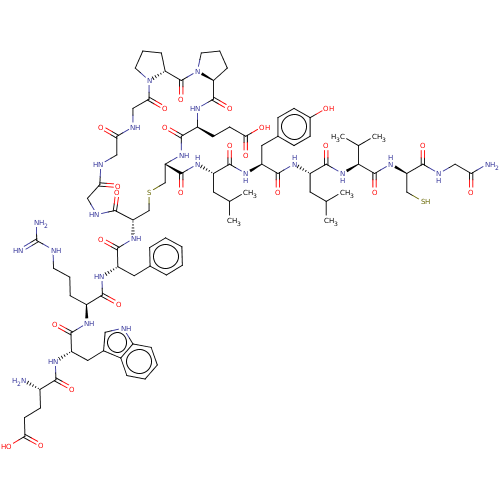
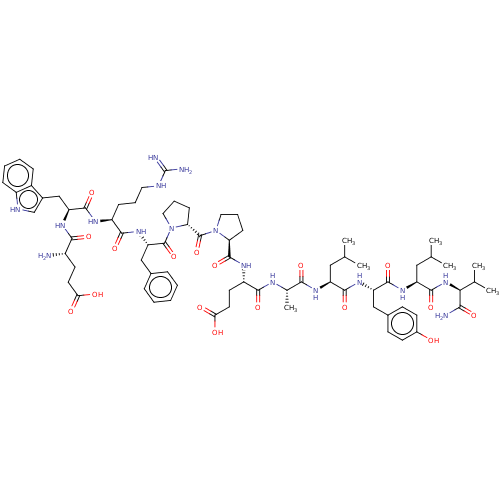
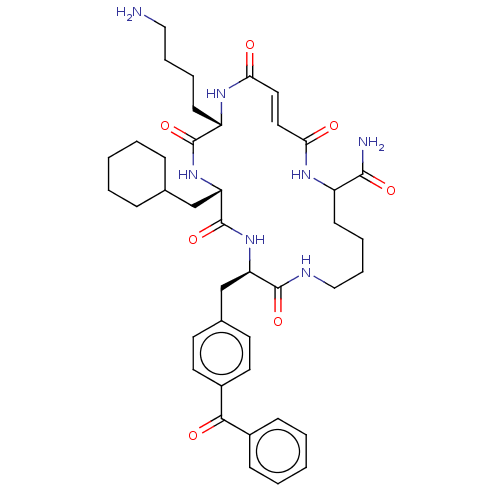
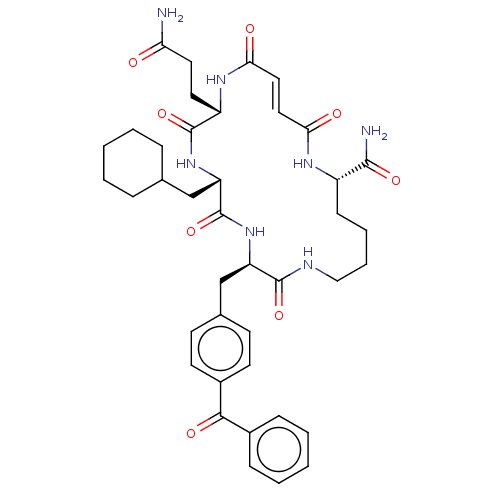
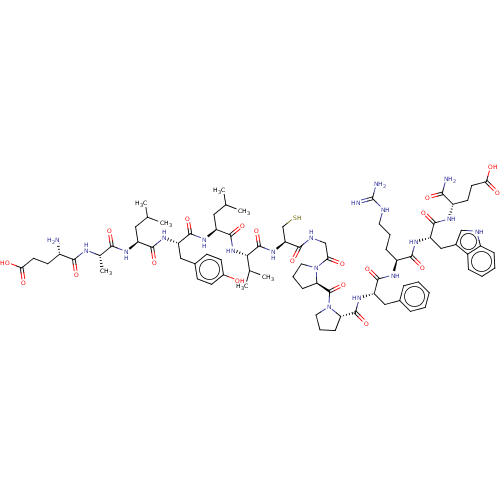
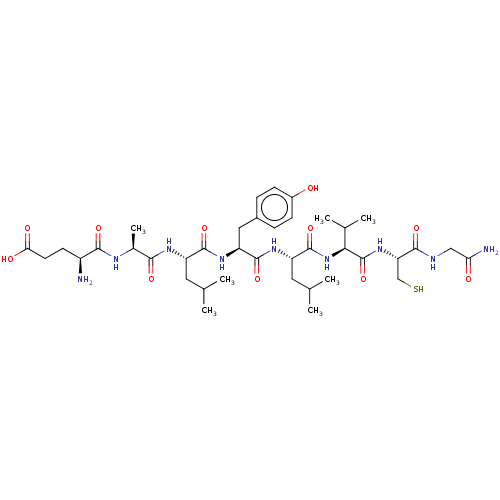
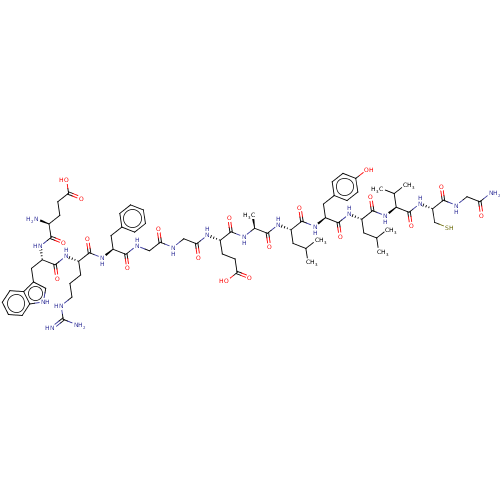
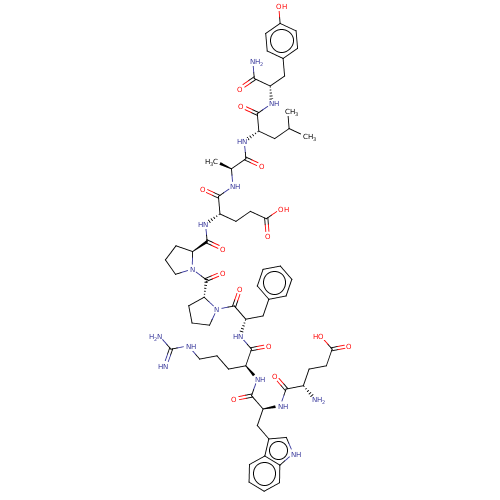
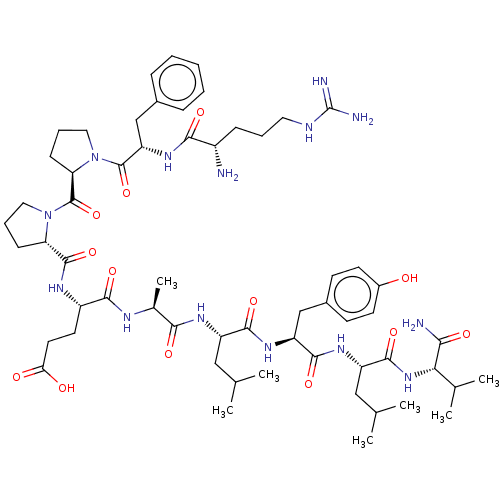
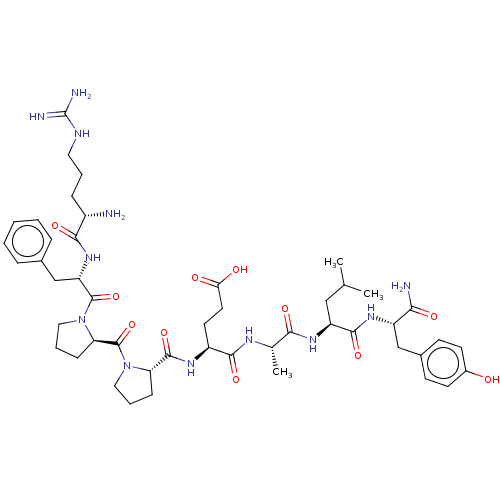
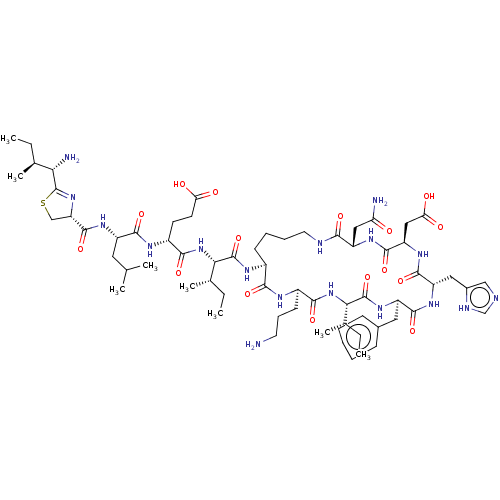
Affinity DataIC50: 22nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 44nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of N-terminal His6-tagged mouse recombinant IDE Escherichia coli BL21 (DE3) cells using fluorogenic peptide Mca-RPPGFSAFK(Dnp)-OH as subst...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 74nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 86nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMAssay Description:Macrocycles were assayed for IDE inhibition by monitoring cleavage of a substrate peptide containing a fluorophore-quencher pair such that cleavage o...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMAssay Description:Macrocycles were assayed for IDE inhibition by monitoring cleavage of a substrate peptide containing a fluorophore-quencher pair such that cleavage o...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 601nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of mouse His-tagged IDE (42 to 1019 residues) using fluorophore/quencher-tagged RPPGFSAFK(Dnp)-OH peptide as substrate preincubated for 10...More data for this Ligand-Target Pair