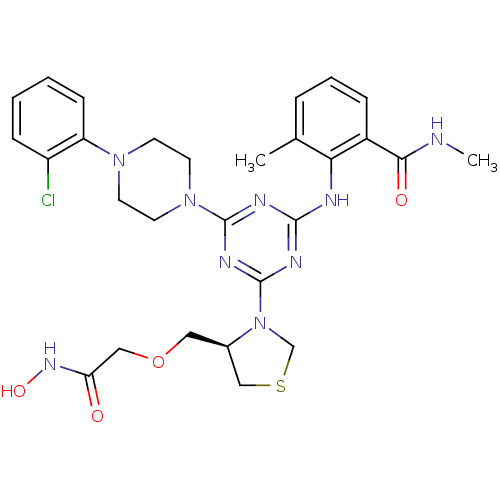
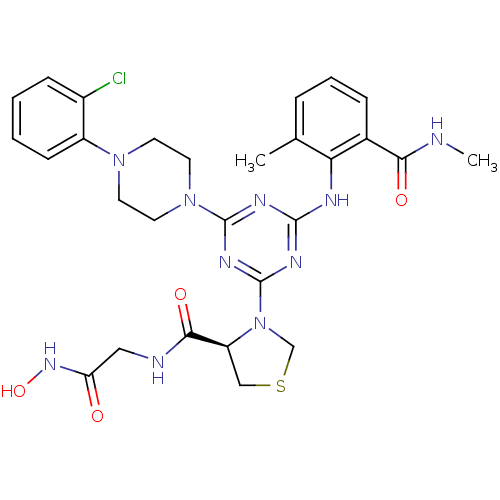
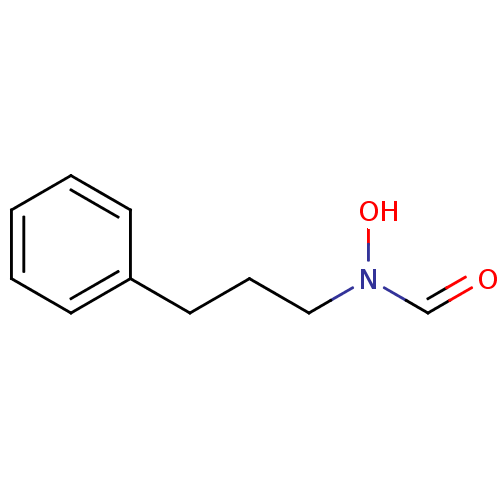
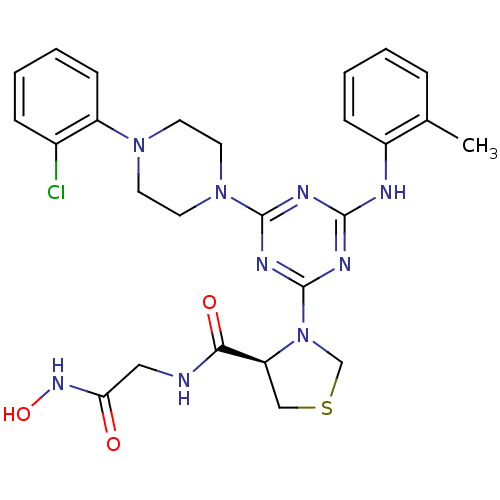
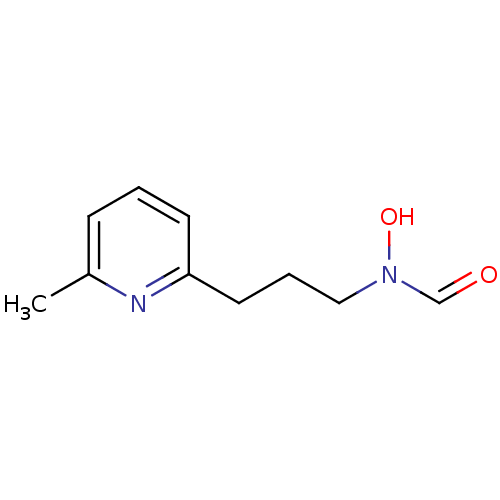
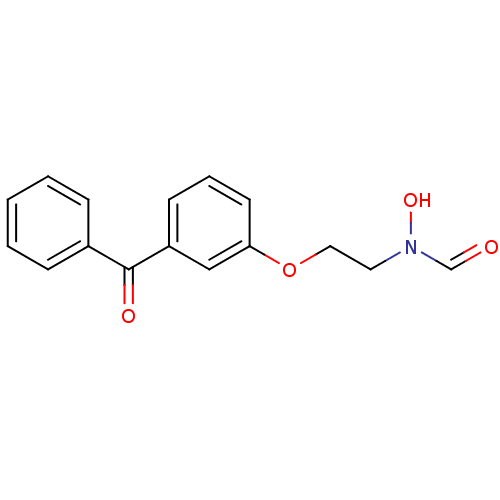
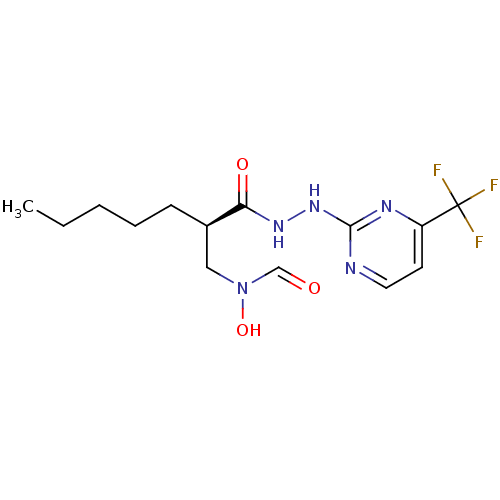
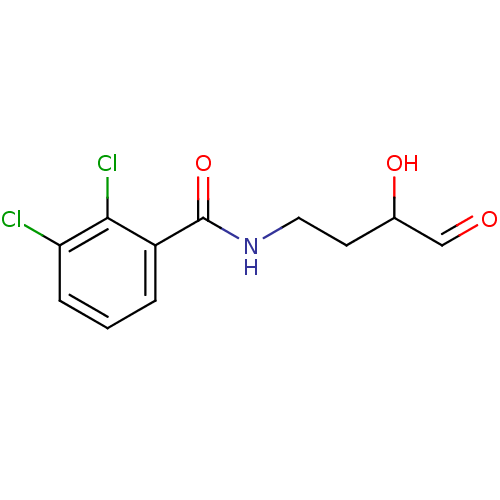
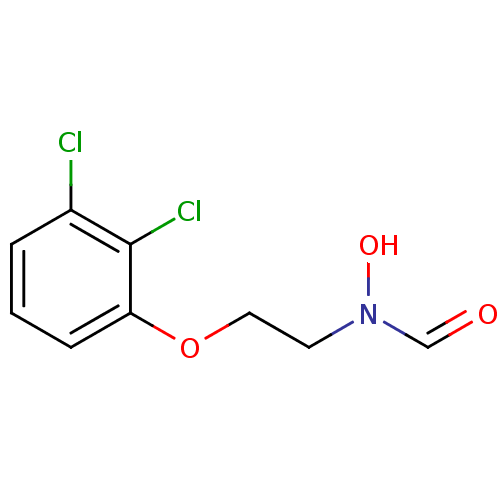
Affinity DataKi: 53.7nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
Affinity DataKi: 117nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
Affinity DataKi: 213nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
Affinity DataKi: 334nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
Affinity DataIC50: 400nMpH: 7.6 T: 2°CAssay Description:Enzyme assay was carried out in 96-well microtiter plates with a SpectraMax plate reader (Molecular Devices Corp.). PDF activity was measured using a...More data for this Ligand-Target Pair
Affinity DataKi: 650nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMpH: 7.6 T: 2°CAssay Description:Enzyme assay was carried out in 96-well microtiter plates with a SpectraMax plate reader (Molecular Devices Corp.). PDF activity was measured using a...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+3nMpH: 7.6 T: 2°CAssay Description:Enzyme assay was carried out in 96-well microtiter plates with a SpectraMax plate reader (Molecular Devices Corp.). PDF activity was measured using a...More data for this Ligand-Target Pair
Affinity DataKi: 6.31E+3nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
Affinity DataKi: 2.59E+4nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
Affinity DataKi: 3.17E+4nMAssay Description:Binding assay using PDF inhibitors.More data for this Ligand-Target Pair
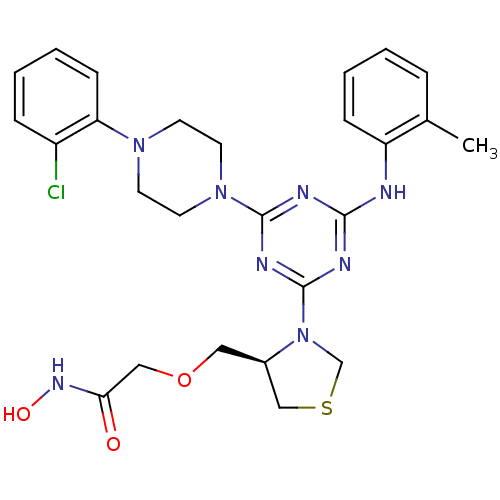
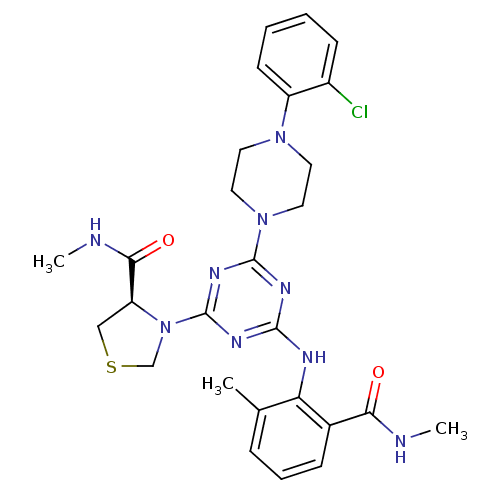
 3D Structure (crystal)
3D Structure (crystal)