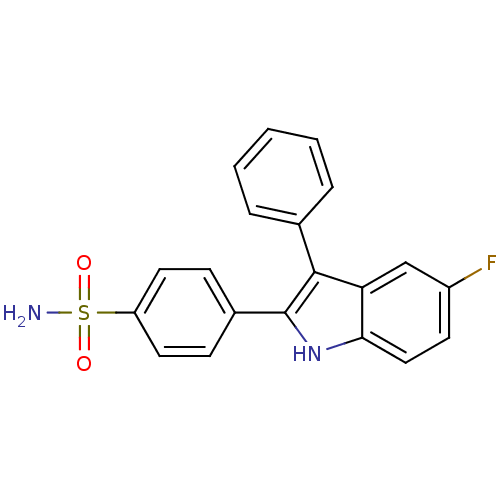
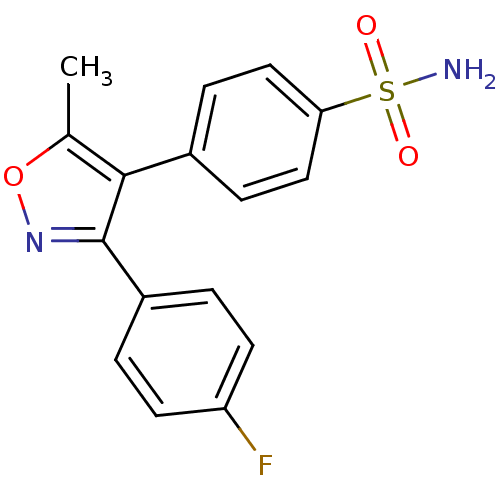
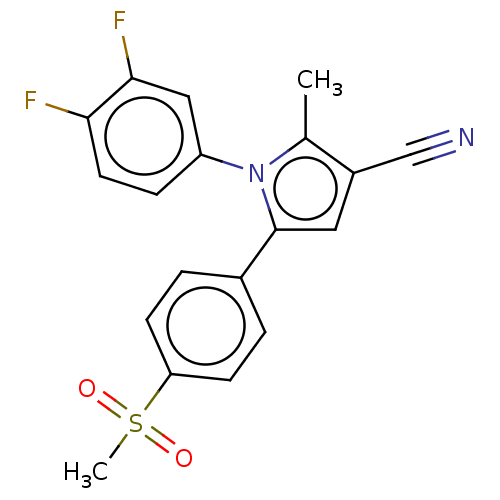
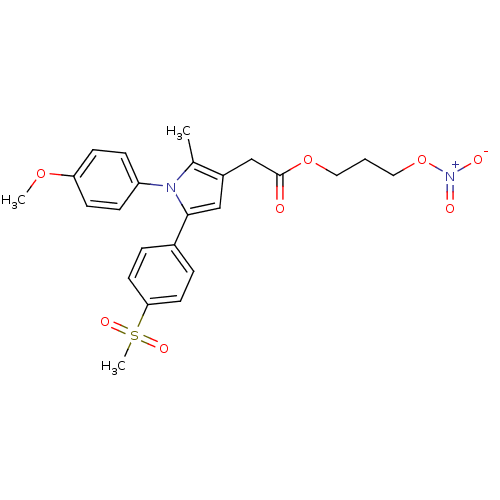
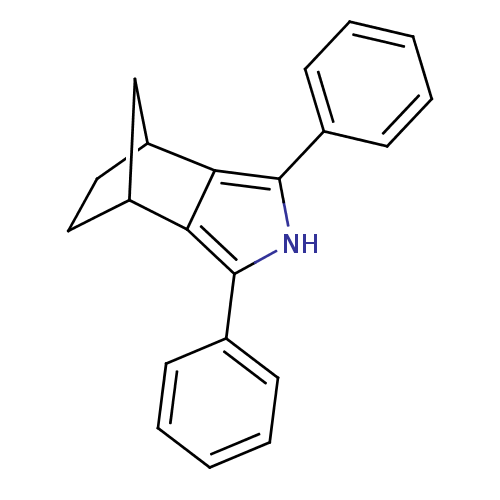
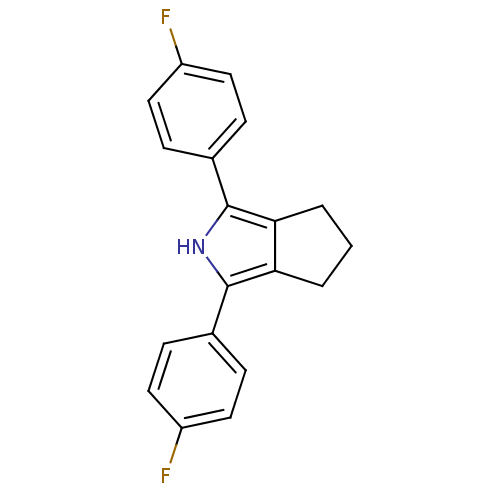
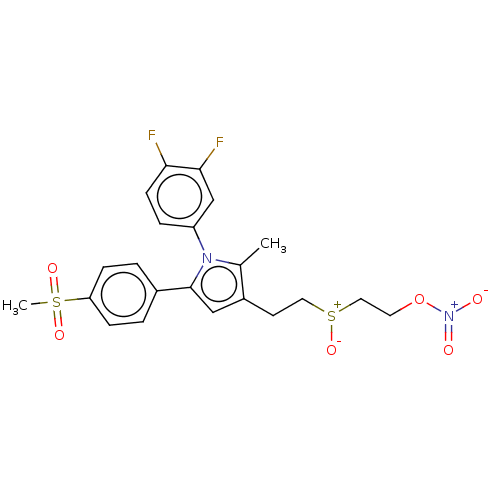
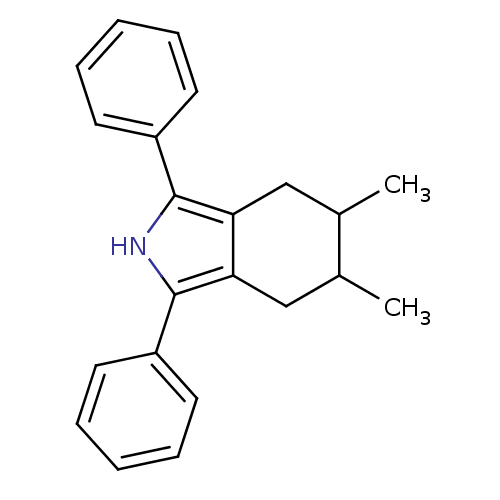
Affinity DataIC50: 0.00603nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
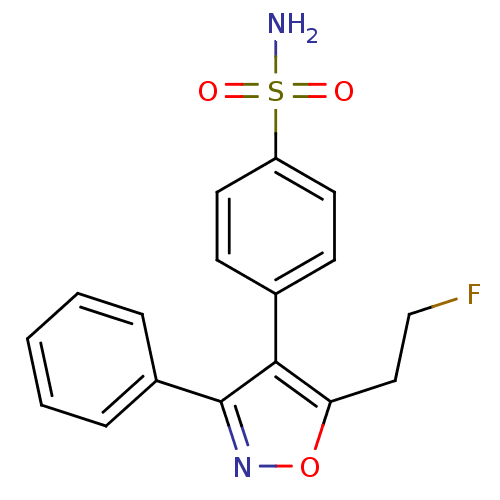
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
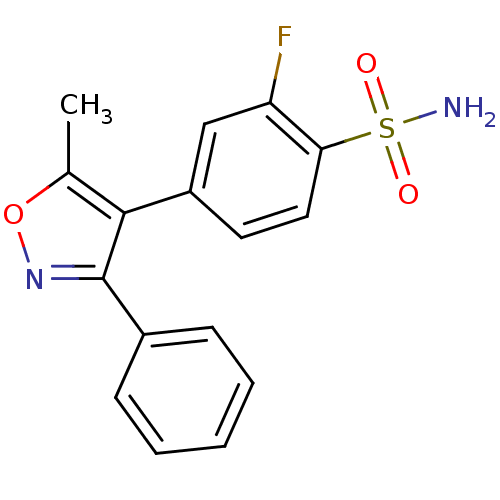
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
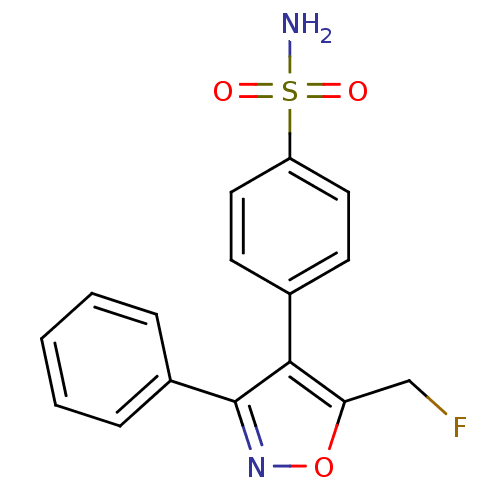
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0692nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0891nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0891nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.141nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.170nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.178nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.282nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.363nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.372nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.510nMAssay Description:Inhibition of LPS-induced COX2 activity in C57BL/6J mouse peritoneal macrophages by RIAMore data for this Ligand-Target Pair
Affinity DataIC50: 0.537nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.601nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 0.603nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.701nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 0.794nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:In vivo inhibition of COX2 in po dosed C57BL/6J mouse model of spontaneous GI-tract tumor Apc-min assessed as chemopreventive effect by measuring red...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 1.67nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMAssay Description:In vivo inhibition of COX2 in po dosed C57BL/6J mouse model of spontaneous GI-tract tumor Apc-min assessed as chemopreventive effect by measuring red...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibitory activity against murine Prostaglandin G/H synthase 2.More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of COX2 in Mus musculus (mouse) peritoneal macrophageMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Tested for prostaglandin E2 production as a function of COX-2 inhibition using endotoxin-treated murine RAW 264.7 macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Tested for prostaglandin E2 production as a function of COX-2 inhibition using endotoxin-treated murine RAW 264.7 macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Tested for prostaglandin E2 production as a function of COX-2 inhibition using endotoxin-treated murine RAW 264.7 macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Tested for prostaglandin E2 production as a function of COX-2 inhibition using endotoxin-treated murine RAW 264.7 macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition of COX2 in mouse J774 cells assessed as inhibition of LPS-induced PGE2 production by radioimmunoassayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40nMAssay Description:Inhibition of COX-2-mediated PGE2 production in LPS-stimulated mouse J774 cells after 24 hrs by radioimmunoassayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90nMAssay Description:Inhibition of mouse macrophage COX2More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of COX2 in LPS-stimulated mouse J774 cells assessed as inhibition of PGE2 production and measured after 24 hrs by enzyme immuno-assay (EIA...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMAssay Description:Inhibition against prostaglandin G/H synthase 2 from mouse resident macrophagesMore data for this Ligand-Target Pair
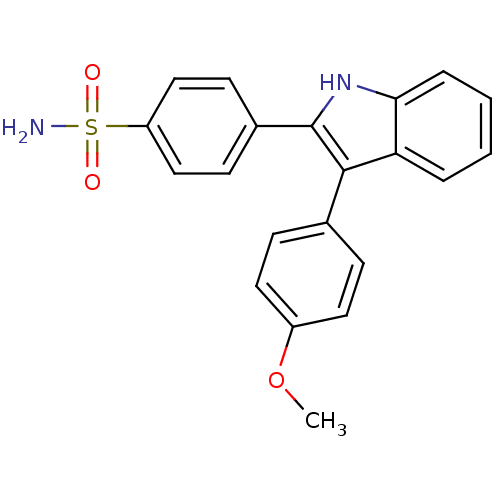
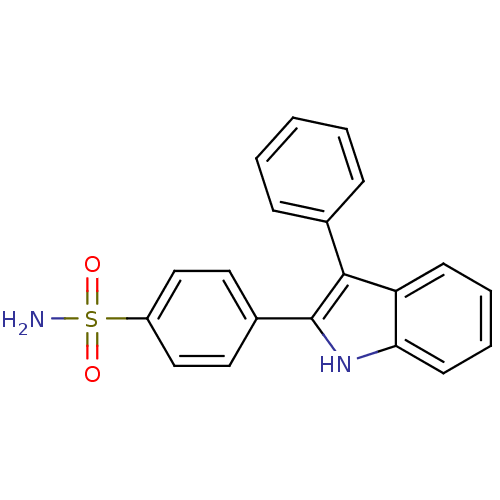
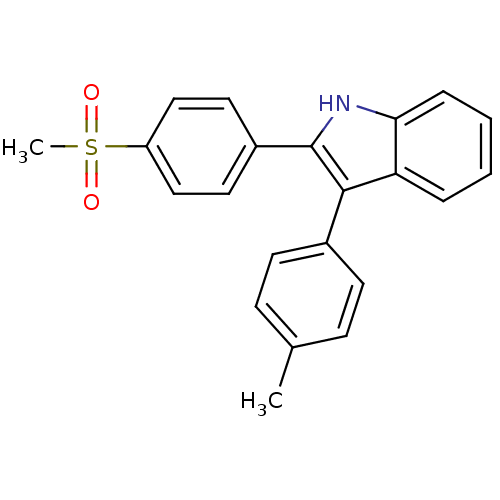
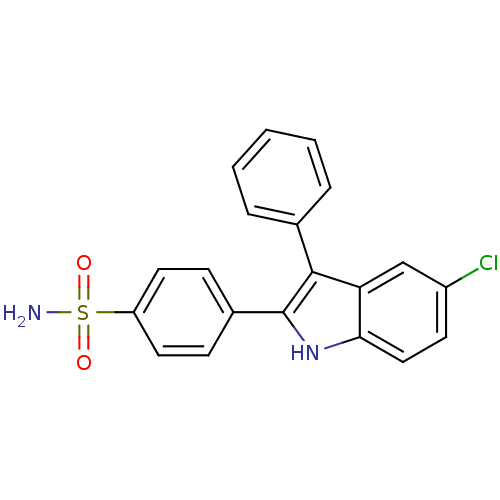
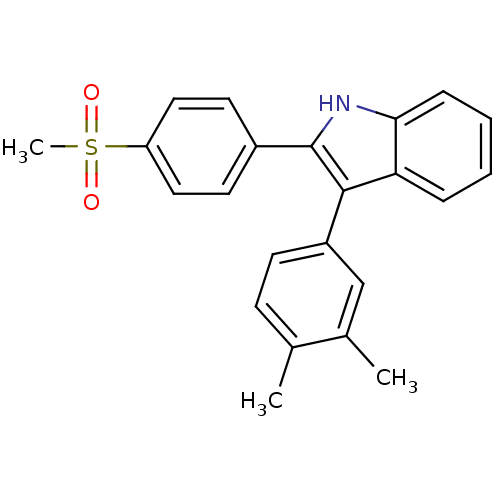
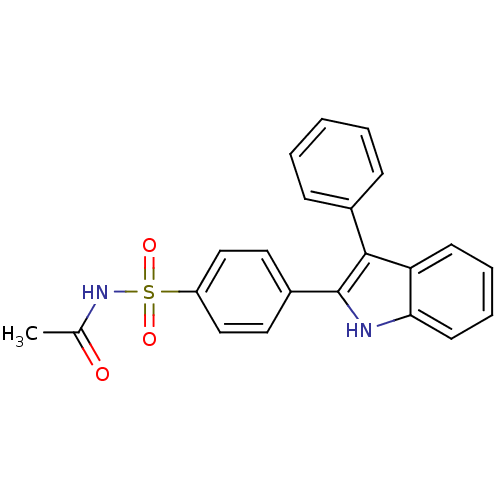
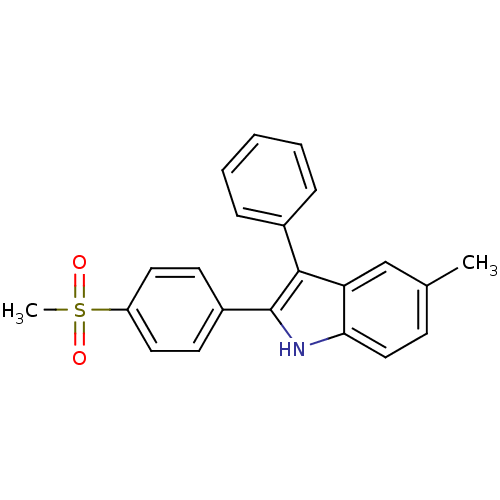
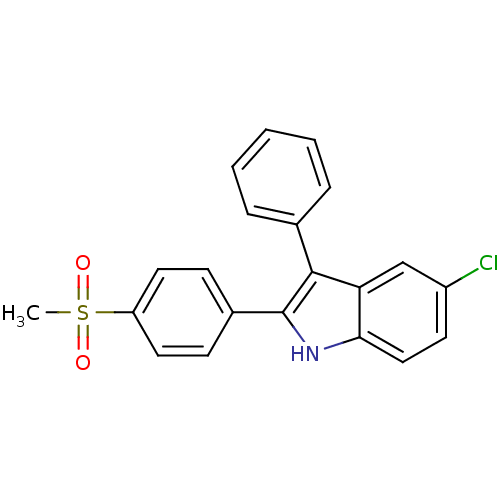
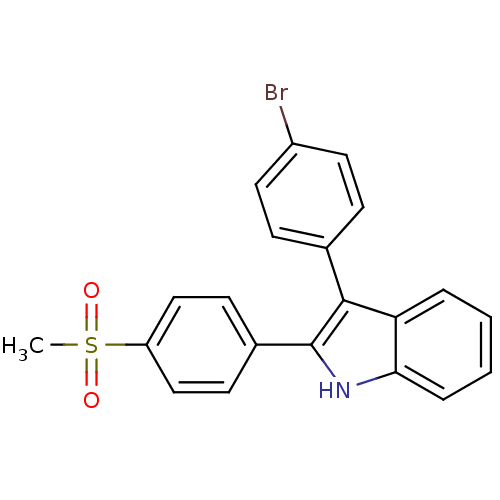
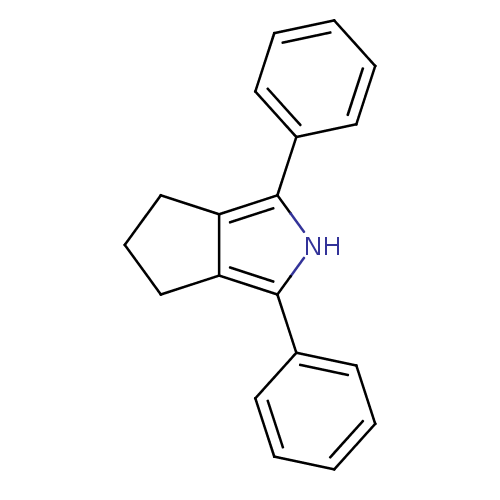
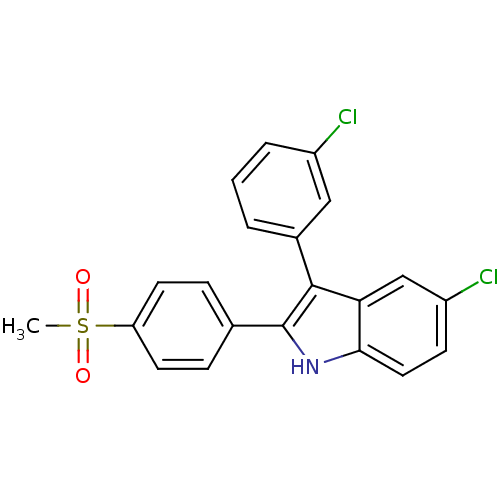
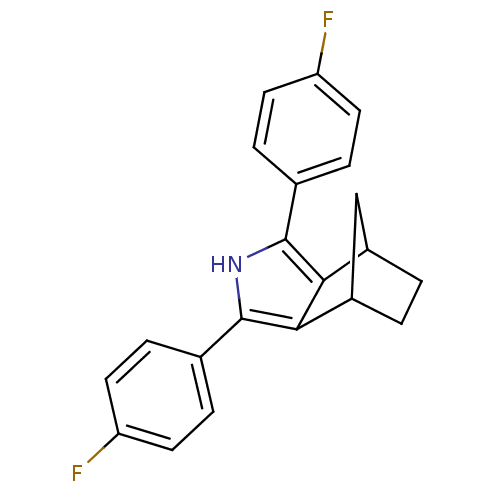
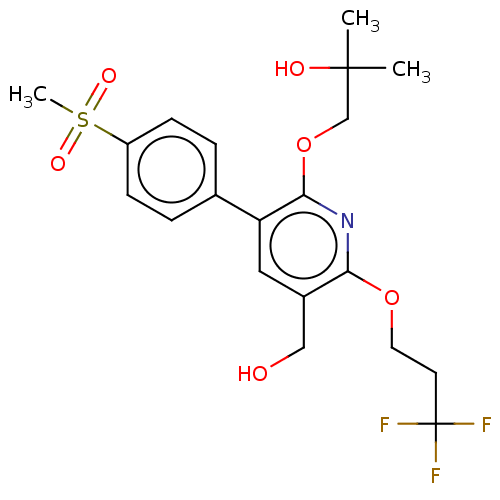
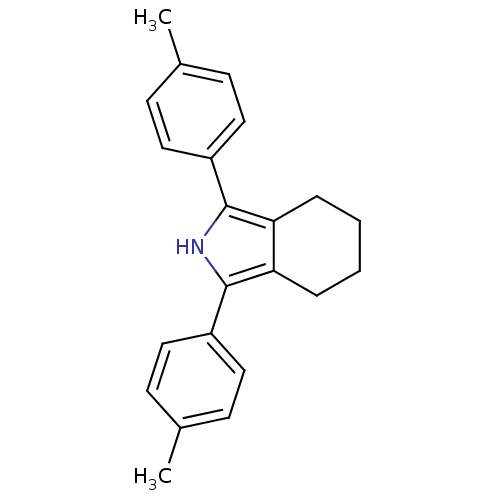
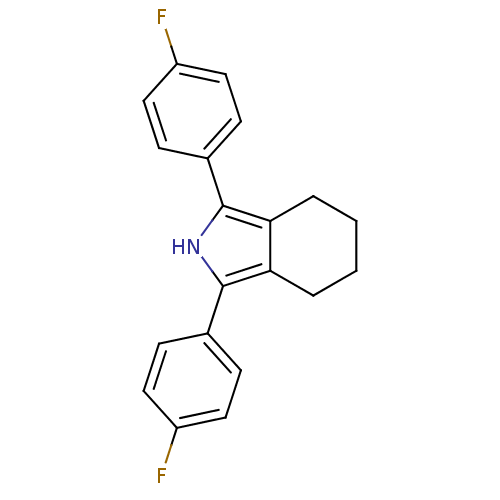
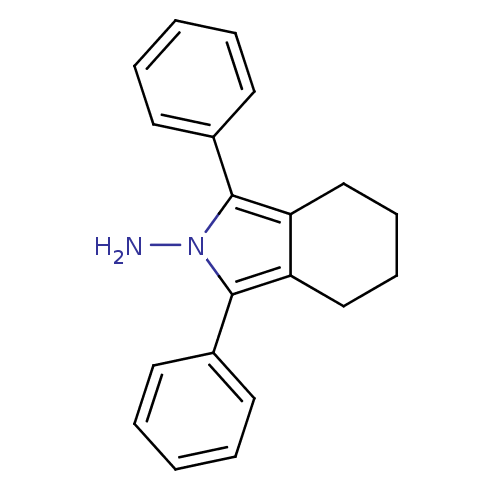
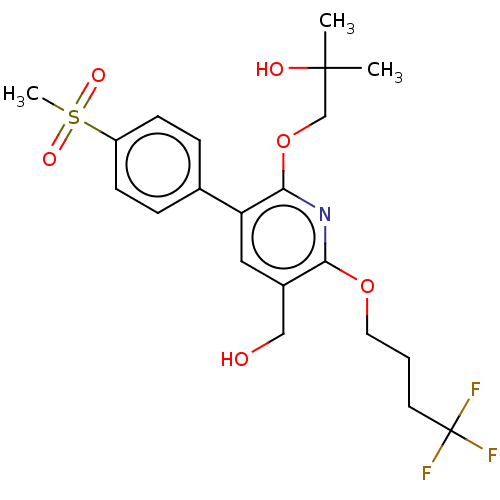
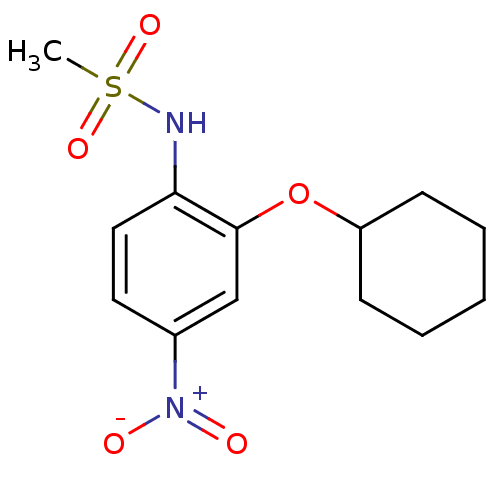
 3D Structure (crystal)
3D Structure (crystal)