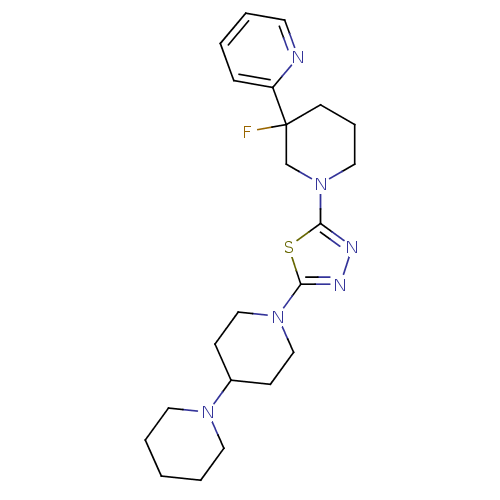
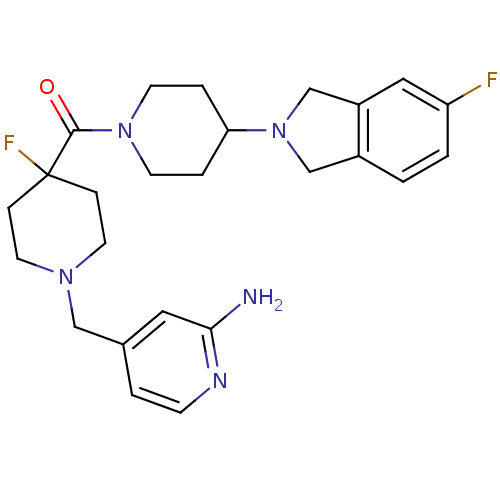
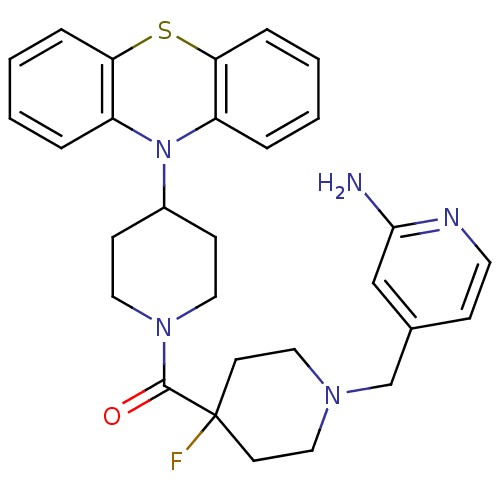
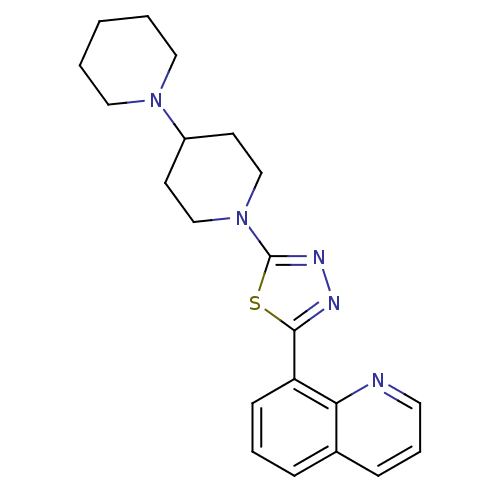
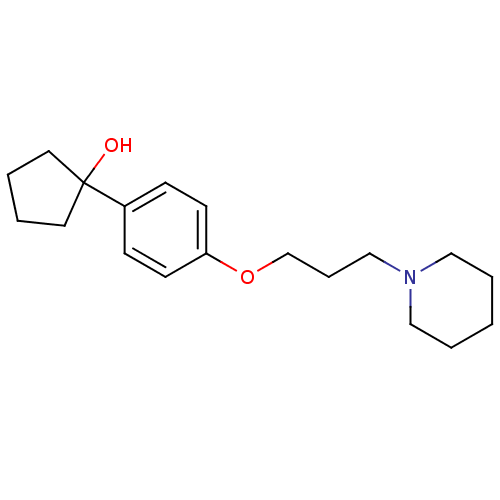
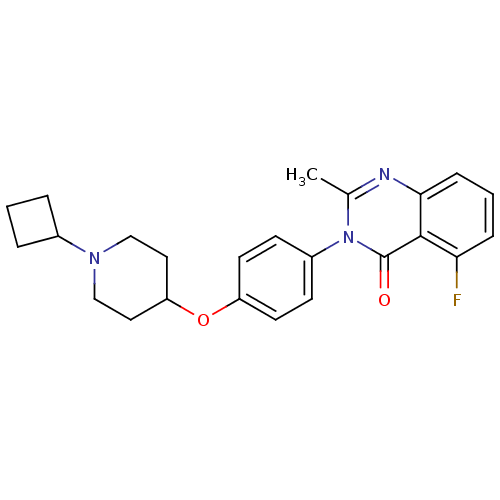
Affinity DataEC50: 0.100nMAssay Description:Agonist activity at mouse H3R expressed on HEK293T cells assessed as inhibition of forskolin-induced CRE-driven luciferase activity co-incubated with...More data for this Ligand-Target Pair
Affinity DataKi: 0.280nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.330nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.520nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:Displacement of 3-([1,1,1-3H]methyl)-2-(4-{[3-(1-pyrrolidinyl)propyl]oxy}phenyl)-4(3H)-quinazolinone from mouse histamine H3 receptor expressed in HE...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Antagonist activity against mouse histamine H3 receptor expressed in human HT1080 cells assessed as inhibition of (R)-alpha-methylhistamine-induced i...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]NAMH from mouse H3R expressed in HEK293T cells incubated for 2 hrs by microbeta scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [125I]-iodoproxyfan from histamine H3 receptor in mouse brain cortex after 1 hr by gamma counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataEC50: 1.70nMAssay Description:Inverse agonist activity at mouse histamine H3 receptor overexpressed in HEK293 cells by [35S]GTPgammaS binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Binding affinity to mouse histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of 3-([1,1,1-3H]methyl)-2-(4-{[3-(1-pyrrolidinyl)propyl]oxy}phenyl)-4(3H)-quinazolinone from mouse histamine H3 receptor expressed in HE...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 2.30nMAssay Description:Displacement of [125I]Iodoproxyfan from mouse brain cortex histamine H3 receptor after 60 minsMore data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Displacement of 3-([1,1,1-3H]methyl)-2-(4-{[3-(1-pyrrolidinyl)propyl]oxy}phenyl)-4(3H)-quinazolinone from mouse histamine H3 receptor expressed in HE...More data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Displacement of [125I]-iodoproxyfan from histamine H3 receptor in mouse brain cortex after 1 hr by gamma counting analysisMore data for this Ligand-Target Pair
Affinity DataKd: 2.60nMAssay Description:Binding affinity to mouse histamine H3 receptor overexpressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.60nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKd: 3.10nMAssay Description:Binding affinity to histamine H3 receptor in wild type mouse brain membraneMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse recombinant histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse recombinant histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from mouse recombinant histamine H3 receptor after 30 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from mouse recombinant histamine H3 receptor after 30 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Displacement of [125I]-iodoproxyfan from histamine H3 receptor in mouse brain cortex after 1 hr by gamma counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Displacement of 3-([1,1,1-3H]methyl)-2-(4-{[3-(1-pyrrolidinyl)propyl]oxy}phenyl)-4(3H)-quinazolinone from mouse histamine H3 receptor expressed in HE...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from mouse recombinant histamine H3 receptor after 30 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 6.30nMAssay Description:Displacement of [125I]-iodoproxyfan from histamine H3 receptor in mouse brain cortex after 1 hr by gamma counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Displacement of [125I]-iodoproxyfan from histamine H3 receptor in mouse brain cortex after 1 hr by gamma counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]-N-alpha-methylhistamine from mouse recombinant histamine H3 receptor after 30 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptor after 30 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Displacement of [125I]-iodoproxyfan from histamine H3 receptor in mouse brain cortex after 1 hr by gamma counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.40nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from mouse recombinant histamine H3 receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 9.60nMAssay Description:Inverse agonist activity at mouse histamine H3 receptor overexpressed in HEK293 cells by [35S]GTPgammaS binding assayMore data for this Ligand-Target Pair
Affinity DataEC50: 11nMAssay Description:Inverse agonist activity at mouse histamine H3 receptor overexpressed in HEK293 cells by [35S]GTPgammaS binding assayMore data for this Ligand-Target Pair
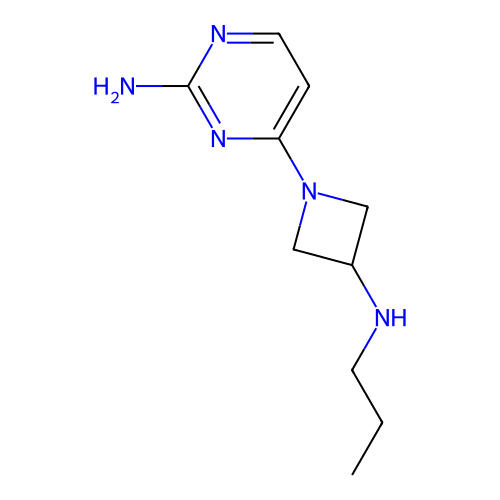
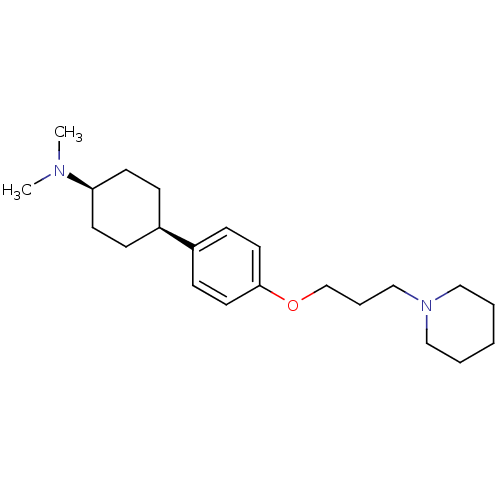
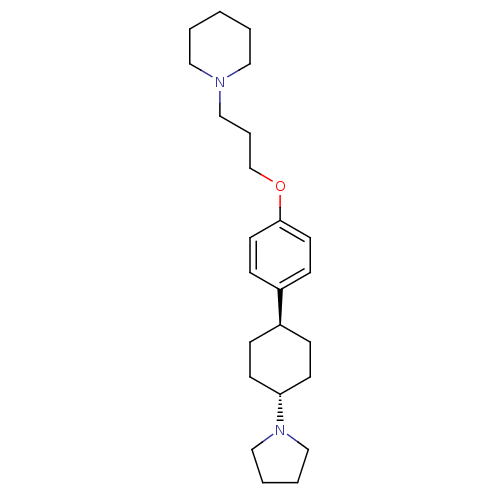
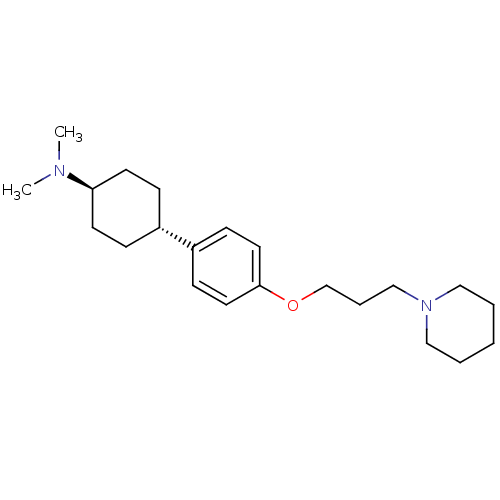
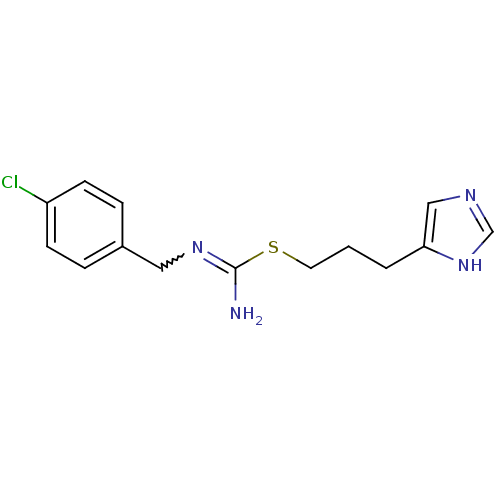
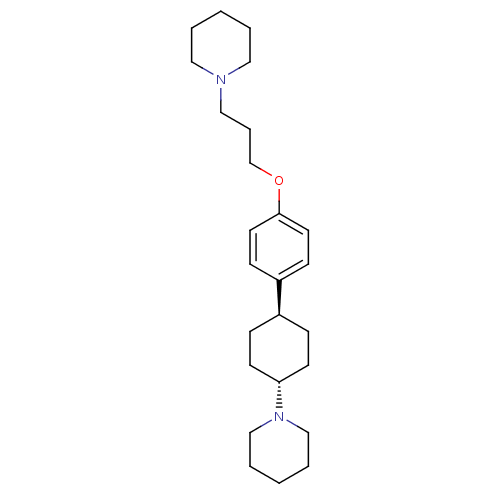
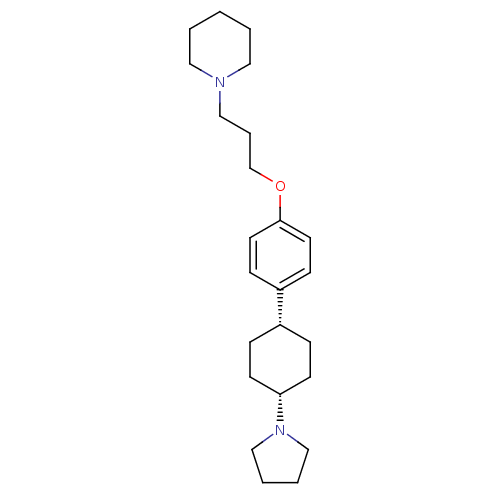
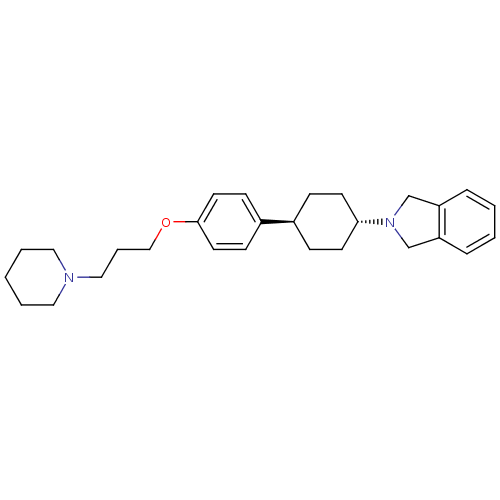
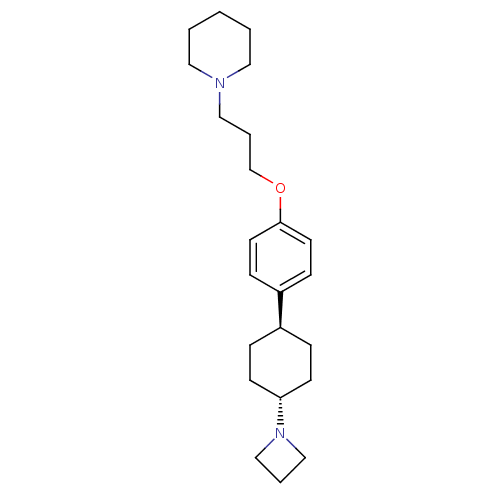
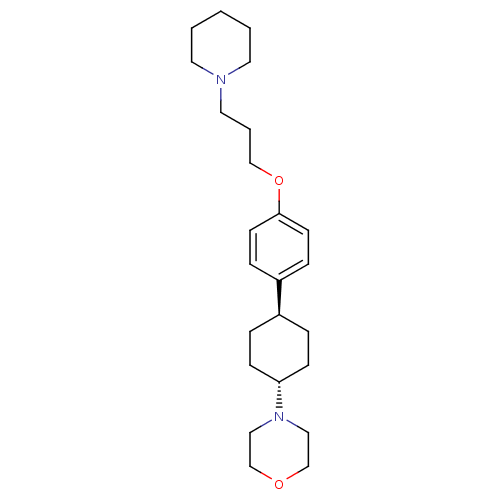
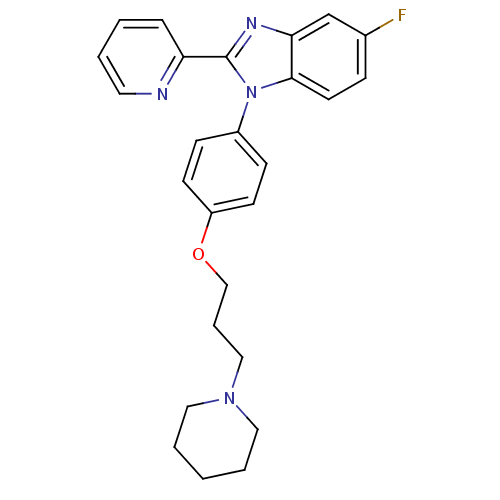
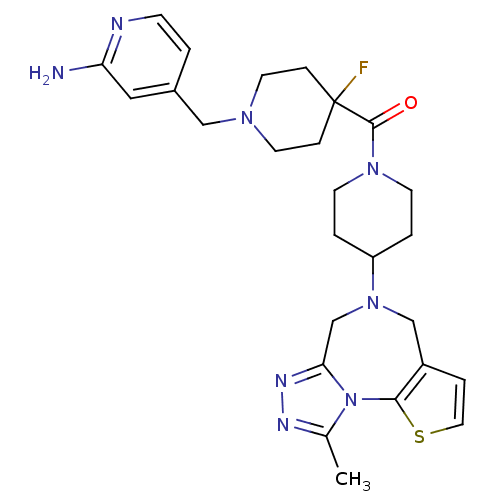
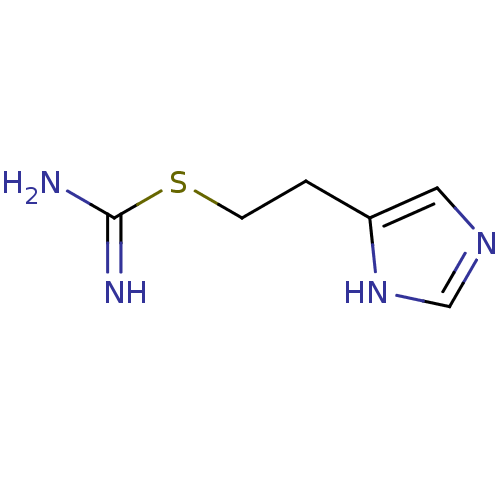
 3D Structure (crystal)
3D Structure (crystal)