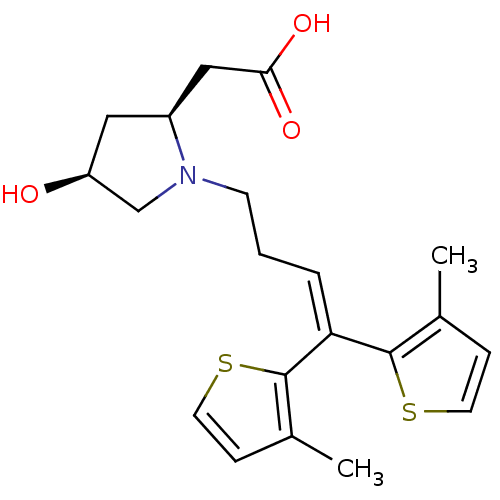
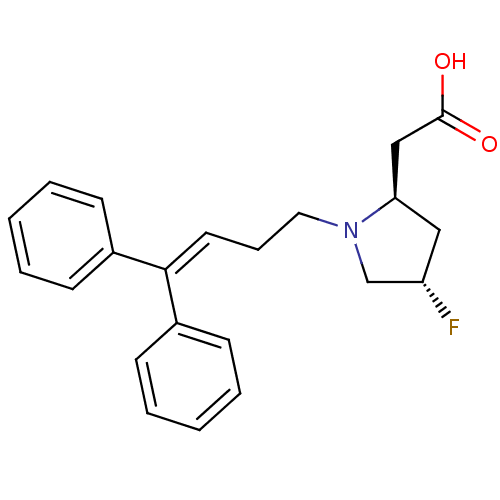
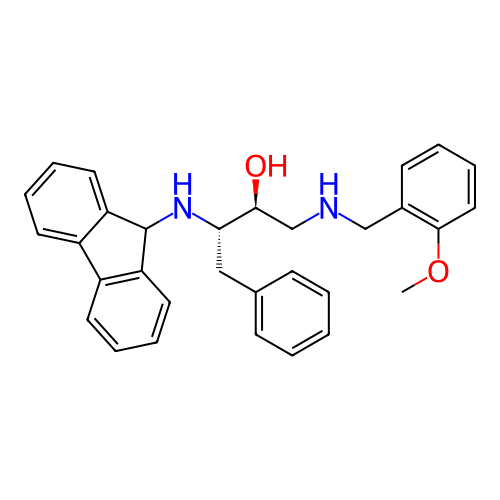
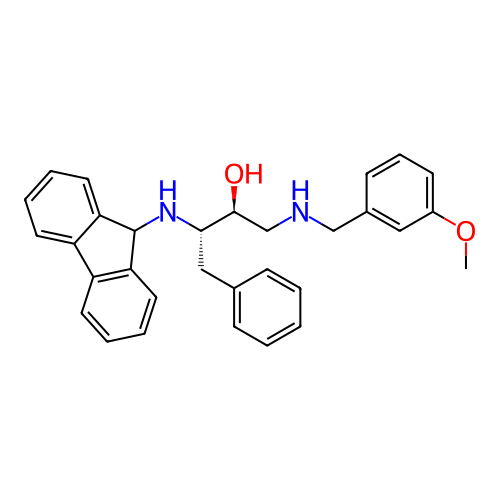
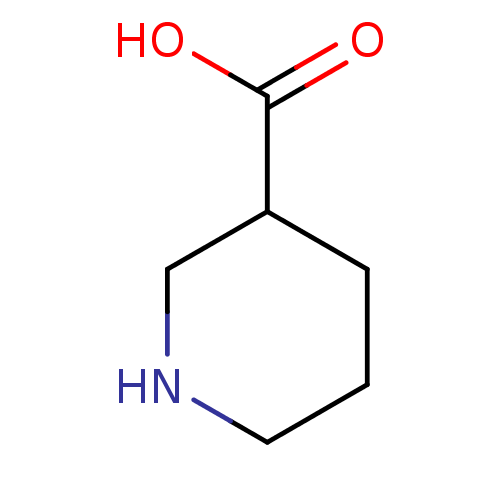
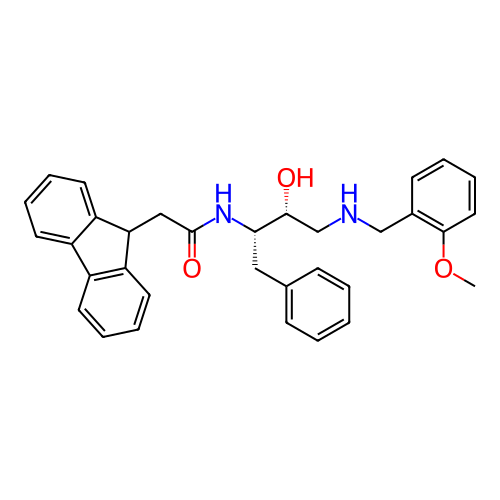
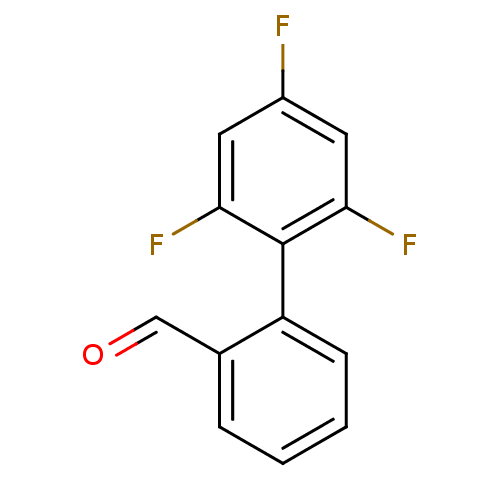
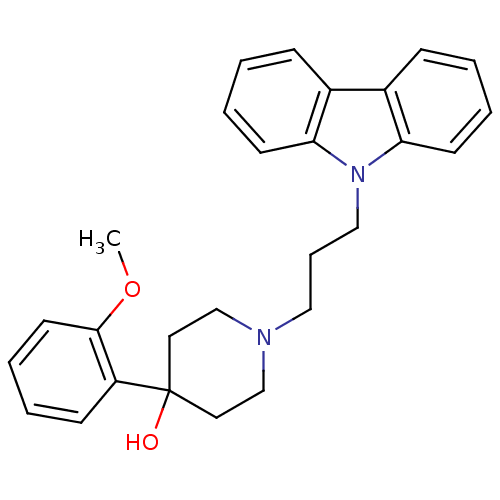
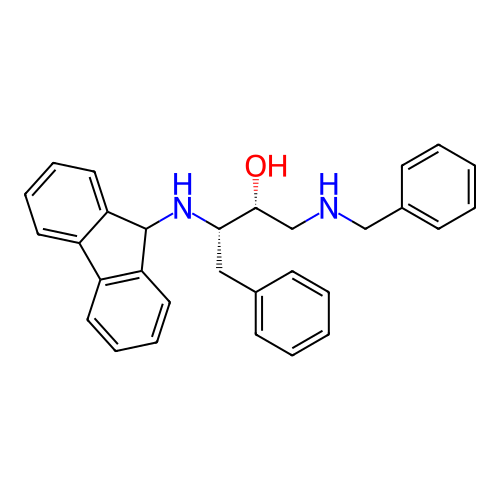
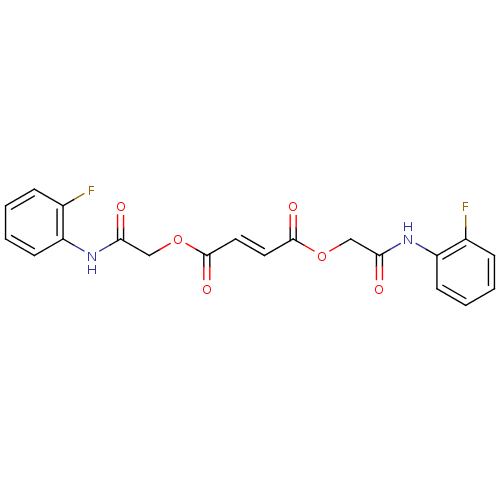
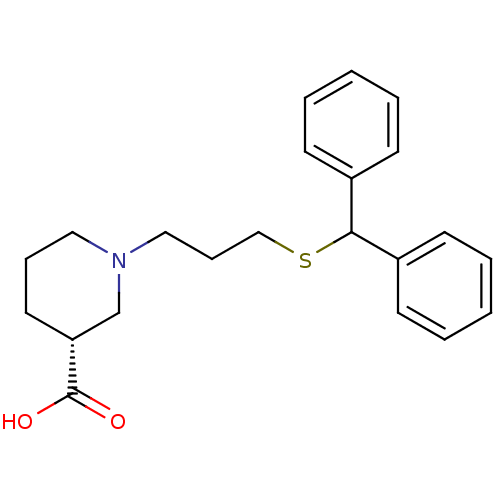
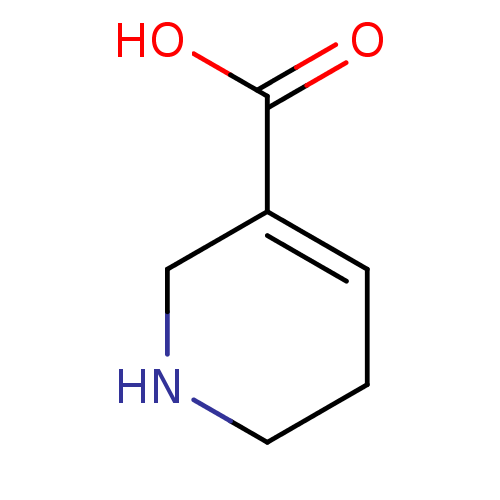
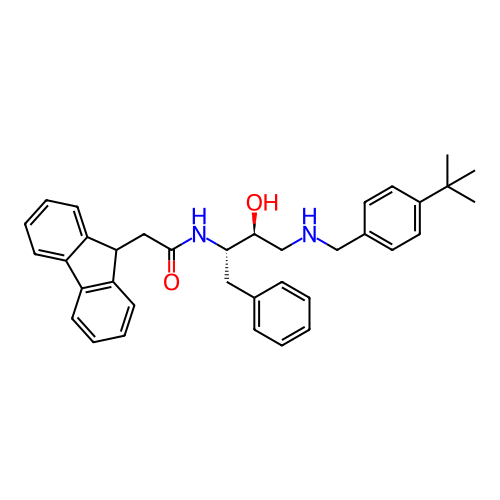
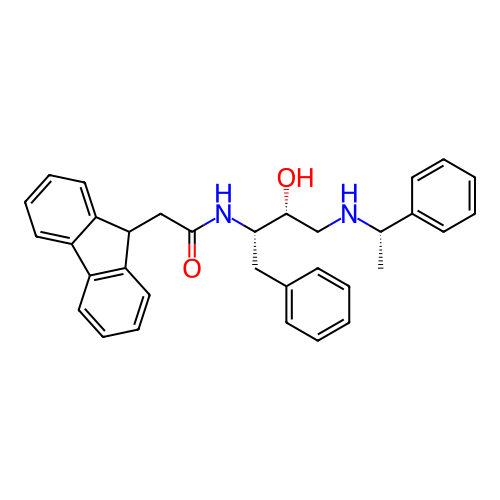
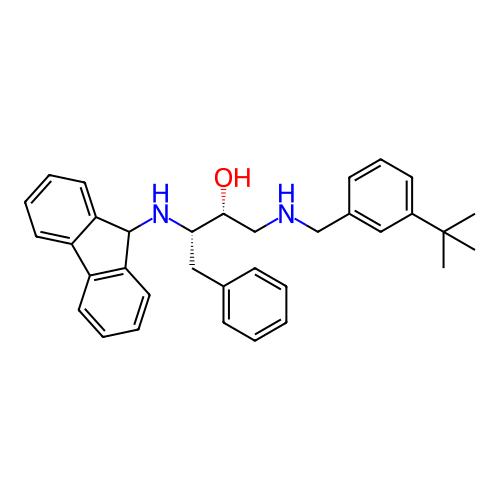TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 40nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
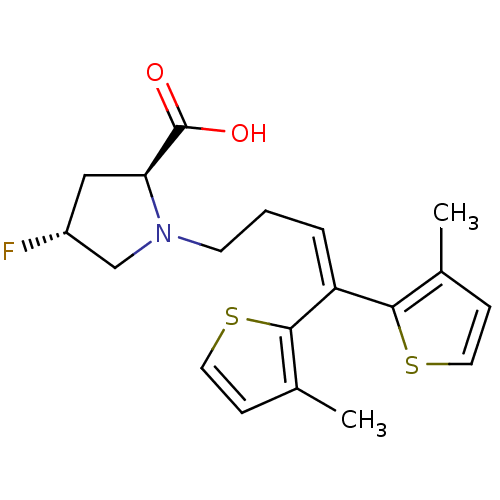
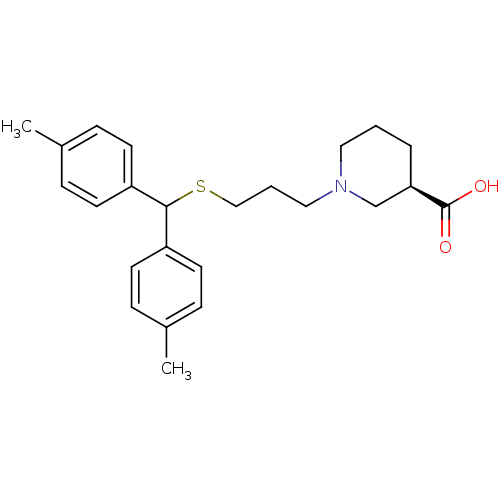
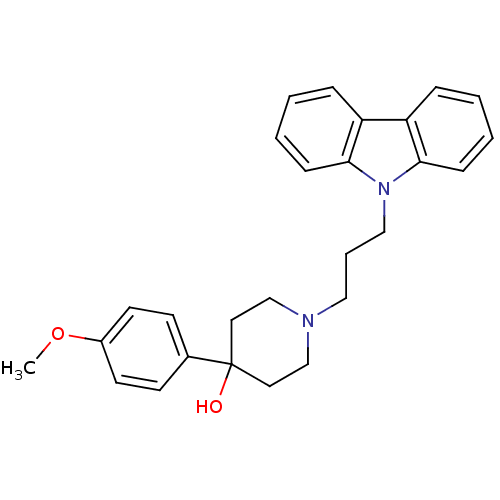
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 55nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
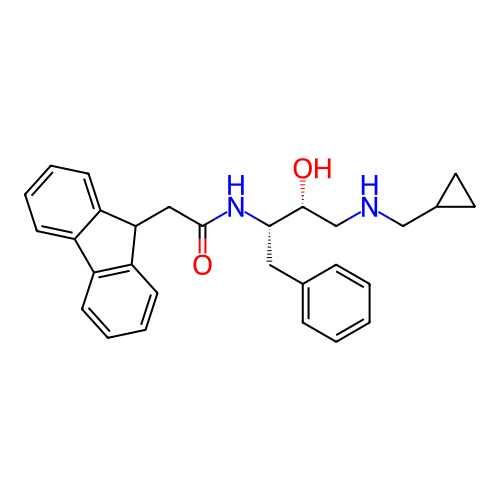
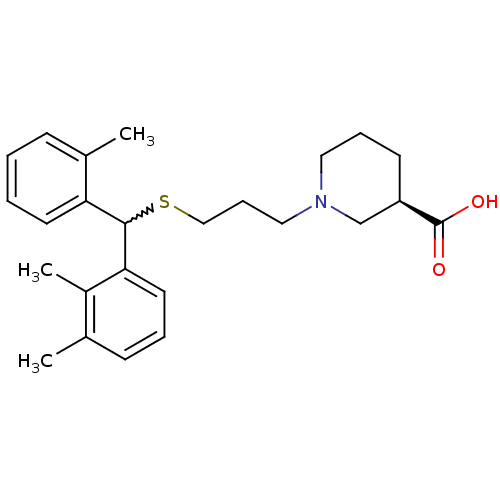
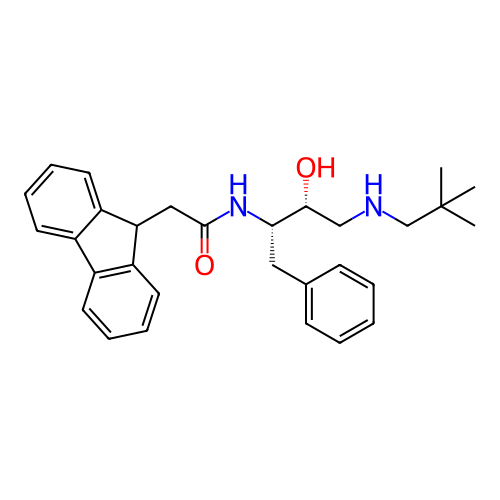
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 63.1nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
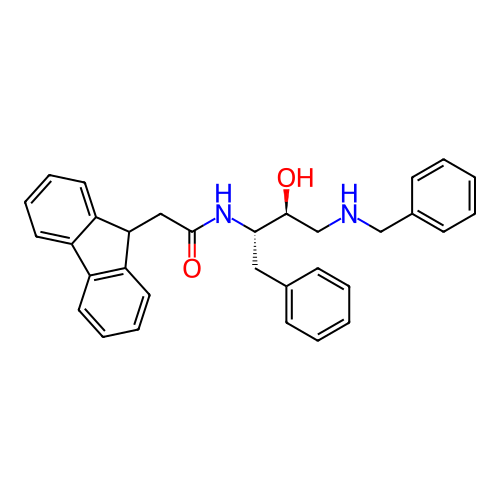
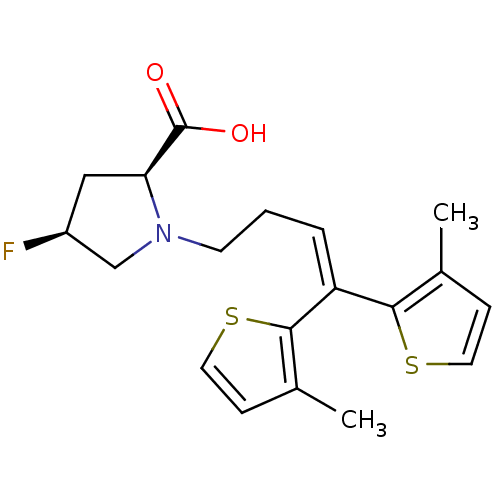
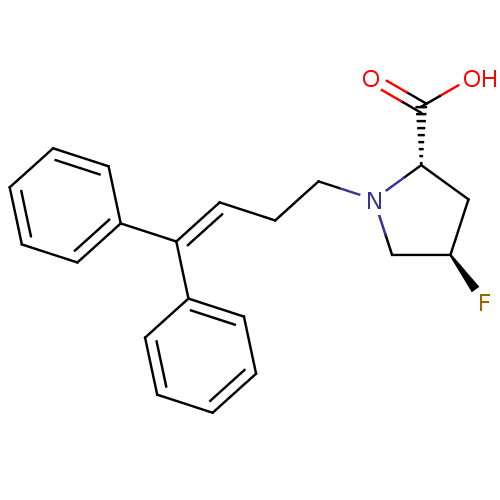
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in D8 cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 191nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 340nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 650nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 800nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 3 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 920nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in D8 cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.18E+3nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in D8 cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 2.56E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 2.97E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 3.15E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 3.29E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 3.96E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of mouse GAT1-mediated [3H]GABA uptake expressed in human HEK cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 4.92E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 4.94E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 5.14E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 5.81E+3nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 6.31E+3nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 3 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 7.08E+3nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 7.24E+3nMAssay Description:Inhibition of mouse GAT1-mediated [3H]GABA uptake expressed in human HEK cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 9.55E+3nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity to mouse GAT1 expressed in HEK293 cells membranes using LC-ESI-MS/MS analysis by [2H10]NO711 binding inhibition assayMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.02E+4nMAssay Description:Inhibition of mouse GAT1-mediated [3H]GABA uptake expressed in human HEK cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.10E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.10E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.16E+4nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in D8 cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in HEK293 cells after 25 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.26E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.28E+4nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.35E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.41E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.48E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.58E+4nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in D8 cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.69E+4nMAssay Description:Inhibition of [3H]GABA uptake at mouse GAT1 expressed in D8 cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.86E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.90E+4nMAssay Description:Inhibition of mouse GAT1-mediated [3H]GABA uptake expressed in human HEK cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 1.91E+4nMAssay Description:Inhibition of mouse GAT1-mediated [3H]GABA uptake expressed in BHK cellsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 4.79E+4nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 1(Mouse)
Ludwig Maximilians University At Munich
Curated by ChEMBL
Ludwig Maximilians University At Munich
Curated by ChEMBL
Affinity DataIC50: 6.74E+4nMAssay Description:Inhibition of GAT1 in mouse D8 cells assessed as [3H]GABA transport by liquid scintillation countingMore data for this Ligand-Target Pair
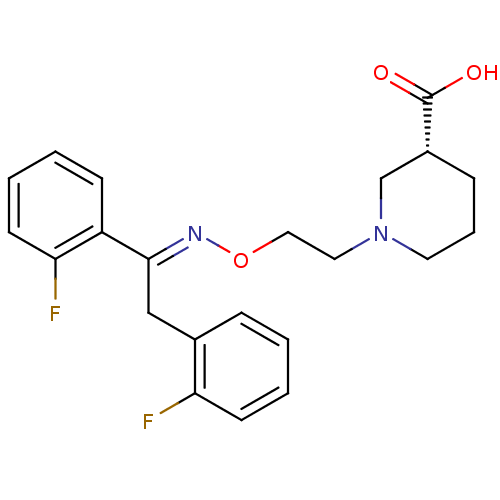
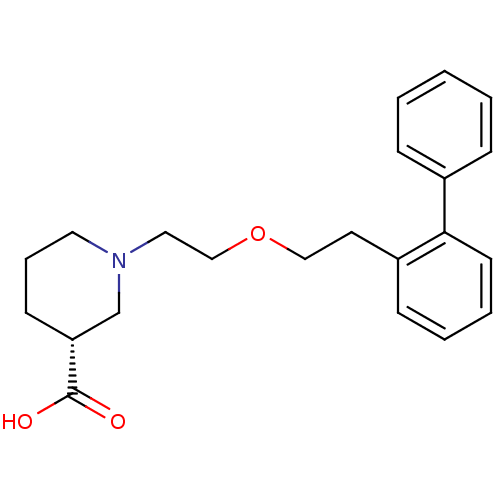
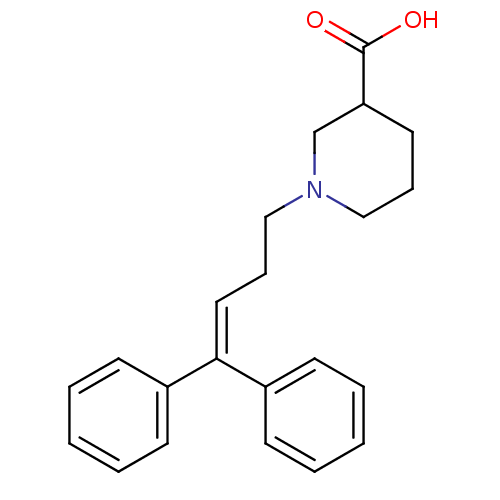
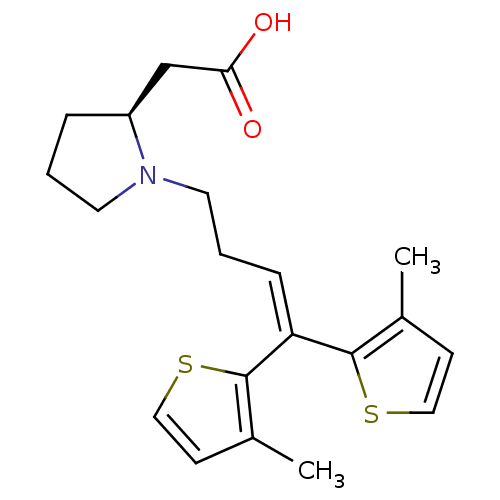
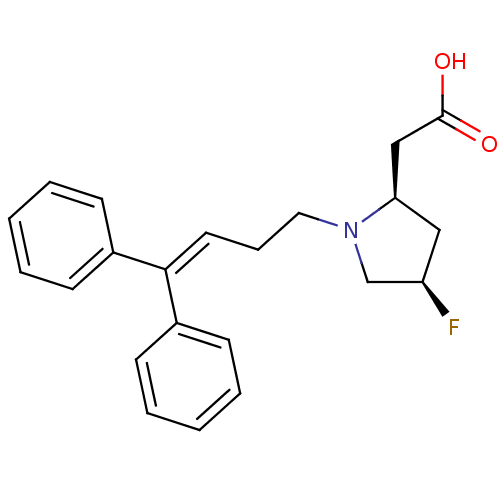
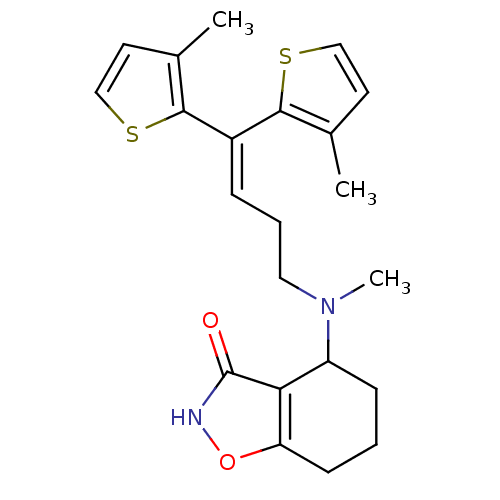
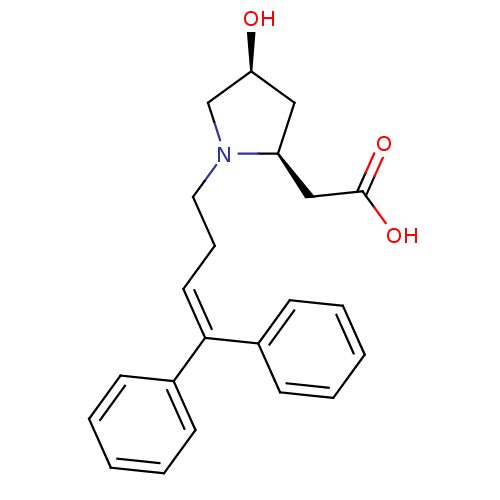
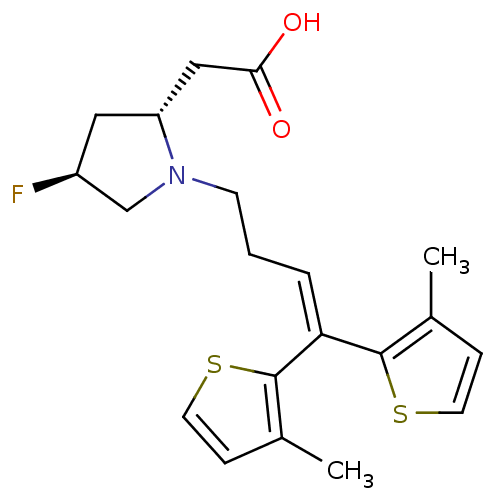
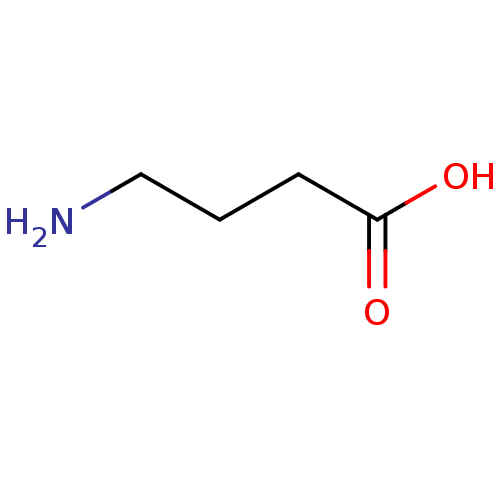
 3D Structure (crystal)
3D Structure (crystal)