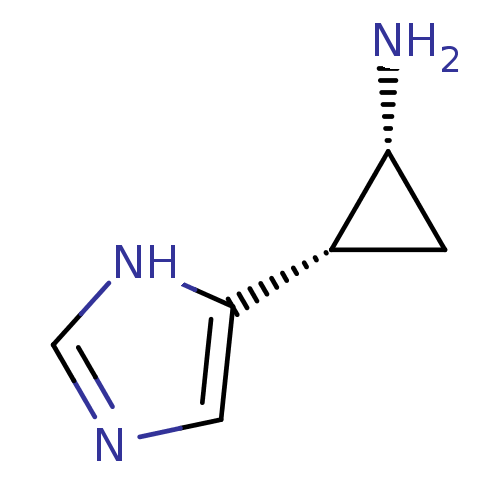
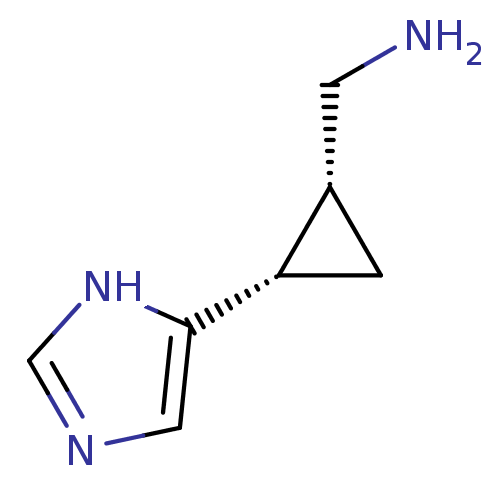
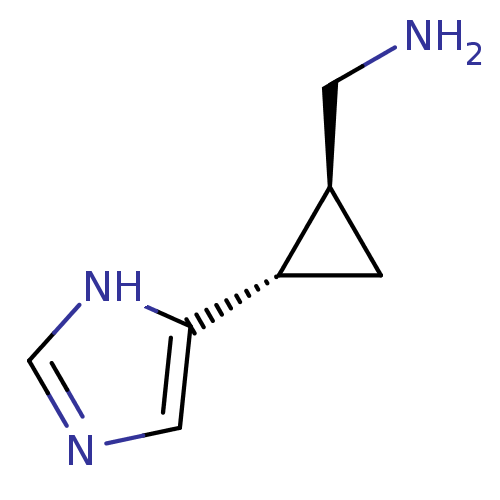
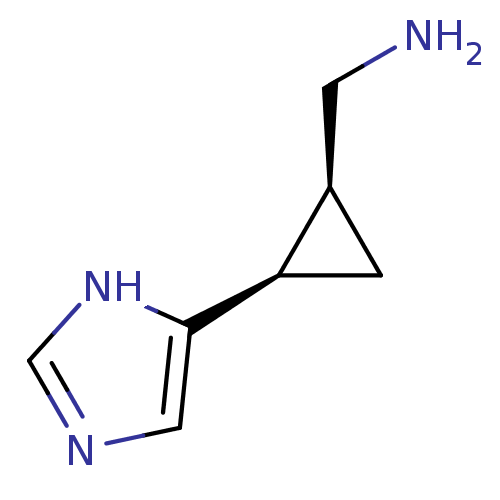
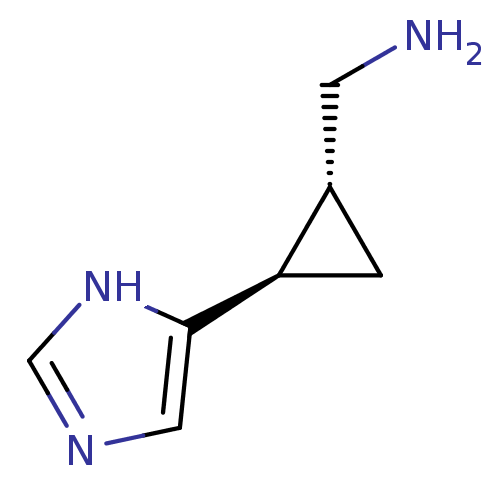
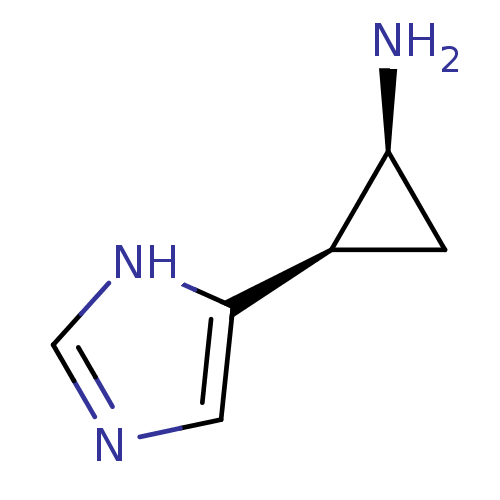
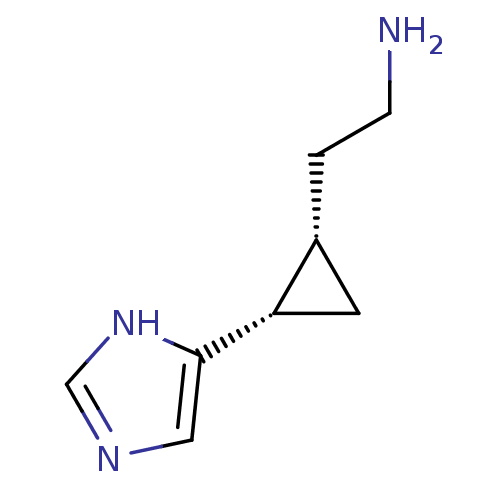
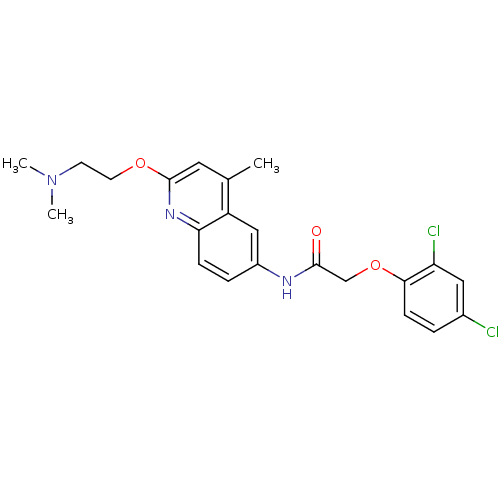
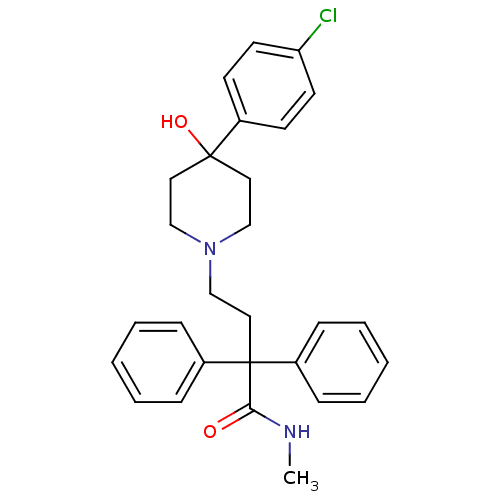
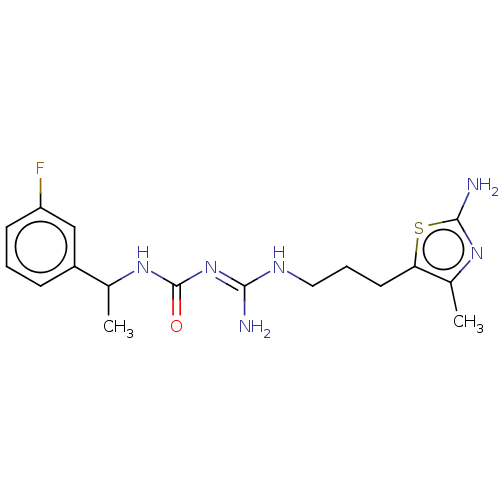
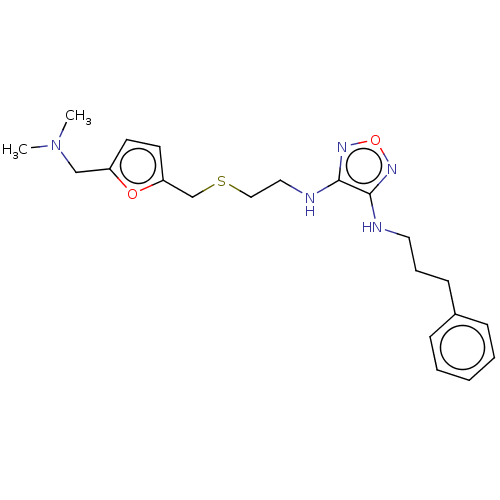
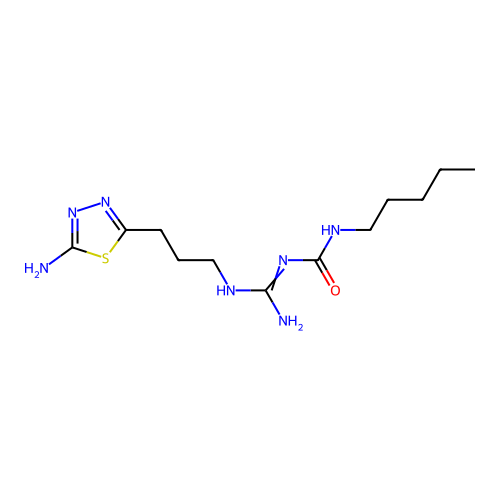
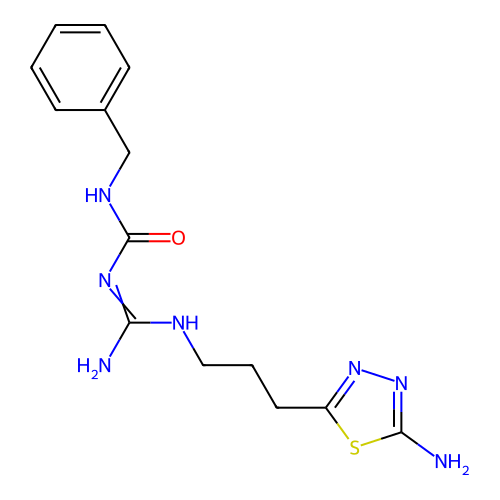
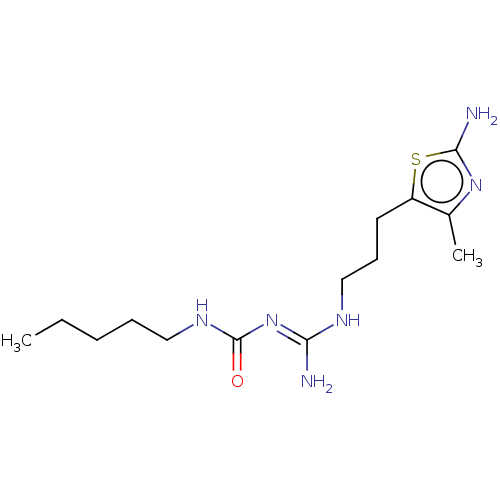
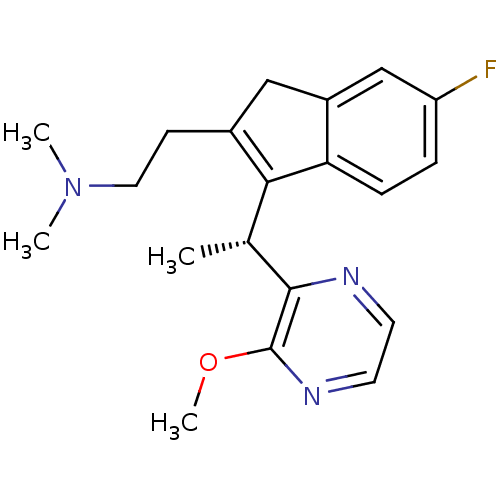
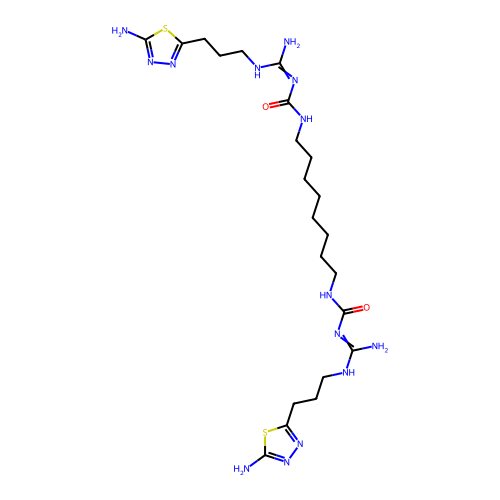
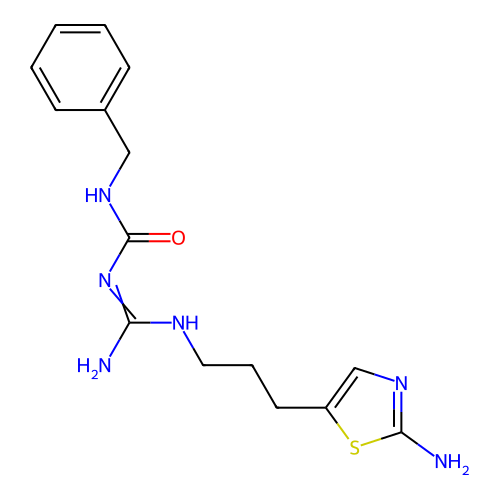
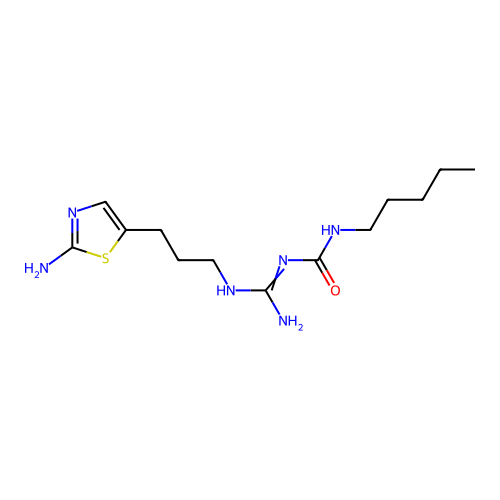
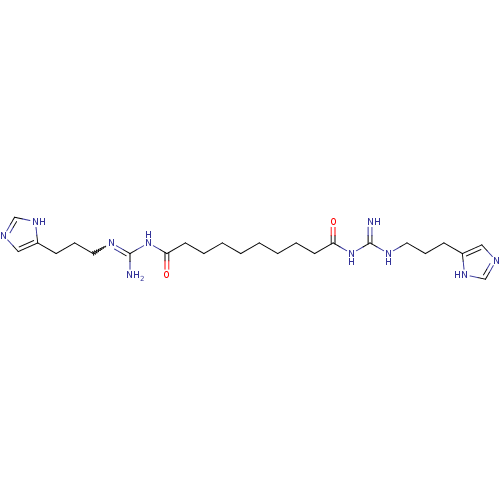
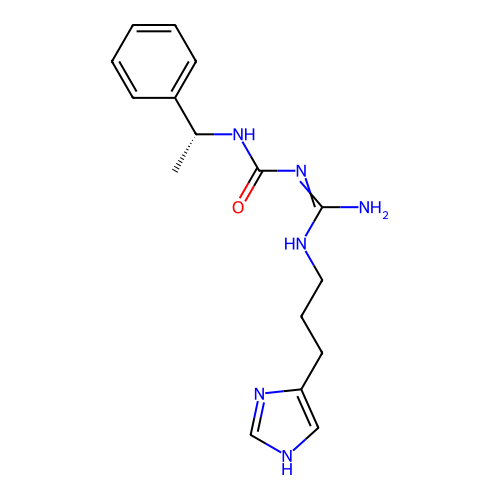
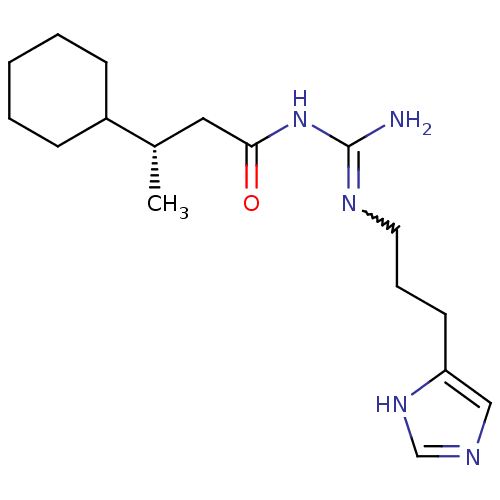
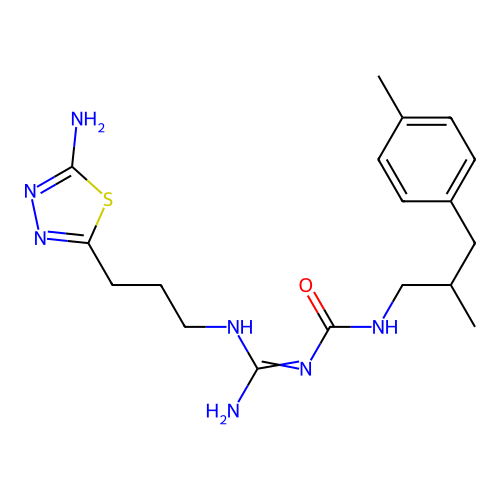
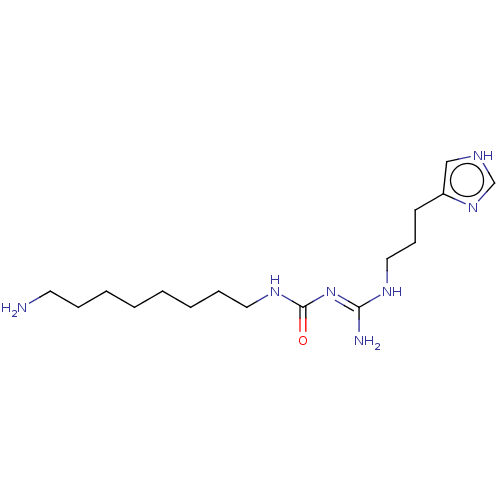
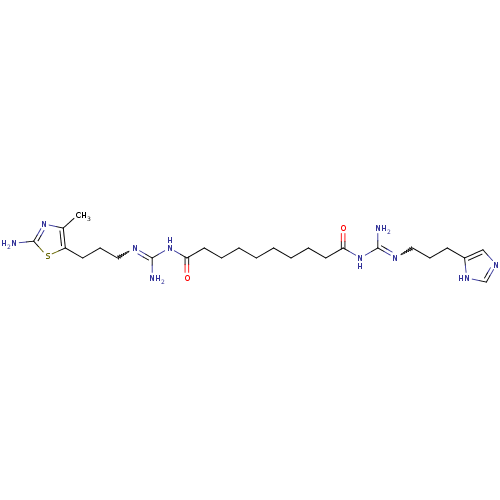
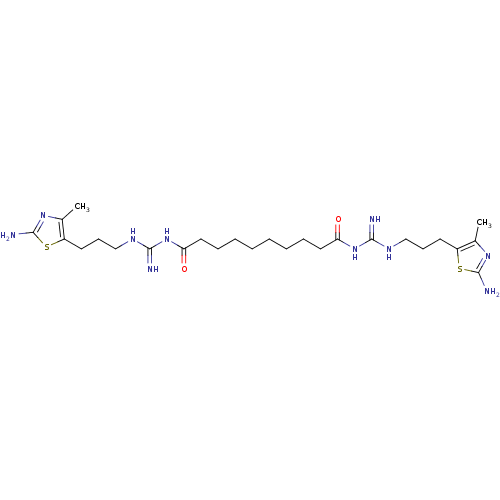
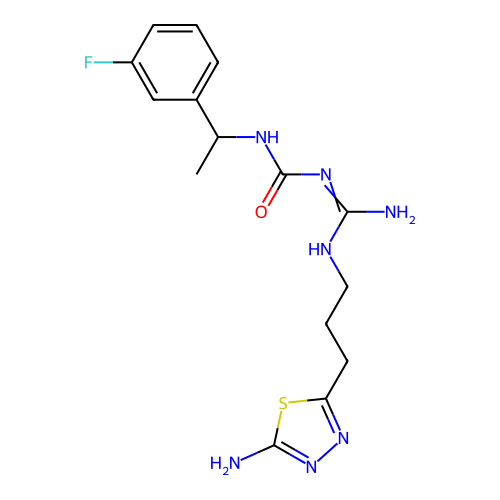
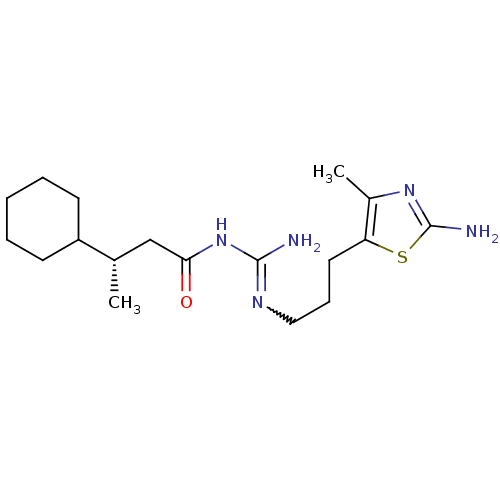
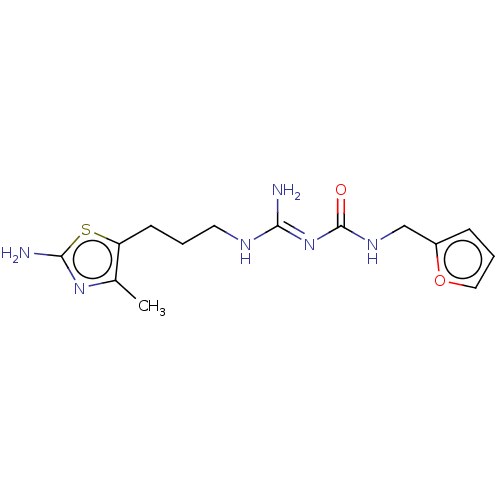
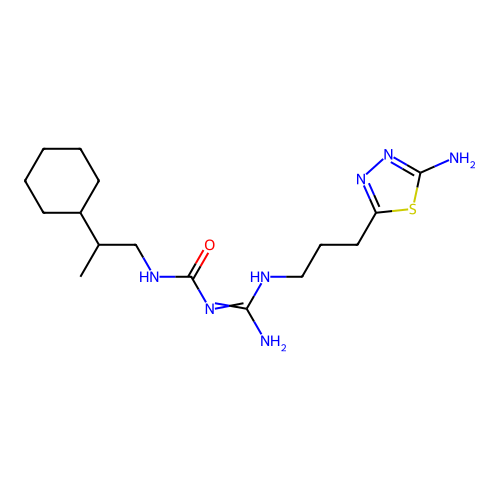
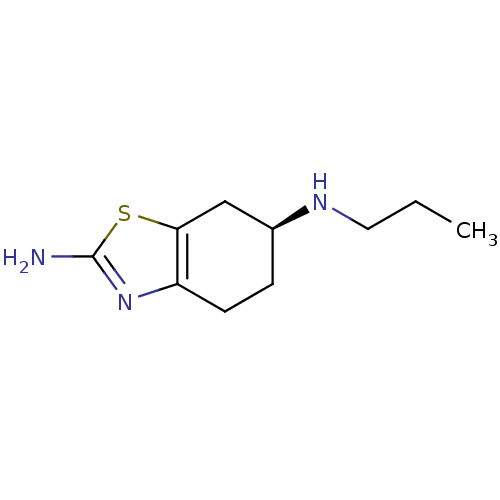
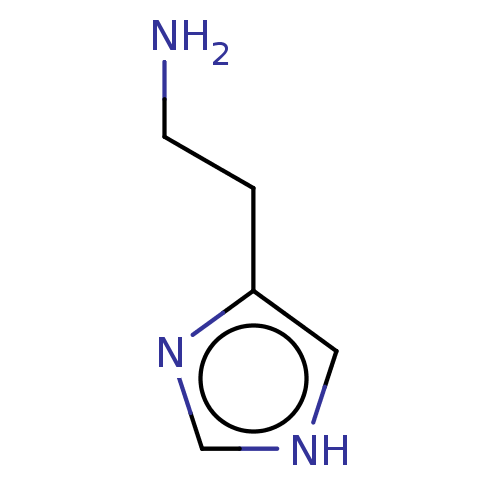
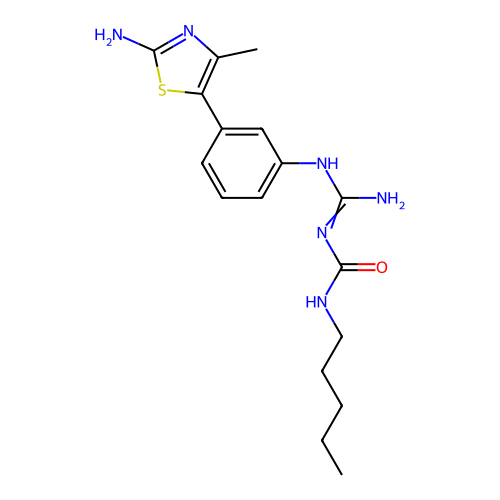
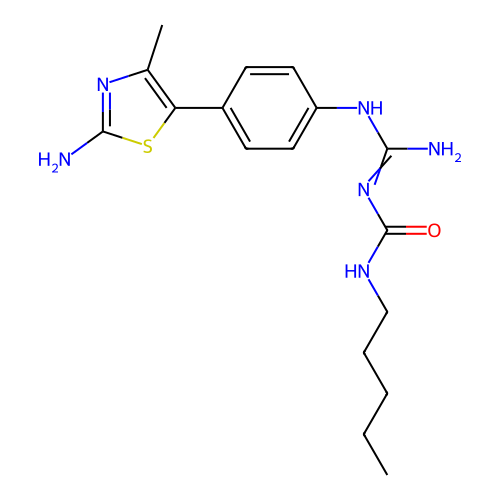
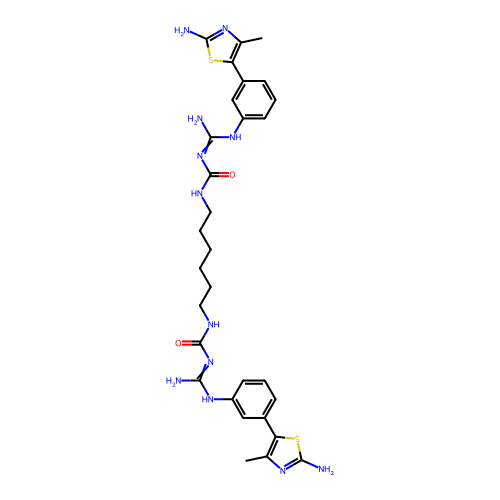
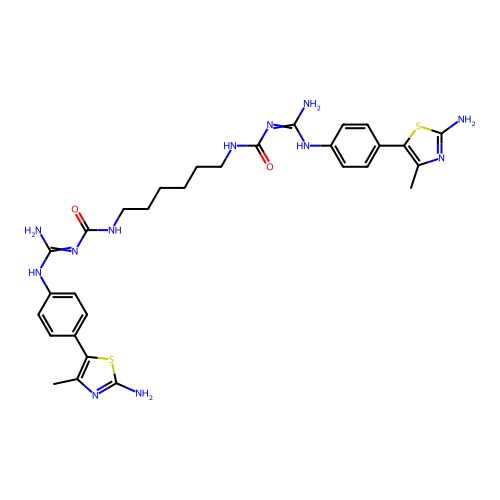
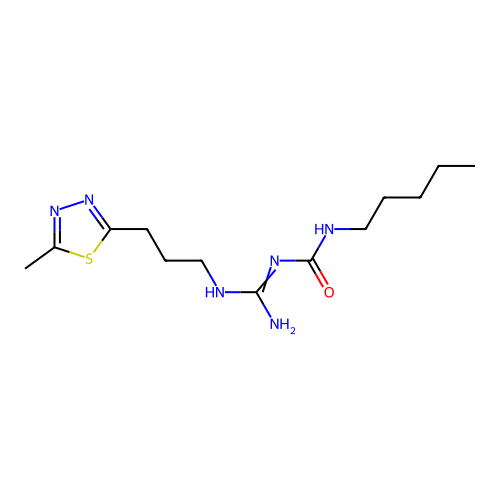
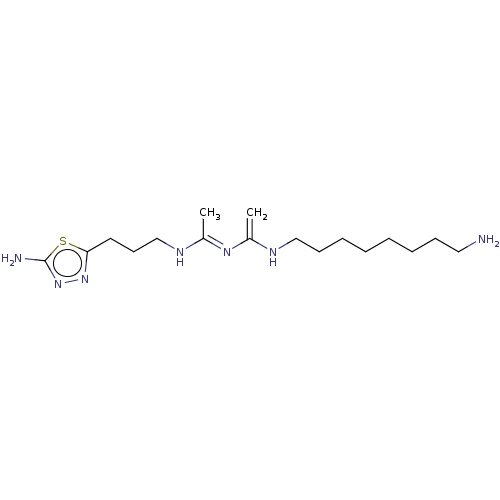Affinity DataEC50: >0.000100nMAssay Description:Inhibition of human Histamine H2 receptor using [3H]tiotidineMore data for this Ligand-Target Pair
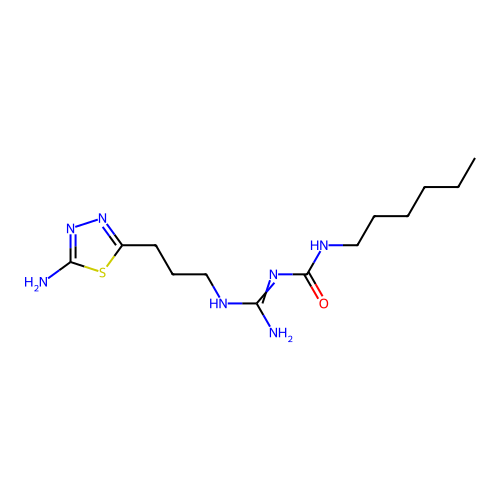
Affinity DataEC50: >0.000100nMAssay Description:Inhibition of human Histamine H2 receptor using [3H]tiotidineMore data for this Ligand-Target Pair
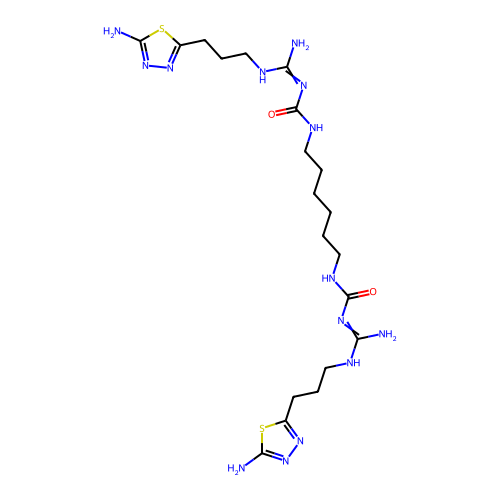
Affinity DataEC50: >0.000100nMAssay Description:Inhibition of human Histamine H2 receptor using [3H]tiotidineMore data for this Ligand-Target Pair
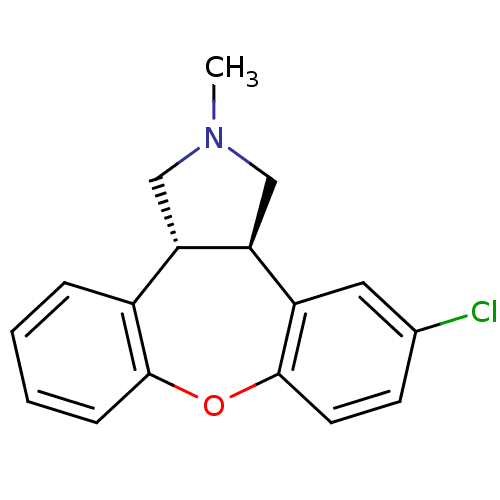
Affinity DataEC50: >0.000100nMAssay Description:Inhibition of human Histamine H2 receptor using [3H]tiotidineMore data for this Ligand-Target Pair
Affinity DataEC50: >0.000100nMAssay Description:Inhibition of human Histamine H2 receptor using [3H]tiotidineMore data for this Ligand-Target Pair
Affinity DataEC50: >0.000100nMAssay Description:Inhibition of human Histamine H2 receptor using [3H]tiotidineMore data for this Ligand-Target Pair
Affinity DataEC50: >0.000100nMAssay Description:Inhibition of human Histamine H2 receptor using [3H]tiotidineMore data for this Ligand-Target Pair
Affinity DataIC50: 0.870nMAssay Description:Inhibitory concentration against histamine H2 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Displacement of [3H]tiotidine from human histamine H2 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor expressed in sf9 insect cell membranes co-expressing GSalphaS incubated for 60 mins by ...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Antagonist activity at histamine H2 receptor (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataEC50: 3.30nMAssay Description:Agonist activity at human histamine H2 receptor stably expressed in HEK293T cells co-expressing NlucN-mGs/hH2R-NlucC assessed as induction of mini-Gi...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor expressed in sf9 insect cell membranes co-expressing GSalphaS incubated for 60 mins by ...More data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Inhibition of H2 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataEC50: 4.90nMAssay Description:Agonist activity at human histamine H2 receptor stably expressed in HEK293T cells co-expressing NlucN-mGs/hH2R-NlucC assessed as induction of mini-Gi...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKi: 5.10nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKi: 5.70nMAssay Description:Displacement of [3H]Cimetidine from histamine H2 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataEC50: 5.80nMAssay Description:Agonist activity at human histamine H2 receptor stably expressed in HEK293T cells co-expressing NlucN-mGs/hH2R-NlucC assessed as induction of mini-Gi...More data for this Ligand-Target Pair
Affinity DataKi: 5.80nMAssay Description:Displacement of [3H]Cimetidine from histamine H2 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataEC50: 6nMAssay Description:Agonist activity at human histamine H2 receptor stably expressed in HEK293T cells co-expressing NlucN-mGs/hH2R-NlucC assessed as induction of mini-Gi...More data for this Ligand-Target Pair
Affinity DataEC50: 6nMAssay Description:Agonist activity at human histamine H2 receptor stably expressed in HEK293T cells co-expressing NlucN-mGs/hH2R-NlucC assessed as induction of mini-Gi...More data for this Ligand-Target Pair
Affinity DataEC50: 6.17nMAssay Description:Agonist activity at human H2R-Gsalphas expressed in Sf9 cells at 0.1 nM to 10 uM by steady state GTPase activity assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.20nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataEC50: 6.46nMAssay Description:Agonist activity at GsalphaS-fused human histamine H2 receptor expressed in Sf9 cells by [35S]GTPgammaS binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.5nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataEC50: 7.40nMAssay Description:Agonist activity at human H2 receptor expressed in HEK293T-ARRB2 cells assessed as stimulation of beta-arrestin 2 recruitmentMore data for this Ligand-Target Pair
Affinity DataEC50: 7.59nMAssay Description:Agonist activity at human H2R-Gsalphas expressed in Sf9 cells at 0.1 nM to 10 uM by steady state GTPase activity assayMore data for this Ligand-Target Pair
Affinity DataEC50: 7.60nMAssay Description:Agonist activity at human H2 receptor expressed in HEK293T-ARRB2 cells assessed as stimulation of beta-arrestin 2 recruitmentMore data for this Ligand-Target Pair
Affinity DataEC50: 7.76nMAssay Description:Agonist activity at human H2R-Gsalphas expressed in Sf9 cells at 0.1 nM to 10 uM by steady state GTPase activity assayMore data for this Ligand-Target Pair
Affinity DataEC50: 8.10nMAssay Description:Agonist activity at human histamine H2 receptor stably expressed in HEK293T cells co-expressing NlucN-mGs/hH2R-NlucC assessed as induction of mini-Gi...More data for this Ligand-Target Pair
Affinity DataKi: 8.10nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataEC50: 9.33nMAssay Description:Agonist activity at GsalphaS-fused human histamine H2 receptor expressed in Sf9 cells by [35S]GTPgammaS binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor expressed in sf9 insect cell membranes co-expressing GSalphaS incubated for 60 mins by ...More data for this Ligand-Target Pair
Affinity DataEC50: 11nMAssay Description:Agonist activity at human histamine H2 receptor stably expressed in HEK293T cells co-expressing NlucN-mGs/hH2R-NlucC assessed as induction of mini-Gi...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [3H]UR-DE257 from human histamine H2 receptor stably expressed in baculovirus infected Sf9 cell membrane co-expressing RG-Salpha S in...More data for this Ligand-Target Pair