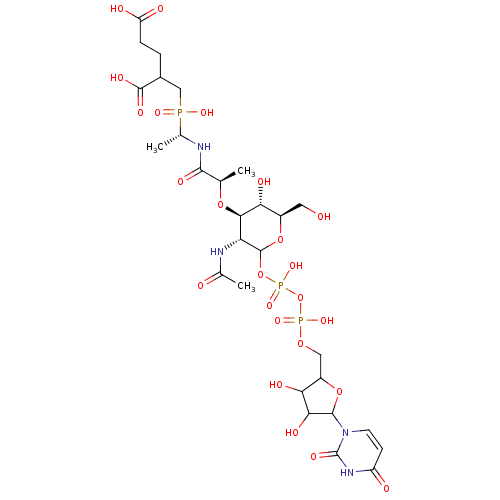
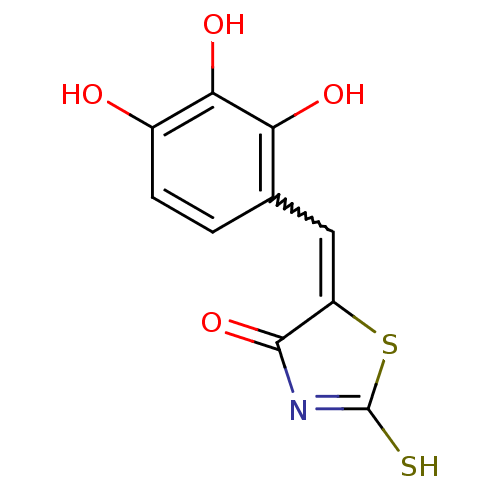
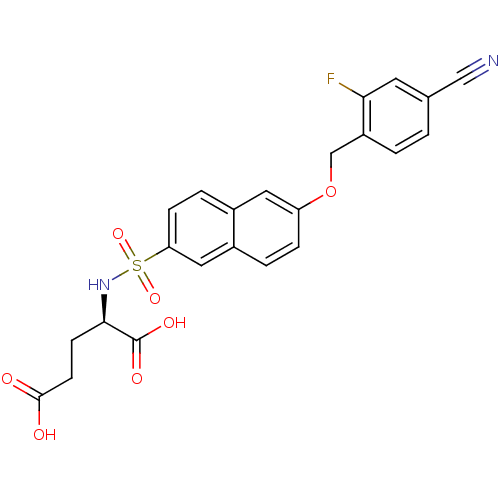
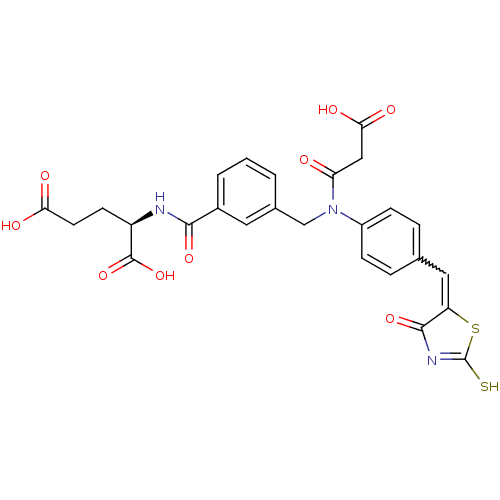
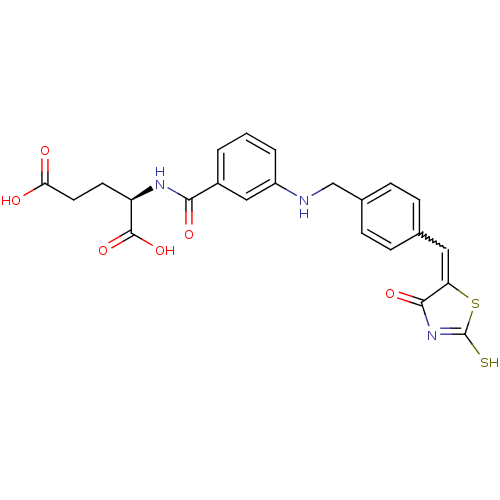
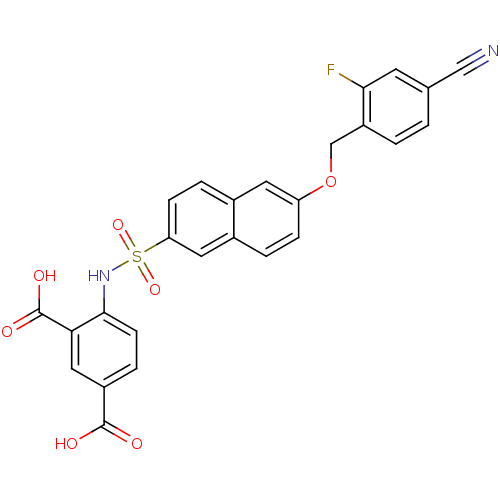
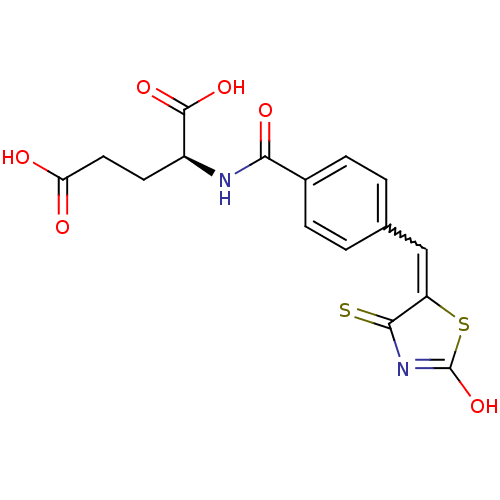
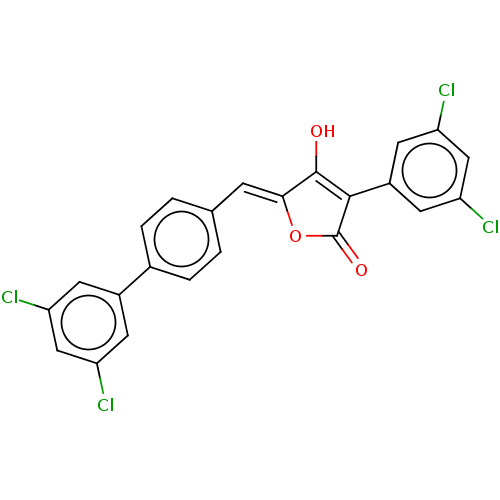
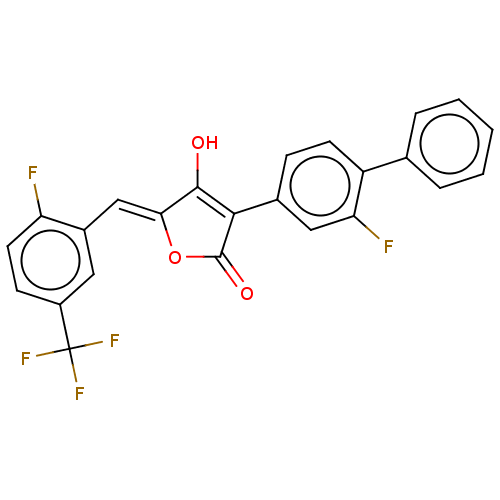
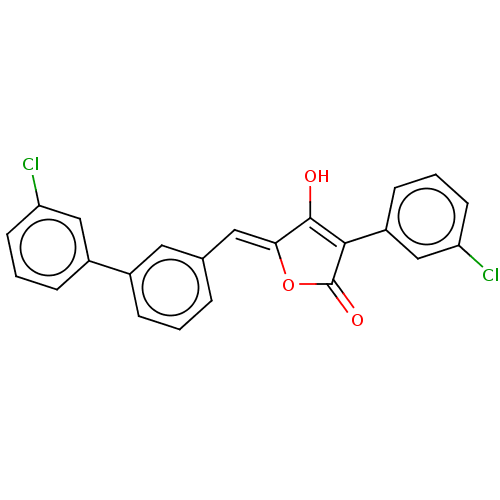
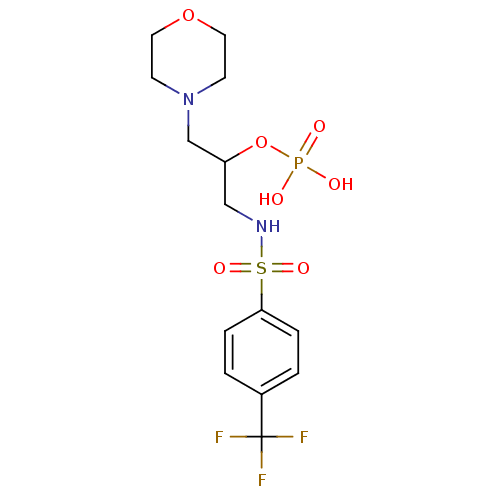
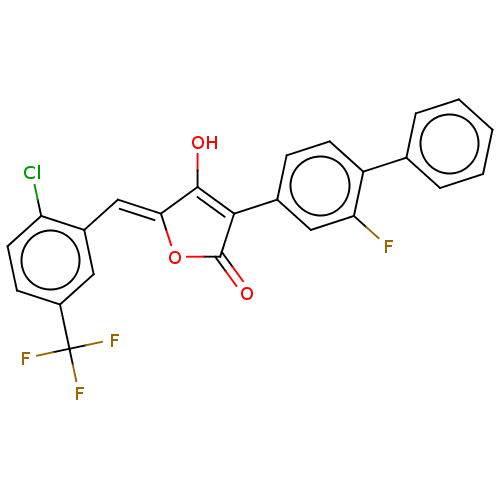
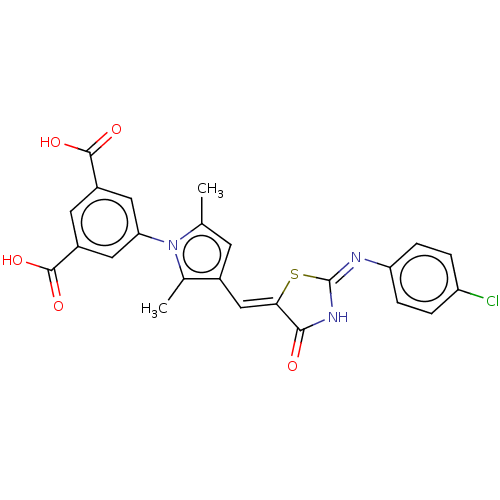
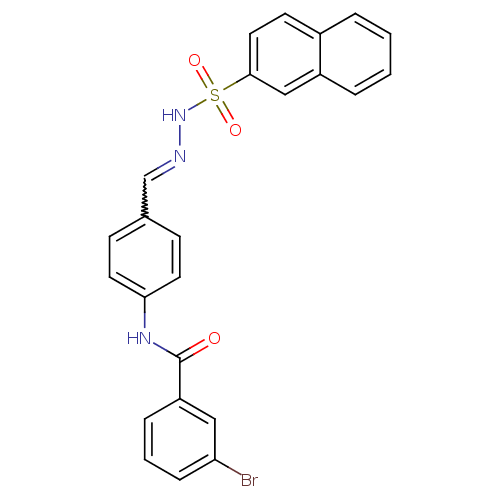
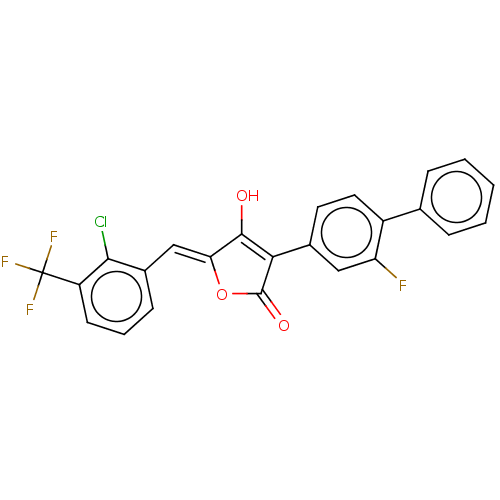
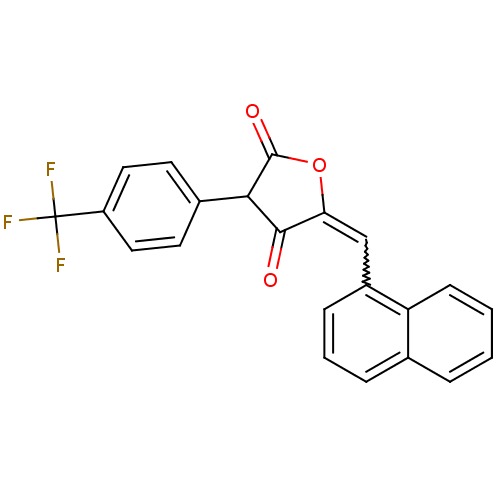
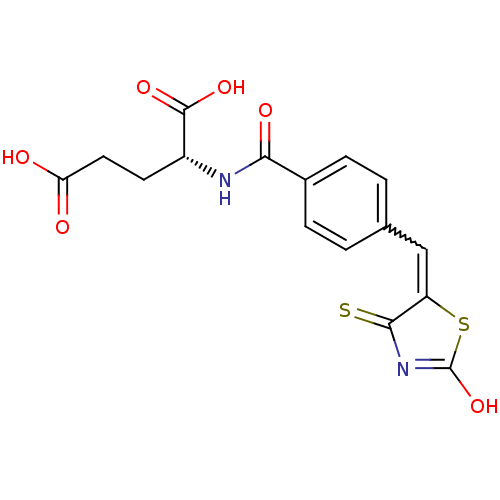
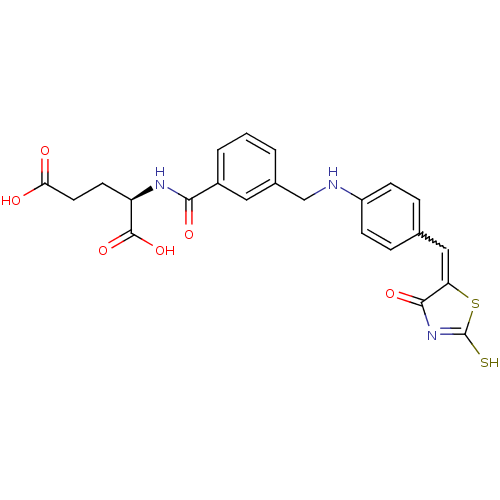
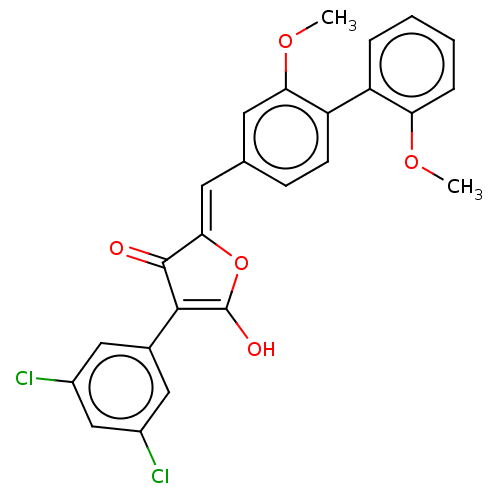
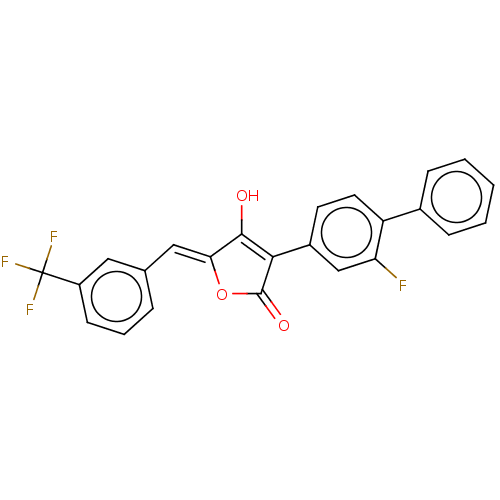
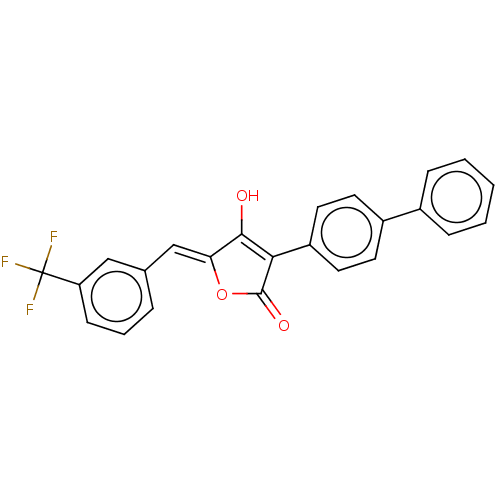
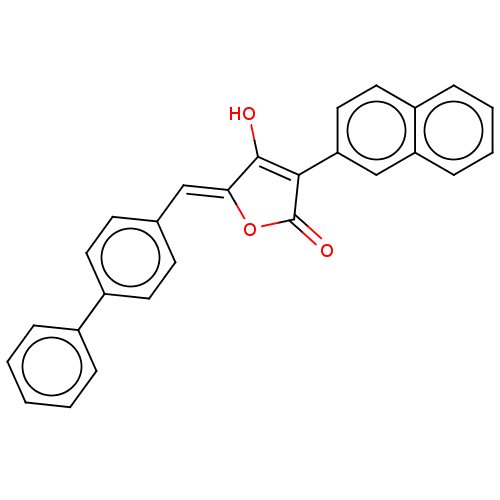
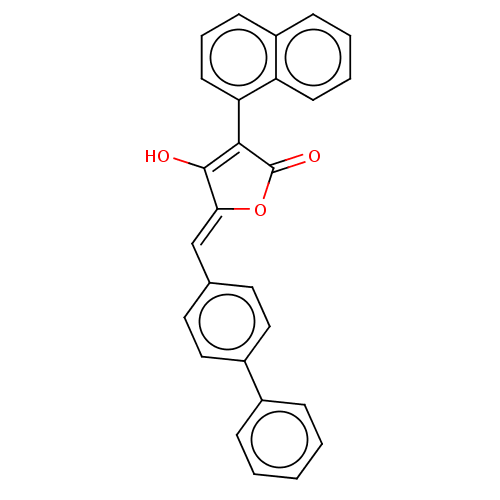
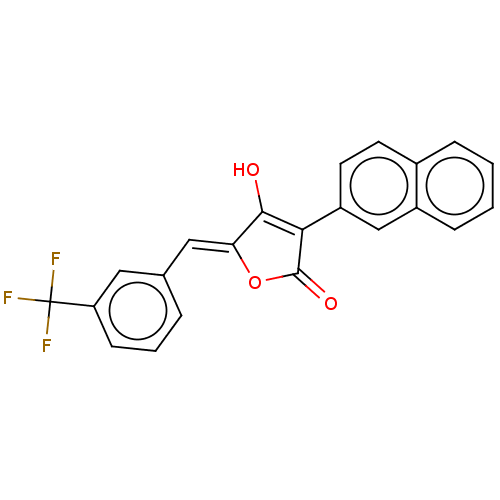
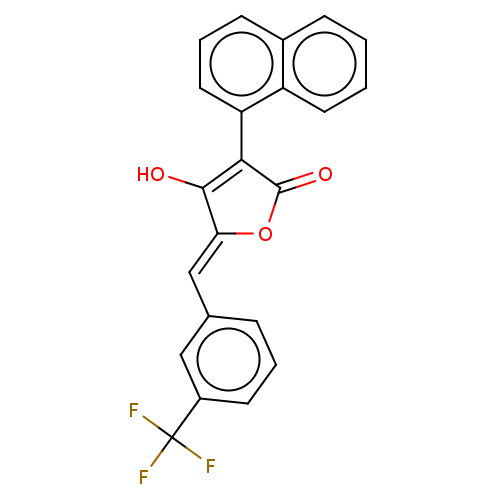
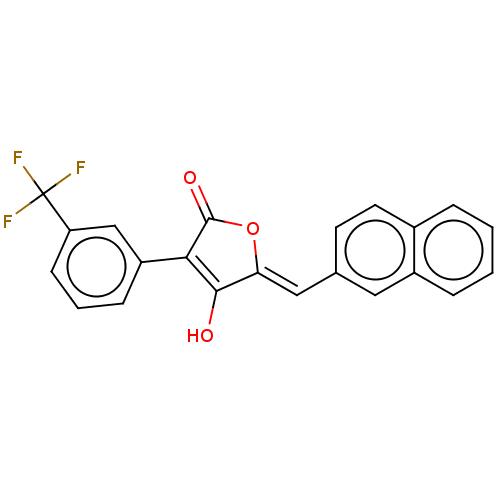
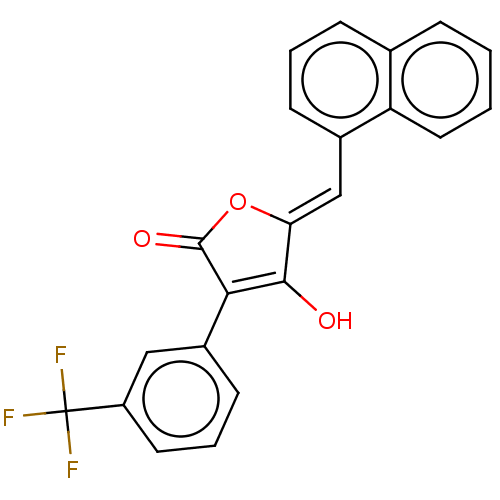
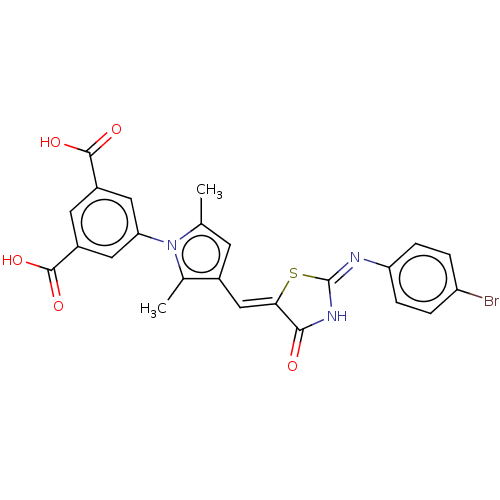
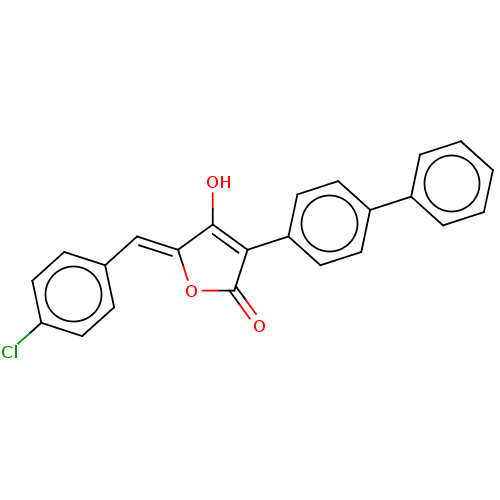
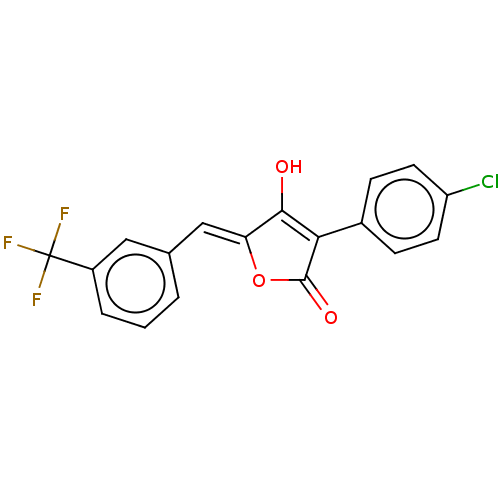
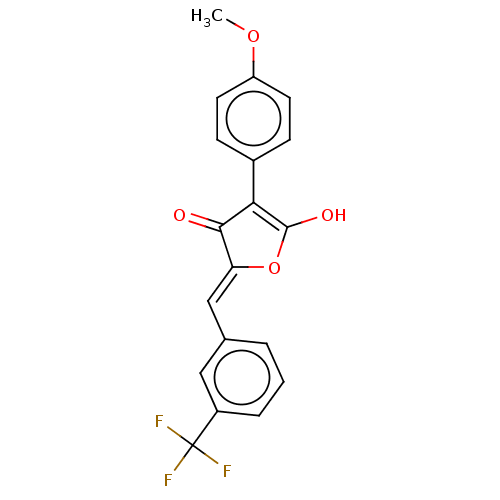
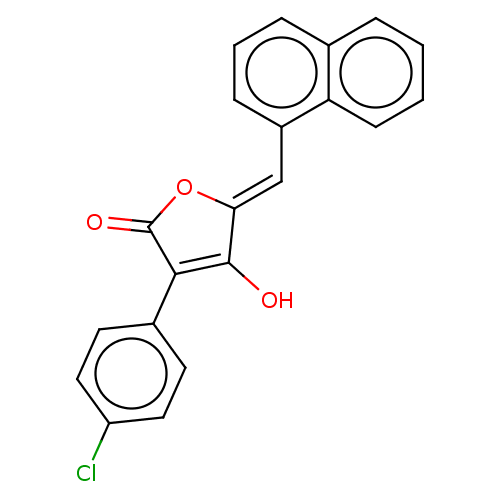
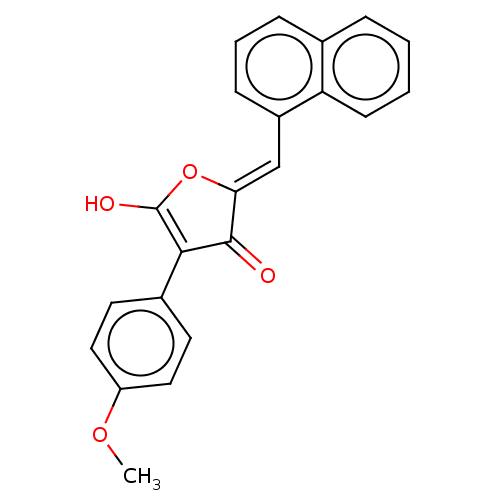
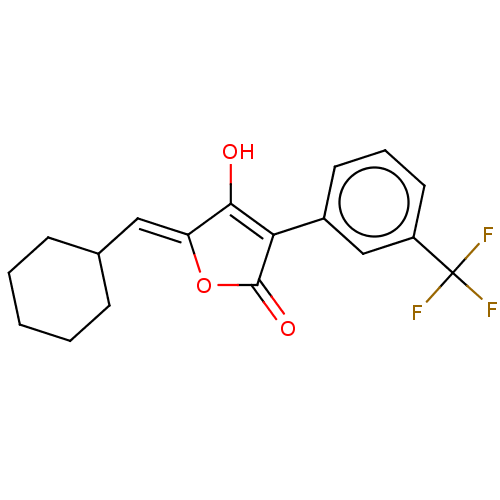
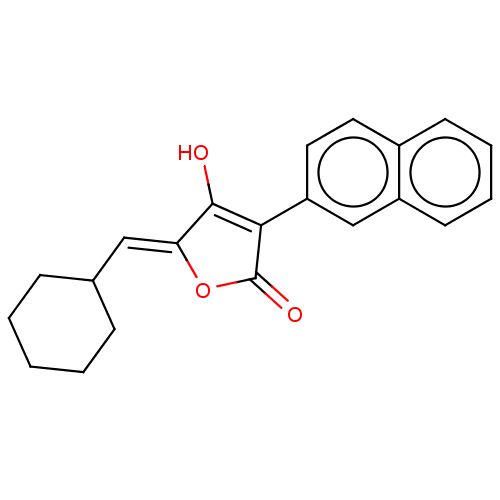
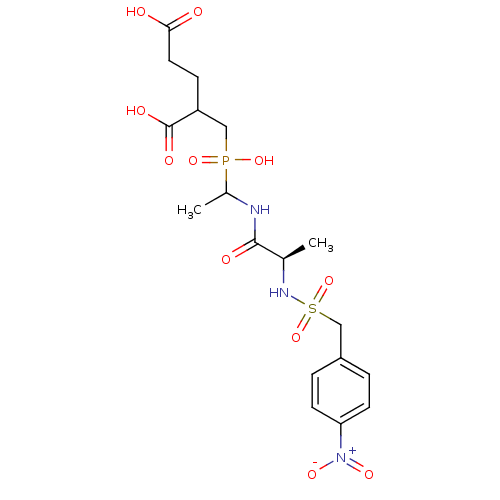
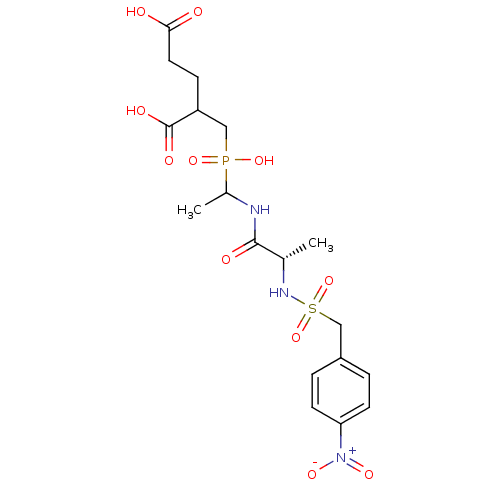
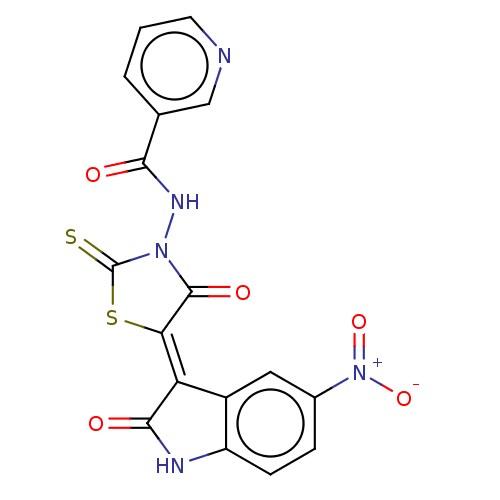
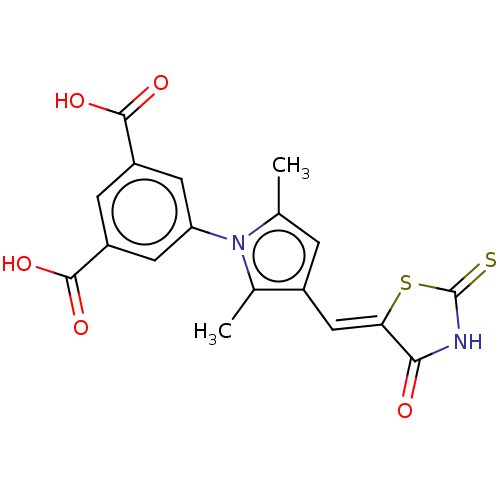
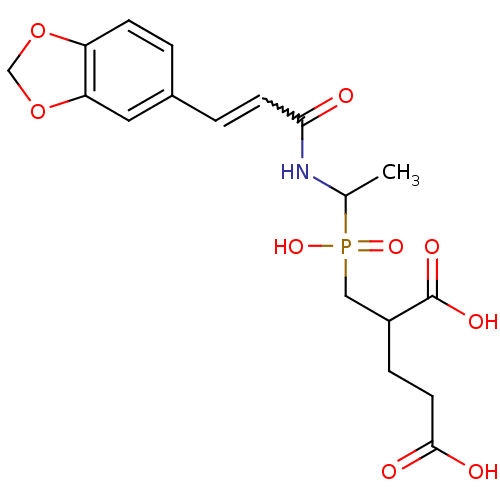
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKd: 2.20E+3nMAssay Description:Binding affinity to 6x His-tagged and 15N/13C-labeled Escherichia coli MurD ligase measured at NMR signal 9 by 1H/13C-HSQC 2D NMR spectroscopyMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKd: 5.50E+3nMAssay Description:Binding affinity to 6x His-tagged and 15N/13C-labeled Escherichia coli MurD ligase measured at NMR signal 56 by 1H/13C-HSQC 2D NMR spectroscopyMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKd: 7.00E+3nMAssay Description:Binding affinity to 6x His-tagged and 15N/13C-labeled Escherichia coli MurD ligase measured at NMR signal 13 by 1H/13C-HSQC 2D NMR spectroscopyMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.35E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.46E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.95E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKi: 2.10E+4nMAssay Description:Competitive inhibition of Escherichia coli MurD using D-Glu substrate by steady-state enzyme kinetics assayMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKi: 2.90E+4nMAssay Description:Partial non-competitive inhibition of Escherichia coli MurD using UMA substrate by steady-state enzyme kinetics assayMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 3.47E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKi: 3.90E+4nMAssay Description:Partial un-competitive inhibition of Escherichia coli MurD using ATP substrate by steady-state enzyme kinetics assayMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 5.32E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 5.86E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.00E+4nMAssay Description:Inhibition of Escherichia coli MurD after 15 mins in presence of 0.005% Triton X-114 by Malachite green assayMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.12E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.40E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.40E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.53E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.53E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.53E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.53E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.60E+4nMAssay Description:Inhibition of Escherichia coli MurD after 15 mins in presence of 0.005% Triton X-114 by Malachite green assayMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.66E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.81E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.90E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 7.16E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 7.25E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 7.38E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 7.80E+4nMAssay Description:Inhibitory activity against MurD in Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 7.80E+4nMAssay Description:Inhibitory activity against MurD from Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKd: 7.90E+4nMAssay Description:Binding affinity to 6x His-tagged and 15N/13C-labeled Escherichia coli MurD ligase measured at NMR signal 43 by 1H/13C-HSQC 2D NMR spectroscopyMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 8.50E+4nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 9.50E+4nMAssay Description:Inhibition of Escherichia coli MurD after 15 mins in presence of 0.005% Triton X-114 by Malachite green assayMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 9.50E+4nMAssay Description:Inhibitory activity against MurD from Escherichia coliMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataKd: 9.80E+4nMAssay Description:Binding affinity to 6x His-tagged and 15N/13C-labeled Escherichia coli MurD ligase by 1H/13C-HSQC 2D NMR spectroscopyMore data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.00E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair